

Chem. J. Chinese Universities ›› 2018, Vol. 39 ›› Issue (2): 299.doi: 10.7503/cjcu20170402
• Physical Chemistry • Previous Articles Next Articles
XIN Meiling1,2, CHU Zhenhua1,2, LI Yu1,2,*( )
)
Received:2017-06-26
Online:2018-02-10
Published:2017-12-25
Contact:
LI Yu
E-mail:liyuxx@jlu.edu.cn
CLC Number:
TrendMD:
XIN Meiling, CHU Zhenhua, LI Yu. Molecular Modification of Polychlorinated Biphenyl Dihydroxy Derivatives Through Molecular Docking Associated with CoMSIA/HQSAR Models†[J]. Chem. J. Chinese Universities, 2018, 39(2): 299.
| No. | Compound | Kd | Similarity | No. | Compound | Kd | Similarity |
|---|---|---|---|---|---|---|---|
| 1a | 4,5-2OH-CB35 | 4.32 | 0.58 | 16a | 3,4-2OH-CB76 | 3.53 | 0.63 |
| 2,3-2OH-CB37 | 5.56 | 0.65 | 17a | 4,5-2OH-CB76 | 3.41 | 0.57 | |
| 3a | 5,6-2OH-CB37 | 2.03 | 0.57 | 18b | 2',3'-2OH-CB77 | 4.21 | 0.65 |
| 4b | 4',5'-2OH-CB38 | 4.04 | 0.56 | 19a | 2',3'-2OH-CB81 | 2.74 | 0.64 |
| 5a | 4',5'-2OH-CB55 | 3.82 | 0.58 | 20 | 5,6-2OH-CB105 | 0.98 | 0.39 |
| 6a | 4,5-2OH-CB56 | 4.21 | 0.58 | 21b | 4',5'-2OH-CB106 | 2.95 | 0.57 |
| 7a | 2',3'-2OH-CB60 | 3.75 | 0.66 | 22a | 2',3'-2OH-CB114 | 2.79 | 0.64 |
| 8a | 5,6-2OH-CB60 | 1.25 | 0.55 | 23a | 5',6'-2OH-CB126 | 1.13 | 0.63 |
| 9b | 4',5'-2OH-CB61 | 3.96 | 0.60 | 24 | 5,6-2OH-CB128 | 0.44 | 0.37 |
| 10a | 3,4-2OH-CB61 | 3.37 | 0.60 | 25 | 4',5'-2OH-CB129 | 2.57 | 0.38 |
| 11 | 5,6-2OH-CB66 | 2.32 | 0.39 | 26 | 5',6'-2OH-CB137 | 0.56 | 0.35 |
| 12a | 5',6'-2OH-CB66 | 2.24 | 0.63 | 27 | 5,6-2OH-CB138 | 0.90 | 0.38 |
| 13b | 4',5'-2OH-CB67 | 5.18 | 0.57 | 28 | 3,4-2OH-CB141 | 1.00 | 0.39 |
| 14a | 3,4-2OH-CB68 | 2.64 | 0.62 | 29 | 5,6-2OH-CB157 | 0.81 | 0.30 |
| 15a | 2',3'-2OH-CB74 | 3.14 | 0.64 | 30 | 2',3'-2OH-CB167 | 0.24 | 0.45 |
Table 1 Total score(Kd) of two hydroxy polychlorinated biphenyls
| No. | Compound | Kd | Similarity | No. | Compound | Kd | Similarity |
|---|---|---|---|---|---|---|---|
| 1a | 4,5-2OH-CB35 | 4.32 | 0.58 | 16a | 3,4-2OH-CB76 | 3.53 | 0.63 |
| 2,3-2OH-CB37 | 5.56 | 0.65 | 17a | 4,5-2OH-CB76 | 3.41 | 0.57 | |
| 3a | 5,6-2OH-CB37 | 2.03 | 0.57 | 18b | 2',3'-2OH-CB77 | 4.21 | 0.65 |
| 4b | 4',5'-2OH-CB38 | 4.04 | 0.56 | 19a | 2',3'-2OH-CB81 | 2.74 | 0.64 |
| 5a | 4',5'-2OH-CB55 | 3.82 | 0.58 | 20 | 5,6-2OH-CB105 | 0.98 | 0.39 |
| 6a | 4,5-2OH-CB56 | 4.21 | 0.58 | 21b | 4',5'-2OH-CB106 | 2.95 | 0.57 |
| 7a | 2',3'-2OH-CB60 | 3.75 | 0.66 | 22a | 2',3'-2OH-CB114 | 2.79 | 0.64 |
| 8a | 5,6-2OH-CB60 | 1.25 | 0.55 | 23a | 5',6'-2OH-CB126 | 1.13 | 0.63 |
| 9b | 4',5'-2OH-CB61 | 3.96 | 0.60 | 24 | 5,6-2OH-CB128 | 0.44 | 0.37 |
| 10a | 3,4-2OH-CB61 | 3.37 | 0.60 | 25 | 4',5'-2OH-CB129 | 2.57 | 0.38 |
| 11 | 5,6-2OH-CB66 | 2.32 | 0.39 | 26 | 5',6'-2OH-CB137 | 0.56 | 0.35 |
| 12a | 5',6'-2OH-CB66 | 2.24 | 0.63 | 27 | 5,6-2OH-CB138 | 0.90 | 0.38 |
| 13b | 4',5'-2OH-CB67 | 5.18 | 0.57 | 28 | 3,4-2OH-CB141 | 1.00 | 0.39 |
| 14a | 3,4-2OH-CB68 | 2.64 | 0.62 | 29 | 5,6-2OH-CB157 | 0.81 | 0.30 |
| 15a | 2',3'-2OH-CB74 | 3.14 | 0.64 | 30 | 2',3'-2OH-CB167 | 0.24 | 0.45 |
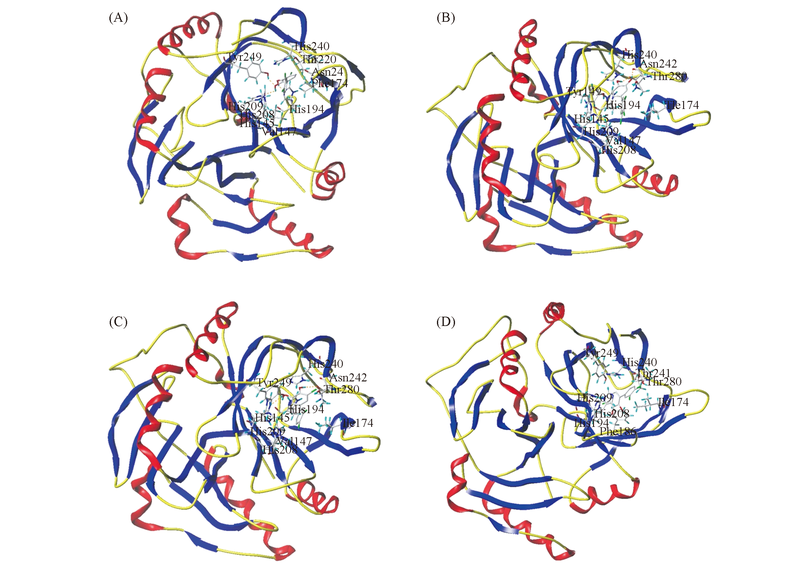
Fig.1 Binding pattern diagram of 2,3-2OH-CB37(A), 3,4-2OH-CB61(B), 4,5-2OH-CB35(C) and 5,6-2OH-CB37(D) with BphcThe three-dimensional structure of IKW6 represented by the ribbon structure, docking ligands and amino acid residues represented by the stick model, hydrogen bond represented by yellow dashed lines.
| No. | Internal validation | External validation | ||||||
|---|---|---|---|---|---|---|---|---|
| n | q2 | r2 | SEE | F | SEP | |||
| 1 | 5 | 0.569 | 0.985 | 0.134 | 108.792 | 0.982 | 0.135 | 0.779 |
| 2 | 7 | 0.566 | 0.985 | 0.136 | 133.500 | 0.935 | 0.273 | 0.824 |
| 3 | 5 | 0.553 | 0.983 | 0.137 | 102.504 | 0.945 | 0.327 | 0.745 |
| 4 | 6 | 0.535 | 0.973 | 0.150 | 133.385 | 0.931 | 0.328 | 0.801 |
| 5 | 6 | 0.574 | 0.984 | 0.128 | 101.032 | 0.977 | 0.172 | 0.771 |
| 6 | 6 | 0.501 | 0.994 | 0.132 | 104.524 | 0.965 | 0.154 | 0.682 |
| 7 | 7 | 0.598 | 0.980 | 0.126 | 112.334 | 0.982 | 0.163 | 0.679 |
| 8 | 6 | 0.624 | 0.962 | 0.142 | 124.231 | 0.973 | 0.182 | 0.724 |
| 9 | 5 | 0.572 | 0.986 | 0.156 | 141.529 | 0.932 | 0.209 | 0.754 |
| 10 | 7 | 0.529 | 0.991 | 0.134 | 105.010 | 0.967 | 0.311 | 0.593 |
| 11 | 6 | 0.535 | 0.990 | 0.145 | 131.204 | 0.985 | 0.345 | 0.628 |
| 12 | 7 | 0.567 | 0.968 | 0.127 | 112.574 | 0.971 | 0.178 | 0.721 |
Table 2 Primary parameters of the internal and external validation for CoMSIA model
| No. | Internal validation | External validation | ||||||
|---|---|---|---|---|---|---|---|---|
| n | q2 | r2 | SEE | F | SEP | |||
| 1 | 5 | 0.569 | 0.985 | 0.134 | 108.792 | 0.982 | 0.135 | 0.779 |
| 2 | 7 | 0.566 | 0.985 | 0.136 | 133.500 | 0.935 | 0.273 | 0.824 |
| 3 | 5 | 0.553 | 0.983 | 0.137 | 102.504 | 0.945 | 0.327 | 0.745 |
| 4 | 6 | 0.535 | 0.973 | 0.150 | 133.385 | 0.931 | 0.328 | 0.801 |
| 5 | 6 | 0.574 | 0.984 | 0.128 | 101.032 | 0.977 | 0.172 | 0.771 |
| 6 | 6 | 0.501 | 0.994 | 0.132 | 104.524 | 0.965 | 0.154 | 0.682 |
| 7 | 7 | 0.598 | 0.980 | 0.126 | 112.334 | 0.982 | 0.163 | 0.679 |
| 8 | 6 | 0.624 | 0.962 | 0.142 | 124.231 | 0.973 | 0.182 | 0.724 |
| 9 | 5 | 0.572 | 0.986 | 0.156 | 141.529 | 0.932 | 0.209 | 0.754 |
| 10 | 7 | 0.529 | 0.991 | 0.134 | 105.010 | 0.967 | 0.311 | 0.593 |
| 11 | 6 | 0.535 | 0.990 | 0.145 | 131.204 | 0.985 | 0.345 | 0.628 |
| 12 | 7 | 0.567 | 0.968 | 0.127 | 112.574 | 0.971 | 0.178 | 0.721 |
| Model | Fragment distinction | Fragment size | r2 | q2 | SEE | n | HL | |
|---|---|---|---|---|---|---|---|---|
| 1 | A/B/C | 4—7 | 0.976 | 0.867 | 0.912 | 0.243 | 6 | 59 |
| 2 | A/B/C/H | 4—7 | 0.987 | 0.926 | 0.924 | 0.172 | 6 | 59 |
| 3 | A/B/C/Ch | 4—7 | 0.976 | 0.867 | 0.912 | 0.241 | 6 | 59 |
| 4 | A/B/C/DA | 4—7 | 0.923 | 0.782 | 0.896 | 0.386 | 4 | 71 |
| 5 | A/B/C/H/Ch | 4—7 | 0.987 | 0.926 | 0.924 | 0.173 | 6 | 59 |
| 6 | A/B/C/H/DA | 4—7 | 0.989 | 0.900 | 0.935 | 0.164 | 6 | 59 |
| 7 | A/B/C/Ch/DA | 4—7 | 0.958 | 0.806 | 0.900 | 0.301 | 6 | 59 |
| 8 | A/B/C/H/Ch/DA | 4—7 | 0.980 | 0.852 | 0.913 | 0.204 | 6 | 59 |
Table 3 HQSAR model for distinguishing parameters of different fragments
| Model | Fragment distinction | Fragment size | r2 | q2 | SEE | n | HL | |
|---|---|---|---|---|---|---|---|---|
| 1 | A/B/C | 4—7 | 0.976 | 0.867 | 0.912 | 0.243 | 6 | 59 |
| 2 | A/B/C/H | 4—7 | 0.987 | 0.926 | 0.924 | 0.172 | 6 | 59 |
| 3 | A/B/C/Ch | 4—7 | 0.976 | 0.867 | 0.912 | 0.241 | 6 | 59 |
| 4 | A/B/C/DA | 4—7 | 0.923 | 0.782 | 0.896 | 0.386 | 4 | 71 |
| 5 | A/B/C/H/Ch | 4—7 | 0.987 | 0.926 | 0.924 | 0.173 | 6 | 59 |
| 6 | A/B/C/H/DA | 4—7 | 0.989 | 0.900 | 0.935 | 0.164 | 6 | 59 |
| 7 | A/B/C/Ch/DA | 4—7 | 0.958 | 0.806 | 0.900 | 0.301 | 6 | 59 |
| 8 | A/B/C/H/Ch/DA | 4—7 | 0.980 | 0.852 | 0.913 | 0.204 | 6 | 59 |
| No. | Compound | Frequency/cm-1 | Kd | Similarity | Change rate of Kd(%) | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Target molecule | 36.46 | 1.25 | 0.55 | |||||||
| 1 | 38.09 | 2.42 | 0.61 | 94 | ||||||
| 2 | 38.49 | 2.30 | 0.54 | 84 | ||||||
| 3 | 36.68 | 2.06 | 0.51 | 65 | ||||||
| 4 | 0.92 | 0.05 | ||||||||
| 5 | 30.09 | 2.40 | 0.55 | 92 | ||||||
| No. | Compound | Frequency/cm-1 | Kd | Similarity | Change rate ofKd(%) | |||||
| 6 | 24.94 | 2.32 | 0.53 | 86 | ||||||
| 7 | 0.27 | 0.55 | ||||||||
| 8 | 34.55 | 2.13 | 0.55 | 70 | ||||||
| 9 | 31.54 | 2.47 | 0.60 | 98 | ||||||
| 10 | 30.38 | 3.56 | 0.58 | 185 | ||||||
Table 4 Frequency and total score of new 5,6-2OH-CB60 molecules
| No. | Compound | Frequency/cm-1 | Kd | Similarity | Change rate of Kd(%) | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Target molecule | 36.46 | 1.25 | 0.55 | |||||||
| 1 | 38.09 | 2.42 | 0.61 | 94 | ||||||
| 2 | 38.49 | 2.30 | 0.54 | 84 | ||||||
| 3 | 36.68 | 2.06 | 0.51 | 65 | ||||||
| 4 | 0.92 | 0.05 | ||||||||
| 5 | 30.09 | 2.40 | 0.55 | 92 | ||||||
| No. | Compound | Frequency/cm-1 | Kd | Similarity | Change rate ofKd(%) | |||||
| 6 | 24.94 | 2.32 | 0.53 | 86 | ||||||
| 7 | 0.27 | 0.55 | ||||||||
| 8 | 34.55 | 2.13 | 0.55 | 70 | ||||||
| 9 | 31.54 | 2.47 | 0.60 | 98 | ||||||
| 10 | 30.38 | 3.56 | 0.58 | 185 | ||||||
| No. | Compound | IC50 | Change rate of IC50(%) | BCF | Change rate of BCF(%) | KOA | Change rate of KOA(%) | t1/2 | Change rate of t1/2(%) | ΔE/ (kJ·mol-1) | ΔH/ (kJ·mol-1) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Target molecule | 5,6-2OH-CB60 | 0.52 | 5.73 | 9.23 | 0.84 | ||||||
| 1 | 2-OH-5,6-2OH-CB60 | 0.17 | -67 | 4.18 | -27 | 9.19 | -0.43 | 0.93 | 10.7 | 302.86 | -50.09 |
| 2 | 2-NH2-5,6-2OH-CB60 | 0.33 | -37 | 4.42 | -23 | 9.13 | -1.0 | 0.83 | 1.1 | 738.96 | -124.15 |
| 3 | 2-OCH3-5,6-2OH-CB60 | 0.47 | -10 | 3.85 | -33 | 9.13 | -1.0 | 0.91 | 8.3 | 1178.28 | -87.43 |
| 4 | 2-CN-5,6-2OH-CB60 | ||||||||||
| 5 | 2-NHCH3-5,6-2OH-CB60 | 0.81 | 56 | 4.96 | -13 | 9.22 | -0.11 | 0.87 | 3.5 | 728.56 | -137.60 |
| 6 | 2-CH2CH3-5,6-2OH-CB60 | 0.41 | -21 | 4.66 | -19 | 9.59 | 3.9 | 0.88 | 4.7 | 1159.13 | -168.75 |
| 7 | 2-CH3-5,6-2OH-CB60 | ||||||||||
| 8 | 2-NO-5,6-2OH-CB60 | 0.65 | 25 | 5.49 | 4 | 9.37 | 1,5 | 0.72 | -14.2 | 743.19 | -297.07 |
| 9 | 2-CHO-5,6-2OH-CB60 | 0.089 | -83 | 4.78 | -17 | 9.10 | 1.4 | 0.89 | 5.9 | 967.69 | -98.11 |
| 10 | 2-COOH-5,6-2OH-CB60 | 0.13 | -75 | 4.35 | -24 | 9.32 | 0.97 | 0.84 | 0 | 854.45 | -178.76 |
Table 5 IC50, BCF, KOA, t1/2, ΔE and ΔH of new designed 5,6-2OH-CB60 molecules*
| No. | Compound | IC50 | Change rate of IC50(%) | BCF | Change rate of BCF(%) | KOA | Change rate of KOA(%) | t1/2 | Change rate of t1/2(%) | ΔE/ (kJ·mol-1) | ΔH/ (kJ·mol-1) |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Target molecule | 5,6-2OH-CB60 | 0.52 | 5.73 | 9.23 | 0.84 | ||||||
| 1 | 2-OH-5,6-2OH-CB60 | 0.17 | -67 | 4.18 | -27 | 9.19 | -0.43 | 0.93 | 10.7 | 302.86 | -50.09 |
| 2 | 2-NH2-5,6-2OH-CB60 | 0.33 | -37 | 4.42 | -23 | 9.13 | -1.0 | 0.83 | 1.1 | 738.96 | -124.15 |
| 3 | 2-OCH3-5,6-2OH-CB60 | 0.47 | -10 | 3.85 | -33 | 9.13 | -1.0 | 0.91 | 8.3 | 1178.28 | -87.43 |
| 4 | 2-CN-5,6-2OH-CB60 | ||||||||||
| 5 | 2-NHCH3-5,6-2OH-CB60 | 0.81 | 56 | 4.96 | -13 | 9.22 | -0.11 | 0.87 | 3.5 | 728.56 | -137.60 |
| 6 | 2-CH2CH3-5,6-2OH-CB60 | 0.41 | -21 | 4.66 | -19 | 9.59 | 3.9 | 0.88 | 4.7 | 1159.13 | -168.75 |
| 7 | 2-CH3-5,6-2OH-CB60 | ||||||||||
| 8 | 2-NO-5,6-2OH-CB60 | 0.65 | 25 | 5.49 | 4 | 9.37 | 1,5 | 0.72 | -14.2 | 743.19 | -297.07 |
| 9 | 2-CHO-5,6-2OH-CB60 | 0.089 | -83 | 4.78 | -17 | 9.10 | 1.4 | 0.89 | 5.9 | 967.69 | -98.11 |
| 10 | 2-COOH-5,6-2OH-CB60 | 0.13 | -75 | 4.35 | -24 | 9.32 | 0.97 | 0.84 | 0 | 854.45 | -178.76 |
| [1] | Yang F. F., Xu Y., Prog. Chem., 2005, 4, 740—748 |
| (杨方星, 徐盈. 化学进展, 2005, 4, 740—748) | |
| [2] | Flor S., He X., Lehmler H. J., Ludewig G., Environ. Sci. Pollut. R., 2016, 23, 2186—2200 |
| [3] | Ptak A., Ludewig G., Rak A., Nadolna W., Bochenek M., Environment International, 2010, 36, 935—941 |
| [4] | Schultz T. W., Seward J. R., Sinks G. D., Environ. Toxicol. Chem., 2010, 19, 301—304 |
| [5] | Letcher R.J., Klasson-Wehler E., Bergman A., Methyl Sulfone and Hydroxylated Metabolites of Polychlorinated Biphenyls, Springer,Berlin, 2000 |
| [6] | Sun X., Zhang C., Zhao Y., Bai J., Zhang Q., Environmental Science & Technology, 2012, 46, 8148—8155 |
| [7] | Moore T., Pagni R. M., J. Org. Chem., 1987, 52, 770—773 |
| [8] | Goto M., Sugiura K., Hattori M., Miyagawa T., Okamura M., Chemosphere,1974, 3, 233—238 |
| [9] | Bergman A., Bamford I., Wachtmeister C. A., J. Labelled Compd. Rad., 2010, 18, 1023—1032 |
| [10] | Dasary S. S., Saloni J., Fletcher A., Anjaneyulu Y., Yu H., Inter. J. Environ. Res. Public Health,2010, 7, 3987—4001 |
| [11] | Kong J., Achari G., Langford C. H., J. Environ. Sci. Health Part A, Toxic/hazardous Substances Environ. Engin., 2013, 48, 92—98 |
| [12] | Wang X., Sun Y. M., Tech. Dev. Chem. Ind., 2010, 39, 21—24 |
| (王贤, 孙友敏. 化工技术与开发, 2010, 39, 21—24) | |
| [13] | Field J. A., Sierra-Alvarez R., Environ. Pollut., 2008, 155, 1—12 |
| [14] | Fukuda M., Yasukochi Y., Kikuchi Y., Biochem. Bioph. Res. Co., 1994, 202, 850—856 |
| [15] | Francova K., Macková M., Macek T., Environ. Pollut., 2004, 127, 41—48 |
| [16] | Sylvestre M., Sirois M., Hurtubise Y., Gene,1996, 174, 195—202 |
| [17] | Da C. E., Sippl W., De C. R. T., Antunes O. A. C., Alencastro R. B., Eur. J. Med. Chem., 2009, 44, 4344—4352 |
| [18] | Hu S. Q., Mi S. Q., Jia X. L., Chem. J. Chinese Universities, 2011, 32(10), 2402—2409 |
| (胡松青, 米思奇, 贾晓林. 高等学校化学学报, 2011,32(10), 2402—2409) | |
| [19] | Uiaran D. O. M., Girão A. M., Araújo D. B. M., Lamim B. M., Mendes C. L., Drug Des. Dev. Ther., 2013, 7, 953—962 |
| [20] | Nair P. C., Srikanth K., Sobhia M. E., Bioorg. Med. Chem. Lett., 2008, 18,1323—1330 |
| [21] | Ye M., Dawson M. I., Bioorg. Med. Chem. Lett., 2009, 19, 3310—3315 |
| [22] | Cheng Y., Zhou M., Tung C. H., Ji M., Zhang F., Bioorg. Med. Chem. Lett., 2010, 20, 3329—3337 |
| [23] | Mckinney J. D., Gottschalk K. E., Pedersen L., J. Mol. Struc-Theochem., 1983, 104, 445—450 |
| [24] | Dean J. R., Ma R., Chemosphere,2007, 68, 1399—1407 |
| [25] | Molina L., Cabes M., Díaz-Ferrero J., Chemosphere,2000, 40, 921—927 |
| [26] | Xin M. L., Yang J. W., Li Y., Chem. Cent. J., 2017, 11, 61—75 |
| [27] | Frisch M.J., Trucks G. W., Schlegel H. B., Scuseria G. E., Robb M. A., Cheeseman J. R., Scalmani G., Barone V., Mennucci B., Petersson G. A., Nakatsuji H., Caricato M., Li X., Hratchian H. P., Izmaylov A. F., Bloino J., Zheng G., Sonnenberg J. L., Hada M., Ehara M., Toyota K., Fukuda R., Hasegawa J., Ishida M., Nakajima T., Honda Y., Kitao O., Nakai H., Vreven T., Montgomery J. A., Peralta J. J., Ogliaro F., Bearpark M., Heyd J. J., Brothers E., Kudin K. N., Staroverov V. N., Kobayashi R., Normand J., Raghavachari K., Rendell A., Burant J.C., Iyengar S. S., Tomasi J., Cossi M., Rega N., Millam J. M., Klene M., Knox J. E., Cross J. B., Bakken V., Adamo C., Jaramillo J., Gomperts R., Stratmann R. E., Yazyev O., Austin A. J., Cammi R., Pomelli C., Ochterski J. W., Martin R. L., Morokuma K., Zakrzewski V. G., Voth G. A., Salvador P., Dannenberg J. J., Dapprich S., Daniels A. D., Farkas Ö., Foresman J. B., Ortiz J. V., Cioslowski J. F. D., Gaussian 09, Revision E.01, Gaussian Inc., Wallingford CT, 2009 |
| [28] | Singh S. P., Gogoi B., Konwar B. K., Ramteke A., J. Pharm. Pharmcol., 2013, 7, 443—447 |
| [29] | Chen Y., Li Y., Chemistry Letters, 2016, 45, 1453—1456 |
| [30] | Yang W.H., Three Dimensional Structure Activity Relationship of Endocrine Disrupting Activity of New Persistent Organic Pollutants, China University of Mining and Technology Press, Xuzhou, 2011 |
| (杨伟华. 新型持久性有机污染物内分泌干扰活性的三维构效关系研究, 徐州: 中国矿业大学出版社, 2011) | |
| [31] | Golbraikh A., Tropsha A., Mol. Graph. Modell., 2002, 20, 269—276 |
| [32] | Tong L., Guo L., Lv X., Li Y., J. Mole. Graph. Model., 2017, 71, 1—12 |
| [33] | Li X., Ye L., Wang X., Shi W., Liu H., Chemosphere,2013, 92, 795—802 |
| [34] | Liu H., Liu H., Sun P., Wang Z., Chemosphere,2014, 114, 101—105 |
| [35] | Chen Y., Cai X.Y., Jiang L., Li Y., Ecotoxicology and Environmental Safety, 2016, 124, 202—212 |
| [36] | Xu Z., Chen Y., Qiu Y., Gu W. W., Li Y., Chem. Res. Chinese Universities, 2016, 32(3), 348—356 |
| [1] | LEI Xiaotong, JIN Yiqing, MENG Xuanyu. Prediction of the Binding Site of PIP2 in the TREK-1 Channel Based on Molecular Modeling [J]. Chem. J. Chinese Universities, 2021, 42(8): 2550. |
| [2] | SHUAI Die, ZHAO Meijuan, CHEN Bingnian, WANG Li. Inhibitory Effect of Four Kinds of Keegin-type Phosphomolybdate on Tyrosinase and Melanin Formation and Its Antioxidant Activities [J]. Chem. J. Chinese Universities, 2021, 42(12): 3579. |
| [3] | YANG Ju, SU Lijiao, LI Canhua, LU Jiajia, YANG Junli, GU Jie, YANG Li, YANG Lijuan. Host-guest Complexation Behavior of Nardosinone and Water-soluble Phosphate Salt Pillar[6]arene [J]. Chem. J. Chinese Universities, 2021, 42(10): 3099. |
| [4] | ZHANG Aiqin, WANG Man, SHEN Gangyi, JIN Jun. Interactions Between Polybrominated Diphenyl Ethers and Human Serum Albumin Using SPR and Molecular Docking [J]. Chem. J. Chinese Universities, 2020, 41(9): 2054. |
| [5] | WANG Lianping,LI Qingjie,LIU Xiaoyan,REN Yueying,YANG Xiuwei. Screening of Cholinesterase Inhibitors in Fructus Evodiae Alkaloids Based on UFLC-MS/molecular Simulation † [J]. Chem. J. Chinese Universities, 2020, 41(1): 111. |
| [6] | WANG Xiaoxia, MA Litong, NIE Zhihua, WANG Zhengde, CUI Jinlong, ZHAO Wenyuan, SAI Huazheng. Interaction Between Fulvic Acid and Pepsin Investigated by Multispectral and Molecular Docking Simulation † [J]. Chem. J. Chinese Universities, 2019, 40(9): 1840. |
| [7] | LIU Zhongcheng, LIU Shifang, ZHANG Su, YANG Yanlei, LI Fei, ZHANG Nan, YUAN Xin, ZHANG Yanfen. Structure Prediction and Screening of Oligonucleotide Aptamers Target Cε3-Cε4 Protein† [J]. Chem. J. Chinese Universities, 2019, 40(1): 83. |
| [8] | WANG Yan, CHEN Ping, WANG Yunfei, LIU Guiying, YANG Xi, SU Ying, LI Junyang, LIU Weiwei, LIN Lie. Spectral Characterization of the Interaction Between Methamphetamine and Serum Albumin† [J]. Chem. J. Chinese Universities, 2018, 39(11): 2507. |
| [9] | DUAN Yongbin, YIN Yan, MENG Fanli, ZHAO Lianhua, LIU Yukun, YUAN Zhe, FENG Yangbo. Design, Synthesis and Biological Evaluation of Benzothiazoles as Highly Potent ROCK Inhibitors Through Molecular Docking and Free Energy Calculations† [J]. Chem. J. Chinese Universities, 2017, 38(9): 1568. |
| [10] | WANG Song, GUAN Shanshan, WAN Yongfeng, SHAN Yaming, ZHANG Hao. Molecular Dynamics Simulation Study on the Binding Modes of Angiotensin-converting Enzyme with Inhibitory Peptides† [J]. Chem. J. Chinese Universities, 2017, 38(7): 1216. |
| [11] | TANG Qian, SU Jinhong, CAO Hongyu, WANG Lihao, SHI Fei, WANG Ailing, GONG Tingting, JIN Xiaojun, ZHENG Xuefang. Interaction of Pyrimidine Derivatives with Human Serum Albumin† [J]. Chem. J. Chinese Universities, 2017, 38(11): 1982. |
| [12] | TANG Guanghui, ZHANG Ya, ZHANG Yuping, ZHOU Pengpeng, LIN Zhihua, WANG Yuanqiang. Molecular Docking, QSAR and Molecular Dynamics Simulation on Phosphorus Containing Pyrimidines as CDK9 Inhibitors† [J]. Chem. J. Chinese Universities, 2017, 38(11): 2061. |
| [13] | ZHAO Bin, HUO Jingqian, XING Jihong, QI Meng, ZHANG Jinlin, DONG Jingao. Homologous Modeling of Transketolase AtTKL1 and Its Combination with α-Terthienyl in Arabidopsis Thaliana† [J]. Chem. J. Chinese Universities, 2015, 36(4): 682. |
| [14] | DONG Lu, YI Zhongsheng, WU Zhiwei, WANG Haiyang, ZHANG Aiqian. Mechanism Study on the Interaction Between 2'-Hydroxy-2,4-dibromo Diphenyl Ethers and Human Serum Albumin Based on Spectroscopic Methods and Computional Simulations† [J]. Chem. J. Chinese Universities, 2015, 36(3): 516. |
| [15] | ZHANG Min, ZHANG Yi, LI Chengtao, QIN Jiaxiang, Gao Rang, MA Xiaoning, QIU Jianhui. N435 Enzymatic Degradation Difference and Molecular Modeling of Hydrophilic Modified PBS-based Copolymer† [J]. Chem. J. Chinese Universities, 2015, 36(3): 568. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||