

Chem. J. Chinese Universities ›› 2019, Vol. 40 ›› Issue (1): 83.doi: 10.7503/cjcu20180428
• Organic Chemistry • Previous Articles Next Articles
LIU Zhongcheng1,2, LIU Shifang1,2, ZHANG Su1,2, YANG Yanlei1,2, LI Fei1,2, ZHANG Nan1,2, YUAN Xin1,2, ZHANG Yanfen2,3,*
Received:2018-06-11
Online:2019-01-10
Published:2018-12-03
Contact:
ZHANG Yanfen
Supported by:CLC Number:
TrendMD:
LIU Zhongcheng, LIU Shifang, ZHANG Su, YANG Yanlei, LI Fei, ZHANG Nan, YUAN Xin, ZHANG Yanfen. Structure Prediction and Screening of Oligonucleotide Aptamers Target Cε3-Cε4 Protein†[J]. Chem. J. Chinese Universities, 2019, 40(1): 83.
| No. | Sequence(5'-3') | No. | Sequence(5'-3') |
|---|---|---|---|
| 1 | ACCGACCGT | 25 | TGTCTTGTCTTTAATGGTATCCACA |
| 2 | CCGACCGTGCTGG | 26 | TGTCTTTAATGGTATCCACA |
| 3 | CCGTGCTGG | 27 | CTTGTCTTTAATGGTATCCACAG |
| 4 | CGACCG | 28 | TTGTCTTTAA |
| 5 | ACCGT | 29 | TGTCTTTAATGGTATCCA |
| 6 | GACCGTGCTGGACTC | 30 | GTCTTTAATGGTATCCA |
| 7 | CGTGCTGGACTCTGCCCCG | 31 | TCTTTAATGGTATCCACAGTATGA |
| 8 | GTGCTGGAC | 32 | CTTTAATGGTATCCACAG |
| 9 | GCTGGACTCTGC | 33 | TTTAA |
| 10 | TGGACTCTGCCCCGCTGCCTTGCA | 34 | TAATGGTA |
| 11 | GGACTCTGCCCC | 35 | ATGGTAT |
| 12 | GGACTCTGCC | 36 | GGTATCC |
| 13 | GACTCTGCCCCGCTGCCTTGCATC | 37 | GTATCCAC |
| 14 | ACTCTGCCCCGCTGCCTTGCATCTGT | 38 | TATCCACAGTA |
| 15 | GCCCCGC | 39 | ATCCACAGTAT |
| 16 | GCCCCGCTGC | 40 | TCCACAGTATGA |
| 17 | GCTGCCTTGC | 41 | CACAGTATG |
| 18 | TGCCTTGCA | 42 | CAGTATG |
| 19 | CATCTG | 43 | GCGAGC |
| 20 | TTGCATCTGTCTTGTCTTTAA | 44 | GCGAGCGTTGC |
| 21 | CATCTGTCTTG | 45 | CGAGCG |
| 22 | ATCTGTCTTGTCTTTAAT | 46 | CGAGCGTTGCG |
| 23 | TGTCTTGTCTTTAATGGTATCCA | 47 | GCGTTGC |
| 24 | GTCTTGTCTTTAATGGTATCCACAG | 48 | CGTTGCG |
Table 1 Sequences for DNA aptamers
| No. | Sequence(5'-3') | No. | Sequence(5'-3') |
|---|---|---|---|
| 1 | ACCGACCGT | 25 | TGTCTTGTCTTTAATGGTATCCACA |
| 2 | CCGACCGTGCTGG | 26 | TGTCTTTAATGGTATCCACA |
| 3 | CCGTGCTGG | 27 | CTTGTCTTTAATGGTATCCACAG |
| 4 | CGACCG | 28 | TTGTCTTTAA |
| 5 | ACCGT | 29 | TGTCTTTAATGGTATCCA |
| 6 | GACCGTGCTGGACTC | 30 | GTCTTTAATGGTATCCA |
| 7 | CGTGCTGGACTCTGCCCCG | 31 | TCTTTAATGGTATCCACAGTATGA |
| 8 | GTGCTGGAC | 32 | CTTTAATGGTATCCACAG |
| 9 | GCTGGACTCTGC | 33 | TTTAA |
| 10 | TGGACTCTGCCCCGCTGCCTTGCA | 34 | TAATGGTA |
| 11 | GGACTCTGCCCC | 35 | ATGGTAT |
| 12 | GGACTCTGCC | 36 | GGTATCC |
| 13 | GACTCTGCCCCGCTGCCTTGCATC | 37 | GTATCCAC |
| 14 | ACTCTGCCCCGCTGCCTTGCATCTGT | 38 | TATCCACAGTA |
| 15 | GCCCCGC | 39 | ATCCACAGTAT |
| 16 | GCCCCGCTGC | 40 | TCCACAGTATGA |
| 17 | GCTGCCTTGC | 41 | CACAGTATG |
| 18 | TGCCTTGCA | 42 | CAGTATG |
| 19 | CATCTG | 43 | GCGAGC |
| 20 | TTGCATCTGTCTTGTCTTTAA | 44 | GCGAGCGTTGC |
| 21 | CATCTGTCTTG | 45 | CGAGCG |
| 22 | ATCTGTCTTGTCTTTAAT | 46 | CGAGCGTTGCG |
| 23 | TGTCTTGTCTTTAATGGTATCCA | 47 | GCGTTGC |
| 24 | GTCTTGTCTTTAATGGTATCCACAG | 48 | CGTTGCG |
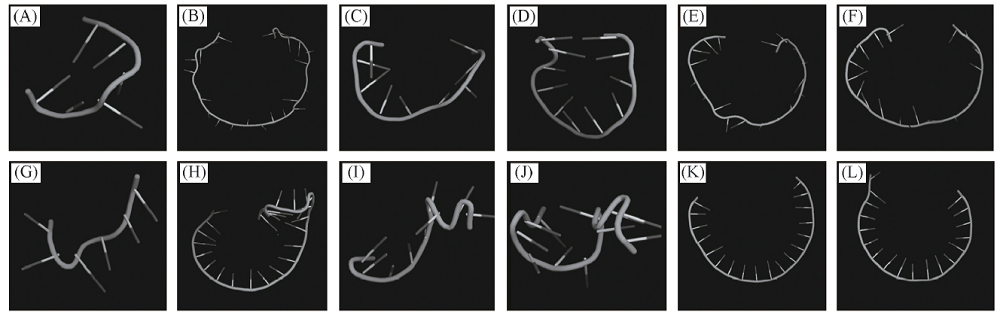
Fig.2 3D structures prediction of DNA short hairpin(A)—(F) 3D structures of sequences 4, 10, 18, 21, 22, 29 by MMB; (G)—(L) 3D structures of sequences 4, 10, 18, 21, 22, 29 by RNA composer.
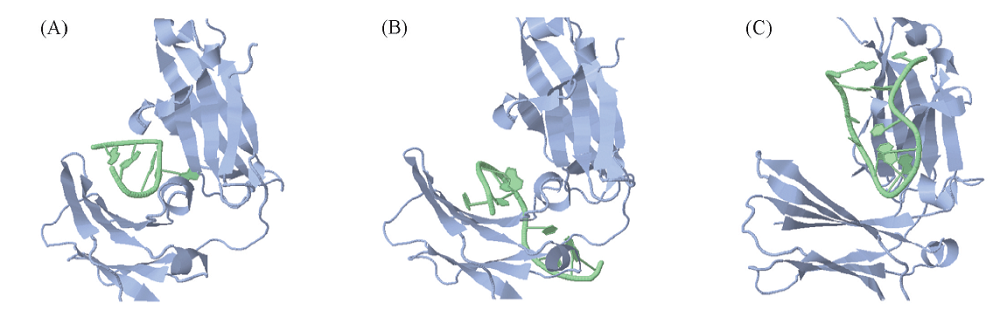
Fig.3 3D conformation of DNA short hairpins and Cε3-Cε4 protein interaction predicted by NPDock(A)—(C) 3D visualization of the 4, 18, 21 best scored complex.
| No. | Sequence | Score | No. | Sequence | Score |
|---|---|---|---|---|---|
| 6 | GACCGTGCTGGACTC | 4.08 | 22 | ATCTGTCTTGTCTTTAAT | 4.17 |
| 11 | GGACTCTGCCCC | 3.87 | 23 | TGTCTTGTCTTTAATGGTATCCA | 3.72 |
| 12 | GGACTCTGCC | 4.85 | 25 | TGTCTTGTCTTTAATGGTATCCACA | 3.97 |
| 20 | TTGCATCTGTCTTGTCTTTAA | 3.72 | 31 | TCTTTAATGGTATCCACAGTATGA | 3.82 |
Table 2 Best sequences of DNA aptamers from docking results
| No. | Sequence | Score | No. | Sequence | Score |
|---|---|---|---|---|---|
| 6 | GACCGTGCTGGACTC | 4.08 | 22 | ATCTGTCTTGTCTTTAAT | 4.17 |
| 11 | GGACTCTGCCCC | 3.87 | 23 | TGTCTTGTCTTTAATGGTATCCA | 3.72 |
| 12 | GGACTCTGCC | 4.85 | 25 | TGTCTTGTCTTTAATGGTATCCACA | 3.97 |
| 20 | TTGCATCTGTCTTGTCTTTAA | 3.72 | 31 | TCTTTAATGGTATCCACAGTATGA | 3.82 |
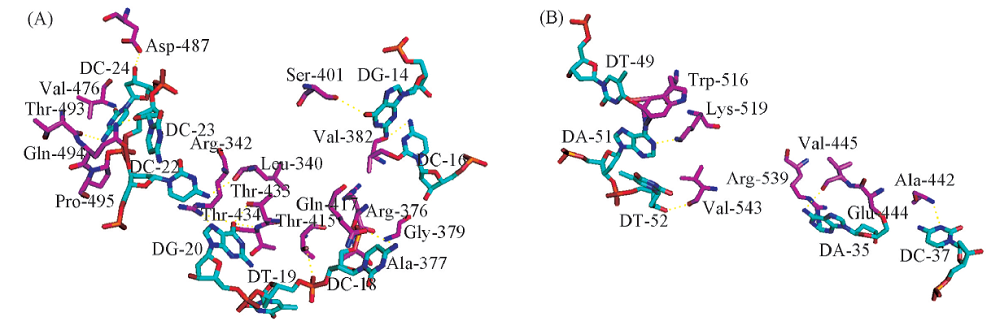
Fig.6 Predictions for protein-DNA complexes(A) The site of interaction of sequence 11 protein complex, the purple are the amino acid residues Thr, Ser, Asp, Lys and His. The cyan are the base T, C and G. (B) The site of interaction of sequence 22-protein complex, the purple are the amino acid residues Ala, Arg, Val, Lys, Glu and Arg. The cyan are the base A, T, C and G.
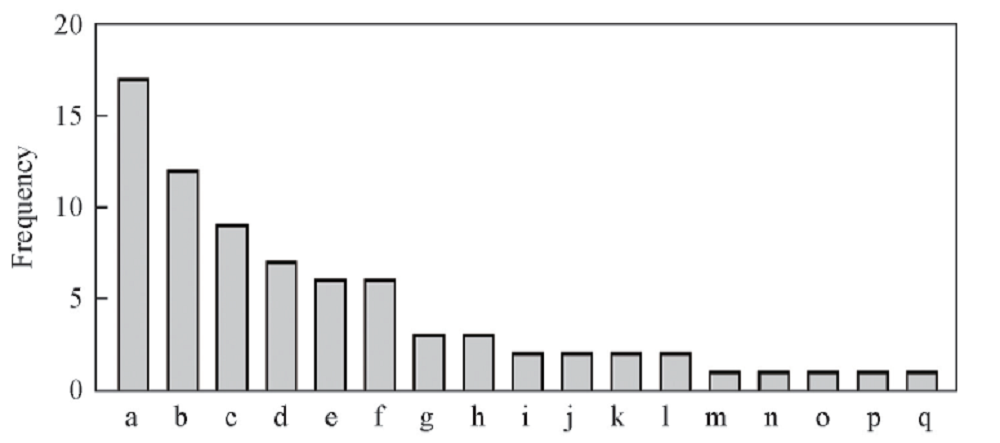
Fig.7 Frequency of amino acid usage at the site of complex interactiona. Thr; b.Arg; c. Ser; d. Gln; e. Lys; f. Val; g. Asp; h. Pro; i. Glu; j. Phe; k. Ala; l. Asn; m. His; n. Ile; o. Leu; p. Met; q. Tyr.
| [1] | Burch B. D., Garrido C., Margolis D. M., Virus Res., 2017, 228, 141—146 |
| [2] | Rozenblum G. T., Lopez V. G., Vitullo A. D., Radrizzani M., Expert Opin. Drug Discov., 2016, 11(2), 127—135 |
| [3] | Zheng K., Hong Y., Zhao X. F., Chem. Res. Chinese Universities,2014, 30(4), 644—649 |
| [4] | Trott O., Olson A. J., J. Comput. Chem.,2010, 31(2), 455—461 |
| [5] | De Vries S. J., van Dijk M., Bonvin A. M., Nat.Protoc., 2010, 5(5), 883—897 |
| [6] | Schneidman-Duhovny D., Inbar Y., Nussinov R., Wolfson H. J., Nucleic Acids Res., 2005, 33(Web Server issue), W363—W367 |
| [7] | Pierce B. G., Wiehe K., Hwang H., Kim B. H., Vreven T., Weng Z., Bioinformatics,2014, 30(12), 1771—1773 |
| [8] | Ritchie D. W., Kemp G. J., Proteins,2000, 39(2), 178—194 |
| [9] | Wei H. E., Zhao A., Zou J. J., Chem. Res. Chinese Universities,2018, 34(1), 90—94 |
| [10] | Abolhassani N., Leon J., Sheng Z., Mech. Ageing Dev.,2017, 161(Pt A), 95—104 |
| [11] | Tuszynska I., Magnus M., Jonak K., Dawson W., Bujnicki J. M., Nucleic Acids Res., 2015, 43(W1), W425—430 |
| [12] | Tuszynska I., Bujnicki J. M., BMC Bioinformatics,2011, 12(1), 348—363 |
| [13] | Liu Z. C., Zhao L. J., Zhang Y. F., Shi H. L., Xie Y., Acta Pharmaceutica Sinica,2012, 47(12), 1605—1611 |
| (刘中成, 赵丽君, 张艳芬, 时海浪, 解瑶. 药学学报, 2012, 47(12), 1605—1611) | |
| [14] | Tuszynska I., Matelska D., Magnus M., Methods,2014, 65(3), 310—319 |
| [15] | Popenda M., Szachniuk M., Antczak M., Purzycka K. J., Lukasiak P., Bartol N., Blazewicz J., Adamiak R. W., Nucleic Acids Res., 2012, 40(14), e112 |
| [16] | Tek A., Korostelev A. A., Flores S. C., Nucleic Acids Res., 2016, 44(1), 95—105 |
| [17] | Heiat M., Najafi A., Ranjbar R., Latifi A. M., Rasaee M. J., J. Biotechnol., 2016, 230, 34—39 |
| [18] | Yang Q., Du W. W., Wu N., Yang W., Awan F. M., Fang L., Ma J., Li X., Zeng Y., Yang Z., Dong J., Khorshidi A., Yang B. B., Cell Death Differ., 2017, 24(9), 1609—1620 |
| [19] | Wang M. H., Li D. J., Guangzhou Chemical Industry,2015, 43(15), 52—54 |
| (王明华, 李杜娟. 广州化工, 2015, 43(15), 52—54) | |
| [20] | Wu J. S., Hu D., Yan S. C., J. Southeast Univ. (Natural Science Edition), 2011, 41(4), 778—783 |
| (吴建盛, 胡栋, 晏善成. 东南大学学报(自然科学版), 2011, 41(4), 778—783) | |
| [21] | Zhao L.J., Binding Sites Prediction and Identification of Oligonucleotide Aptamers Targeting Cε3-Cε4, Hebei University,Baoding, 2015 |
| (赵丽健. Cε3-Cε4核酸适配子结合位点预测及验证研究, 保定: 河北大学, 2015) |
| [1] | LEI Xiaotong, JIN Yiqing, MENG Xuanyu. Prediction of the Binding Site of PIP2 in the TREK-1 Channel Based on Molecular Modeling [J]. Chem. J. Chinese Universities, 2021, 42(8): 2550. |
| [2] | SHUAI Die, ZHAO Meijuan, CHEN Bingnian, WANG Li. Inhibitory Effect of Four Kinds of Keegin-type Phosphomolybdate on Tyrosinase and Melanin Formation and Its Antioxidant Activities [J]. Chem. J. Chinese Universities, 2021, 42(12): 3579. |
| [3] | YANG Ju, SU Lijiao, LI Canhua, LU Jiajia, YANG Junli, GU Jie, YANG Li, YANG Lijuan. Host-guest Complexation Behavior of Nardosinone and Water-soluble Phosphate Salt Pillar[6]arene [J]. Chem. J. Chinese Universities, 2021, 42(10): 3099. |
| [4] | ZHANG Aiqin, WANG Man, SHEN Gangyi, JIN Jun. Interactions Between Polybrominated Diphenyl Ethers and Human Serum Albumin Using SPR and Molecular Docking [J]. Chem. J. Chinese Universities, 2020, 41(9): 2054. |
| [5] | WANG Lianping,LI Qingjie,LIU Xiaoyan,REN Yueying,YANG Xiuwei. Screening of Cholinesterase Inhibitors in Fructus Evodiae Alkaloids Based on UFLC-MS/molecular Simulation † [J]. Chem. J. Chinese Universities, 2020, 41(1): 111. |
| [6] | WANG Xiaoxia, MA Litong, NIE Zhihua, WANG Zhengde, CUI Jinlong, ZHAO Wenyuan, SAI Huazheng. Interaction Between Fulvic Acid and Pepsin Investigated by Multispectral and Molecular Docking Simulation † [J]. Chem. J. Chinese Universities, 2019, 40(9): 1840. |
| [7] | XIN Meiling, CHU Zhenhua, LI Yu. Molecular Modification of Polychlorinated Biphenyl Dihydroxy Derivatives Through Molecular Docking Associated with CoMSIA/HQSAR Models† [J]. Chem. J. Chinese Universities, 2018, 39(2): 299. |
| [8] | WANG Yan, CHEN Ping, WANG Yunfei, LIU Guiying, YANG Xi, SU Ying, LI Junyang, LIU Weiwei, LIN Lie. Spectral Characterization of the Interaction Between Methamphetamine and Serum Albumin† [J]. Chem. J. Chinese Universities, 2018, 39(11): 2507. |
| [9] | DUAN Yongbin, YIN Yan, MENG Fanli, ZHAO Lianhua, LIU Yukun, YUAN Zhe, FENG Yangbo. Design, Synthesis and Biological Evaluation of Benzothiazoles as Highly Potent ROCK Inhibitors Through Molecular Docking and Free Energy Calculations† [J]. Chem. J. Chinese Universities, 2017, 38(9): 1568. |
| [10] | WANG Song, GUAN Shanshan, WAN Yongfeng, SHAN Yaming, ZHANG Hao. Molecular Dynamics Simulation Study on the Binding Modes of Angiotensin-converting Enzyme with Inhibitory Peptides† [J]. Chem. J. Chinese Universities, 2017, 38(7): 1216. |
| [11] | TANG Qian, SU Jinhong, CAO Hongyu, WANG Lihao, SHI Fei, WANG Ailing, GONG Tingting, JIN Xiaojun, ZHENG Xuefang. Interaction of Pyrimidine Derivatives with Human Serum Albumin† [J]. Chem. J. Chinese Universities, 2017, 38(11): 1982. |
| [12] | ZHAO Bin, HUO Jingqian, XING Jihong, QI Meng, ZHANG Jinlin, DONG Jingao. Homologous Modeling of Transketolase AtTKL1 and Its Combination with α-Terthienyl in Arabidopsis Thaliana† [J]. Chem. J. Chinese Universities, 2015, 36(4): 682. |
| [13] | DONG Lu, YI Zhongsheng, WU Zhiwei, WANG Haiyang, ZHANG Aiqian. Mechanism Study on the Interaction Between 2'-Hydroxy-2,4-dibromo Diphenyl Ethers and Human Serum Albumin Based on Spectroscopic Methods and Computional Simulations† [J]. Chem. J. Chinese Universities, 2015, 36(3): 516. |
| [14] | ZHANG Min, ZHANG Yi, LI Chengtao, QIN Jiaxiang, Gao Rang, MA Xiaoning, QIU Jianhui. N435 Enzymatic Degradation Difference and Molecular Modeling of Hydrophilic Modified PBS-based Copolymer† [J]. Chem. J. Chinese Universities, 2015, 36(3): 568. |
| [15] | CHEN Linfeng, ZHANG Jing, ZHU Yaxian, ZHANG Yong. Molecular Interactions of 1-Hydroxypyrene with Catalase Revealed by Spectroscopic Methods Combined with Molecular Docking† [J]. Chem. J. Chinese Universities, 2015, 36(12): 2394. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||