

Chem. J. Chinese Universities ›› 2023, Vol. 44 ›› Issue (3): 20220173.doi: 10.7503/cjcu20220173
• Article • Previous Articles Next Articles
ZHANG Qingyi, CAO Jie, SHU Xiao, LIU Jianzhao( )
)
Received:2022-03-21
Online:2023-03-10
Published:2023-03-14
Contact:
LIU Jianzhao
E-mail:liujz@zju.edu.cn
Supported by:CLC Number:
TrendMD:
ZHANG Qingyi, CAO Jie, SHU Xiao, LIU Jianzhao. Effects of Exogenous N6-methyladenosine Incorporation on the Expression of Cellular mRNA Transcripts[J]. Chem. J. Chinese Universities, 2023, 44(3): 20220173.
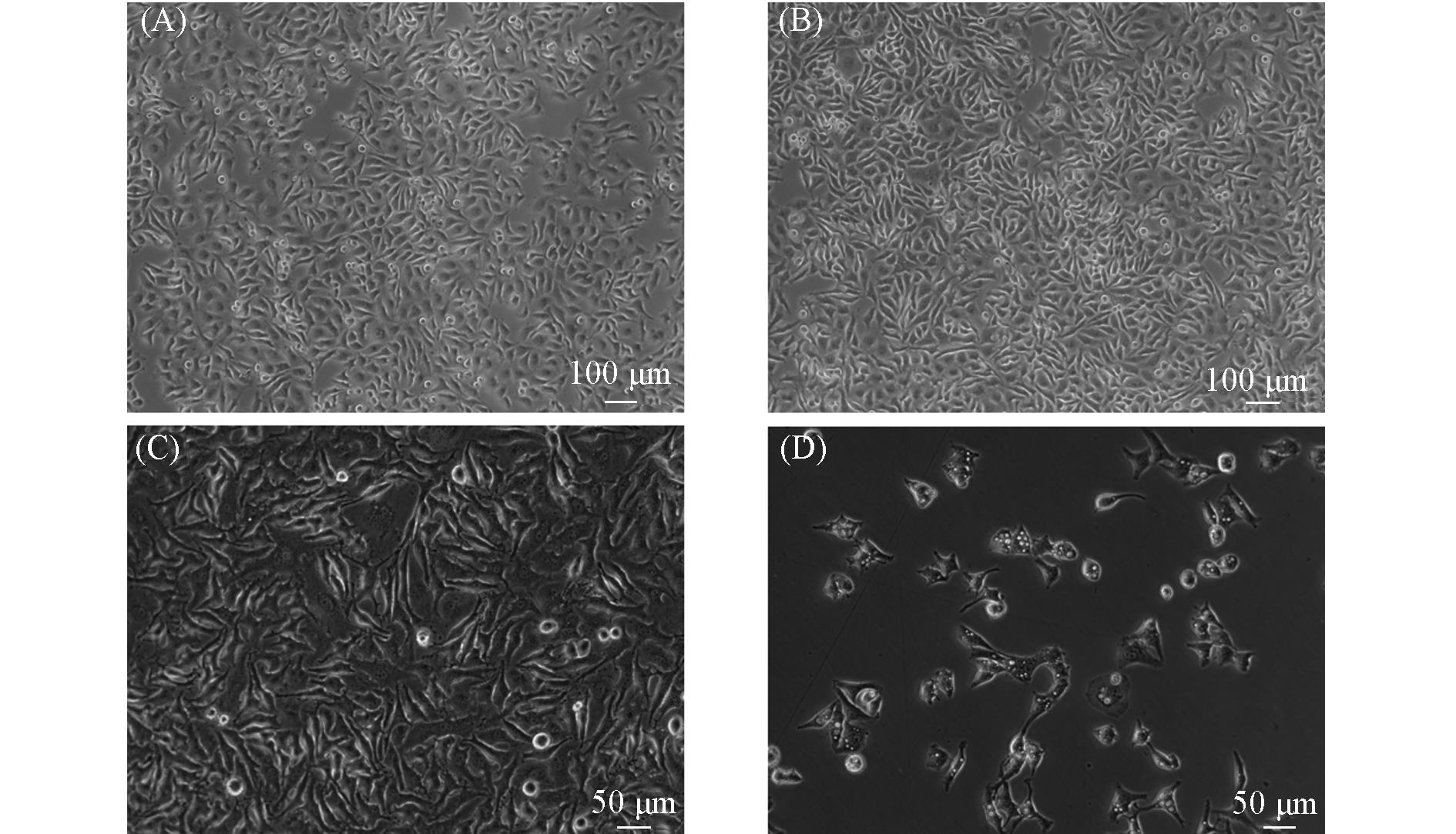

Fig.1 Cell morphology of HeLa cells treated with m6A at different concentrations for 12 h(A) DMSO as control; (B) 50 μmol/L m6A; (C) 500 μmol/L m6A; (D) 1 mmol/L m6A.
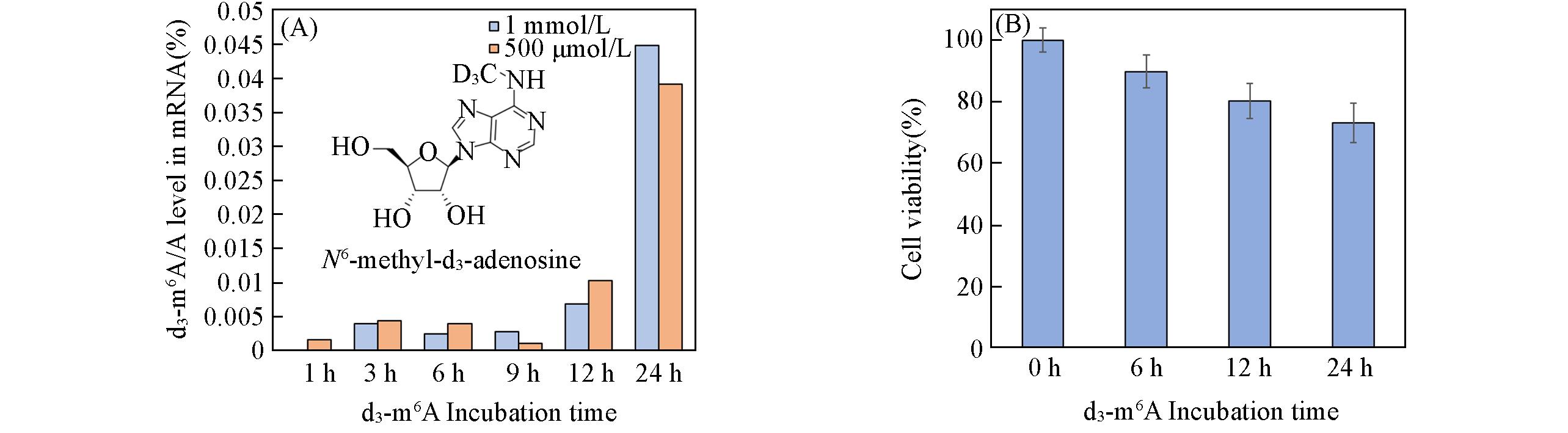
Fig.2 Mass spectrometry characterization of d3⁃m6A/A ratio in mRNA of HeLa cells(A) and cytotoxicity of d3⁃m6A in HeLa cells(B)(A) The cells were treated with 500 μmol/L or 1 mmol/L d3⁃m6A for different time intervals; (B) the cells were treated with 500 μmol/L d3⁃m6A for different time intervals. Data are expressed as mean±SD, n=3.
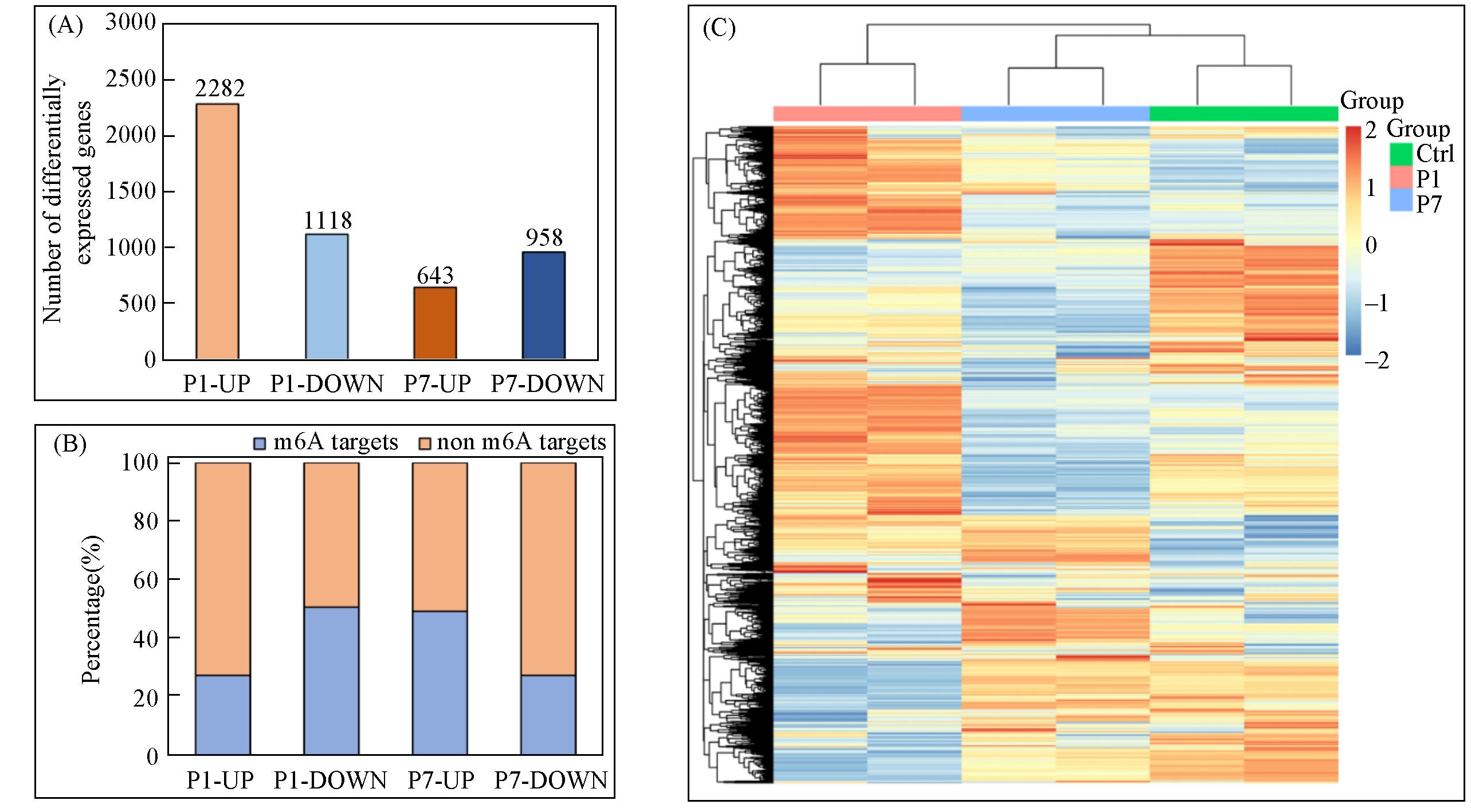
Fig.3 RNA⁃seq analysis showing the abundance change of mRNA transcripts under the metabolic incorporation of d3⁃m6A into mRNA of HeLa cells(A) The number of differentially expressed genes in samples under the metabolic feeding of d3-m6A for different time intervals versus control(Ctrl) without treatment. P1: 500 μmol/L d3-m6A, 24 h treatment; P7: 500 μmol/L d3-m6A, 168 h treatment. UP: up-regulated genes; DOWN: down-regulated genes. (B) The percentage of m6A-containing target genes among the total differentially expressed genes of the samples. (C) The heatmap and cluster analyses of differentially expressed genes in the samples.
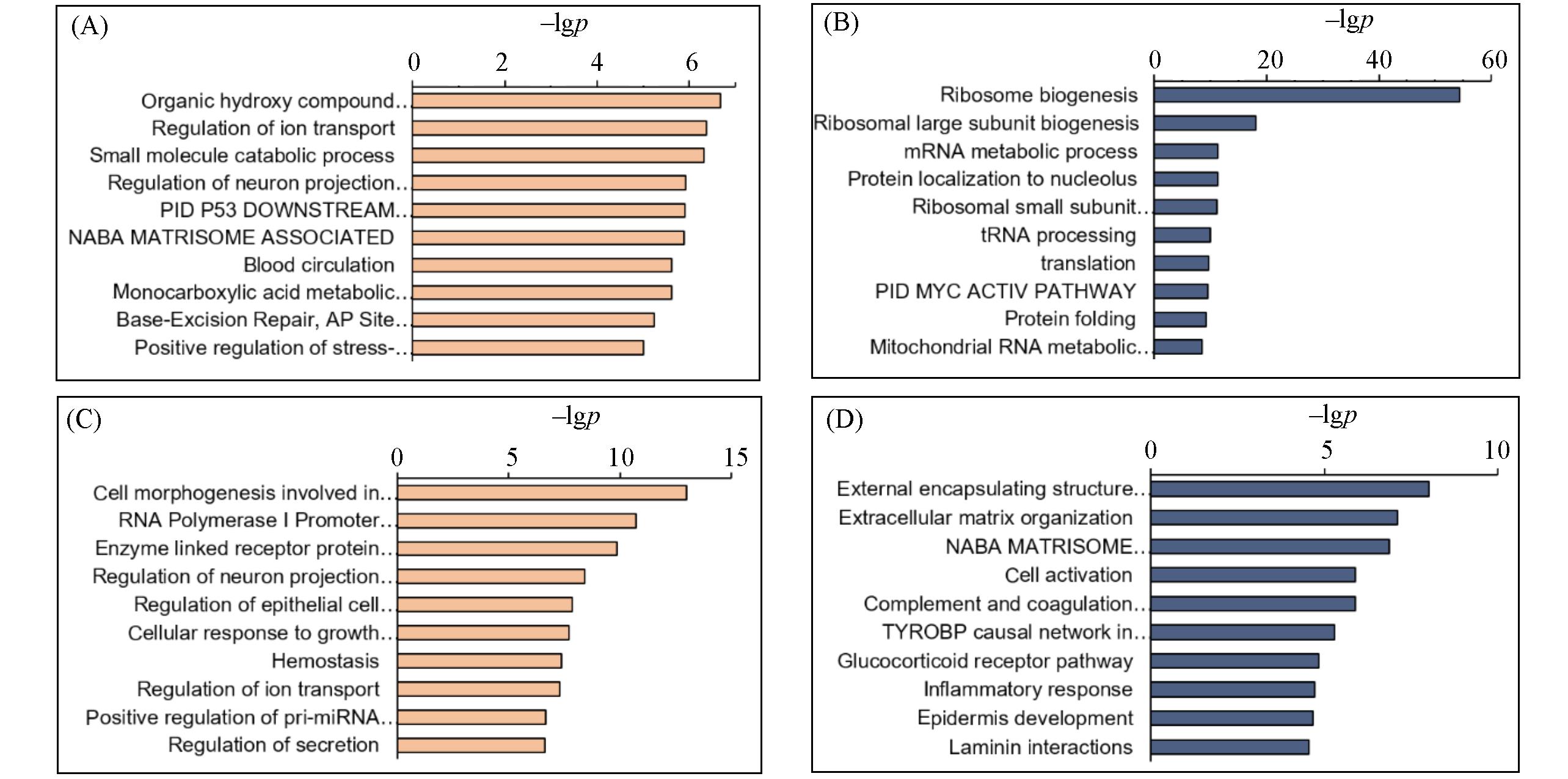
Fig.4 Gene ontology analysis of differentially expressed genes in d3⁃m6A⁃treated HeLa cells for different time intervals(A) P1-UP; (B) P1-DOWN; (C) P7-UP; (D) P7-DOWN. P1: 500 μmol/L d3-m6A, 24 h treatment; P7: 500 μmol/L d3-m6A, 168 h treatment; UP: up-regulated genes; DOWN: down-regulated genes.
| 1 | Zhao B. X. S., Roundtree I. A., He C., Nat. Rev. Mol. Cell Biol., 2017, 18, 31—42 |
| 2 | Meyer K. D., Saletore Y., Zumbo P., Elemento O., Mason C. E., Jaffrey S. R., Cell, 2012, 149, 1635—1646 |
| 3 | Desrosiers R., Friderici K., Rottman F., Proc. Natl. Acad. Sci. USA, 1974, 71, 3971—3975 |
| 4 | Yue Y. N., Liu J. Z., He C., Genes Dev., 2015, 29, 1343—1355 |
| 5 | Roundtree I. A., Evans M. E., Pan T., He C., Cell, 2017, 169, 1187—1200 |
| 6 | Liu J., Harada B. T., He C., Trends Cell Biol., 2019, 29, 487—499 |
| 7 | Liu J. Z., Yue Y. N., Han D. L., Wang X., Fu Y., Zhang L., Jia G. F., Yu M., Lu Z. K., Deng X., Dai Q., Chen W. Z., He C., Nat. Chem. Biol., 2014, 10, 93—95 |
| 8 | Knuckles P., Lence T., Haussmann I. U., Jacob D., Kreim N., Carl S. H., Masiello I., Hares T., Villasenor R., Hess D., rade⁃Navarro M. A., Biggiogera M., Helm M., Soller M., Buhler M., Roignant J. Y., Genes Dev., 2018, 32, 415—429 |
| 9 | Wen J., Lv R. T., Ma H. H., Shen H. J., He C. X., Wang J. H., Jiao F. F., Liu H., Yang P. Y., Tan L., Lan F., Shi Y., He C., Shi Y., Diao J. B., Mol. Cell, 2018, 69, 1028 |
| 10 | Warda A. S., Kretschmer J., Hackert P., Lenz C., Urlaub H., Hobartner C., Sloan K. E., Bohnsack M. T., EMBO Rep., 2017, 18, 2004—2014 |
| 11 | Jia G. F., Fu Y., Zhao X., Dai Q., Zheng G. Q., Yang Y., Yi C. Q., Lindahl T., Pan T., Yang Y. G., He C., Nat. Chem. Biol., 2011, 7, 885—887 |
| 12 | Zheng G. Q., Dahl J. A., Niu Y. M., Fedorcsak P., Huang C. M., Li C. J., Vagbo C. B., Shi Y., Wang W. L., Song S. H., Lu Z. K., Bosmans R. P. G., Dai Q., Hao Y. J., Yang X., Zhao W. M., Tong W. M., Wang X. J., Bogdan F., Furu K., Fu Y., Jia G. F., Zhao X., Liu J., Krokan H. E., Klungland A., Yang Y. G., He C., Mol. Cell, 2013, 49, 18—29 |
| 13 | Patil D. P., Chen C. K., Pickering B. F., Chow A., Jackson C., Guttman M., Jaffrey S. R., Nature, 2016, 537, 369 |
| 14 | Wang T. Y., Kong S., Tao M., Ju S. Q., Mol. Cancer, 2020, 19, 18 |
| 15 | Xiang Y., Laurent B., Hsu C. H., Nachtergaele S., Lu Z. K., Sheng W. Q., Xu C. Y., Chen H., Ouyang J., Wang S. Q., Ling D., Hsu P. H., Zou L., Jambhekar A., He C., Shi Y., Nature, 2017, 552, 430—430 |
| 16 | Batista P. J., Molinie B., Wang J. K., Qu K., Zhang J. J., Li L. J., Bouley D. M., Lujan E., Haddad B., Daneshvar K., Carter A. C., Flynn R. A., Zhou C., Lim K. S., Dedon P., Wernig M., Mullen A. C., Xing Y., Giallourakis C. C., Chang H. Y., Cell Stem Cell, 2014, 15, 707—719 |
| 17 | Geula S., Moshitch⁃Moshkovitz S., Dominissini D., Mansour A. A., Kol N., Salmon⁃Divon M., Hershkovitz V., Peer E., Mor N., Manor Y. S., Ben⁃Haim M. S., Eyal E., Yunger S., Pinto Y., Jaitin D. A., Viukov S., Rais Y., Krupalnik V., Chomsky E., Zerbib M., Maza I., Rechavi Y., Massarwa R., Hanna S., Amit I., Levanon E. Y., Amariglio N., Stern⁃Ginossar N., Novershtern N., Rechavi G., Hanna J. H., Science, 2015, 347, 1002—1006 |
| 18 | Li H. B., Tong J. Y., Zhu S., Batista P. J., Duffy E. E., Zhao J., Bailis W., Cao G. C., Kroehling L., Chen Y. Y., Wang G., Broughton J. P., Chen Y. G., Kluger Y., Simon M. D., Chang H. Y., Yin Z. N., Flavell R. A., Nature, 2017, 548, 338 |
| 19 | Han D. L., Liu J., Chen C. Y., Dong L. H., Liu Y., Chang R. B., Huang X. N., Liu Y. Y., Wang J. Y., Dougherty U., Bissonnette M. B., Shen B., Weichselbaum R. R., Xu M. M., He C., Nature, 2019, 566, 270 |
| 20 | Shu X., Cao J., Cheng M. H., Xiang S. Y., Gao M. S., Li T., Ying X. E., Wang F. Q., Yue Y. N., Lu Z. K., Dai Q., Cui X. L., Ma L. J., Wang Y. Z., He C., Feng X. H., Liu J. Z., Nat. Chem. Biol., 2020, 16, 887 |
| 21 | Zhao W., Qi X. Q., Liu L. N., Ma S. Q., Liu J. W., Wu J., Mol. Ther.⁃Nucl. Acids, 2020, 19, 405—412 |
| 22 | Li X. Y., Xiong X. S., Yi C. Q., Nat. Methods, 2017, 14, 23—31 |
| 23 | Helm M., Motorin Y., Nat. Rev. Genet., 2017, 18, 275—291 |
| 24 | Cao J., Shu X., Feng X. H., Liu J. Z., Curr. Opin. Chem. Biol., 2021, 63, 28—37 |
| 25 | Shu X., Cao J., Liu J., STAR Protocols, 2022, 3, 101096—101096 |
| 26 | Lane A. N., Fan T. W. M., Nucleic Acids Research, 2015, 43, 2466—2485 |
| 27 | Chen M. J., Urs M. J., Sanchez⁃Gonzalez I., Olayioye M. A., Herde M., Witte C. P., Plant Cell, 2018, 30, 1511—1522 |
| 28 | Jiang Z., Wang C., Wu Z., Chen K., Yang W., Deng H., Song H., Zhou X., Nucleic Acids Research, 2021, 49, 12048—12068 |
| 29 | Pertea M., Kim D., Pertea G. M., Leek J. T., Salzberg S. L., Nat. Protoc., 2016, 11, 1650—1667 |
| [1] | KONG Hao, XU Feiyang, WANG Yixiang, ZHANG Yan. Research Progress of CRISPR-Cas9 Functional Regulation System Based on Small Molecule Reaction Tools [J]. Chem. J. Chinese Universities, 2023, 44(3): 20220346. |
| [2] | KUANG Xiaojun, YI Jingwei, FANG Xiaoxia, LAI Dongmei, XU Hong. Preparation of Water-soluble Coumarin Fluorescent Substrate and Its Application in Droplet Based Digital Detection [J]. Chem. J. Chinese Universities, 2021, 42(11): 3537. |
| [3] | XIE Chen, CHEN Na, YANG Yanbing, YUAN Quan. Recent Progress of Aptamer Functionalized Two-dimensional Materials Field Effect Transistor Sensors [J]. Chem. J. Chinese Universities, 2021, 42(11): 3406. |
| [4] | LIU Yuan, DENG Jinqi, ZHAO Shuai, TIAN Fei, LI Yi, SUN Jiashu, LIU Chao. Lateral Flow Assay Based on Molecular Recognition for Diagnosis of Corona Virus Disease 2019 Infection [J]. Chem. J. Chinese Universities, 2021, 42(11): 3390. |
| [5] | MAO Yu, QU Hao, ZHENG Lei. Research Progress on RNA⁃cleaving DNAzyme for the Detection of Pathogenic Bacteria [J]. Chem. J. Chinese Universities, 2021, 42(11): 3445. |
| [6] | ZHANG Xiaorong, CHEN Lanlan, HU Shanwen. Advances in Bacteria Biosensing Based on Molecular Recognition [J]. Chem. J. Chinese Universities, 2021, 42(11): 3468. |
| [7] | XUE Jin, CAO Xiaowei, LIU Yifan, WANG Min. Preparation of Paper Hollow Gold Nanocage SERS Sensor and Its Rapid and Highly Sensitive Detection for miRNAs in Sputum of Patients with Non-small Cell Lung Cancer [J]. Chem. J. Chinese Universities, 2021, 42(8): 2393. |
| [8] | ZHENG Haijiao, JIANG Liyan, JIA Qiong. Arginine Functionalized Magnetic Nanoparticles and Its Application in Phosphopeptides Enrichment [J]. Chem. J. Chinese Universities, 2021, 42(3): 717. |
| [9] | LI Huiyuan, LEI Chunyang, HUANG Yan, NIE Zhou. Structural Modification of Fluorescent Proteins and Their Applications in Biosensing [J]. Chem. J. Chinese Universities, 2020, 41(11): 2324. |
| [10] | ZHANG Zuoran, ZHANG Li, ZHANG Zhiling. On-chip Sorting of Beads with Different Magnetic Responsiveness by Lateral Magnetophoresis † [J]. Chem. J. Chinese Universities, 2020, 41(6): 1243. |
| [11] | ZHANG Xiaofei, WU Lie, LI Shanshan, ZHU Manyu, CHENG Xiaowei, JIANG Xiu’e. Effect of Phase Behavior of Phospholipids on Lipid Membrane Damage Induced by Graphene Oxide † [J]. Chem. J. Chinese Universities, 2020, 41(4): 661. |
| [12] | WEI Manman,LU Haoxiang,YANG Huihua. Research on Boold Species Ide.pngication Algorithm Based on RF_AdaBoost Model † [J]. Chem. J. Chinese Universities, 2020, 41(1): 94. |
| [13] | XIAO Zhourui,HUANG Donghua,BIAN Mengmeng,YUAN Yali,NIE Jinfang. Fluorescence Detection Based on DNA-templated Silver Nanoclusters and the Construction of Multi-level Logic Gate † [J]. Chem. J. Chinese Universities, 2020, 41(1): 102. |
| [14] | ZHENG Shan, LIU Yang, CHEN Piaopiao, XING Yichen, HUANG Chaobiao. Novel Glutathione Photoelectrochemical Sensor Based on PbS QDs/TiO2 NPs † [J]. Chem. J. Chinese Universities, 2019, 40(9): 1866. |
| [15] | XIONG Qin,YANG Wuye,YANG Shaojie,LIU Jing,YANG Shihua,CHEN Wanchao,LI Long,DU Yiping. Rapid Detection of Trace Organophosphorus Pesticides by Membrane Enrichment Combined with Membrane Image Colorimetric† [J]. Chem. J. Chinese Universities, 2019, 40(6): 1135. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||