

高等学校化学学报 ›› 2020, Vol. 41 ›› Issue (6): 1243.doi: 10.7503/cjcu20200080
收稿日期:2020-02-17
出版日期:2020-06-10
发布日期:2020-03-17
通讯作者:
张志凌
E-mail:zlzhang@whu.edu.cn
基金资助:
ZHANG Zuoran,ZHANG Li,ZHANG Zhiling*( )
)
Received:2020-02-17
Online:2020-06-10
Published:2020-03-17
Contact:
Zhiling ZHANG
E-mail:zlzhang@whu.edu.cn
Supported by:摘要:
基于磁性纳米球在微流控芯片上的侧向磁泳, 利用微流控芯片分选了不同磁响应性的磁球. 提出了包含磁性纳米球聚集与偏移的理论模型, 用于分析磁球在芯片上的侧向位移. 在理论分析的基础上设计了芯片系统, 使不同磁响应性的磁纳米球可以在芯片系统上依次被分选. 实验结果表明, 2种磁性纳米球的分选效率均可接受, 且实验操作简单; 磁响应性强的磁球可被完全分离, 这对于珍贵分析样品的分选很有价值. 该分选系统被成功用于同时分选样品中乙型肝炎病毒的DNA与丙型肝炎病毒的反转录DNA, 在生化分析中具有广阔的应用前景.
中图分类号:
TrendMD:
张作然, 张利, 张志凌. 基于侧向磁泳的微流控芯片对两种磁性纳米球的同时分选. 高等学校化学学报, 2020, 41(6): 1243.
ZHANG Zuoran, ZHANG Li, ZHANG Zhiling. On-chip Sorting of Beads with Different Magnetic Responsiveness by Lateral Magnetophoresis †. Chem. J. Chinese Universities, 2020, 41(6): 1243.
| DNA | Sequence |
|---|---|
| Capture-HBV | Biotin-TEG-5'-TGGCTTTCAGTTATA-3' |
| Capture-HCV | Biotin-TEG-5'-TCGTGGATAAACCCG-3' |
| Target-HBV | 5'-ATACCACATCATCCATATAACTGAAAGCCA-3' |
| Target-HCV | 5'-ATCTCCAGGCATTGAGCGGGTTTATCCACGA-3' |
| Probe-HBV | 5'-TGGATGATGTGGTAT-3'-TEG-biotin |
| Probe-HCV | 5'-CTCAATGCCTGGAGAT-3'-TEG-biotin |
Table 1 Sequences of DNA
| DNA | Sequence |
|---|---|
| Capture-HBV | Biotin-TEG-5'-TGGCTTTCAGTTATA-3' |
| Capture-HCV | Biotin-TEG-5'-TCGTGGATAAACCCG-3' |
| Target-HBV | 5'-ATACCACATCATCCATATAACTGAAAGCCA-3' |
| Target-HCV | 5'-ATCTCCAGGCATTGAGCGGGTTTATCCACGA-3' |
| Probe-HBV | 5'-TGGATGATGTGGTAT-3'-TEG-biotin |
| Probe-HCV | 5'-CTCAATGCCTGGAGAT-3'-TEG-biotin |
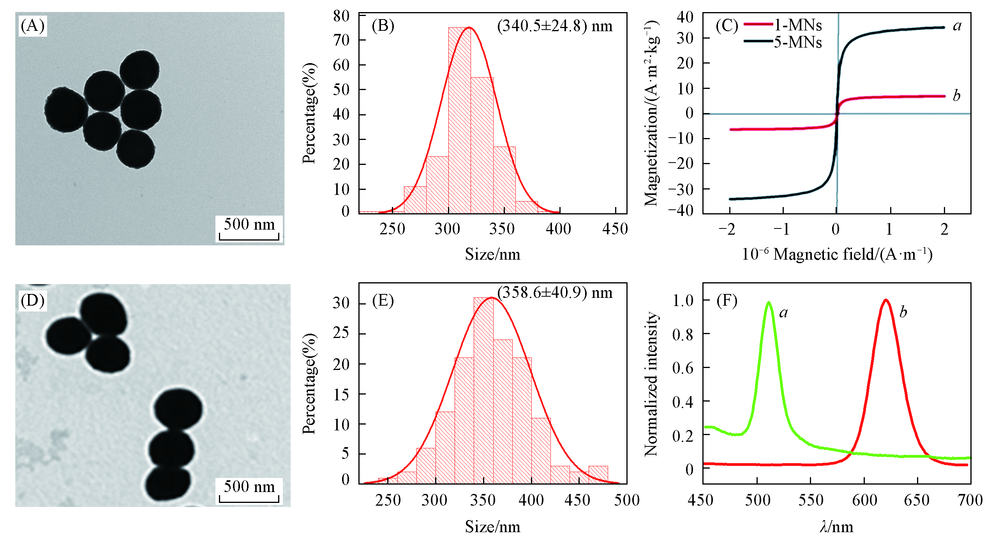
Fig.1 Characterization of beads with strong magnetic responsiveness(5-MNs) and weak magnetic responsiveness(1-MNs) (A, D) TEM images; (B, E) particle size statistics of 1-MNs(A, B) and 5-MNS(D, E); (C) magnetic hysteresis loops for 5-MNs(a) and 1-MNs(b); (F) fluorescence spectra of 1-GMNs(a) and 5-RMNs(b).
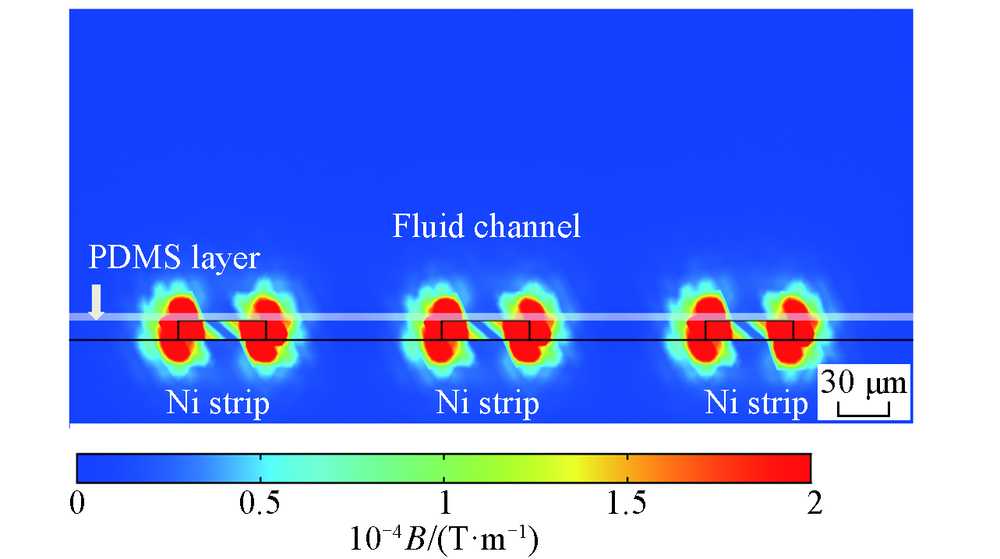
Fig.2 Simulation result of the magnetic field gradient(?B) around Ni strips Each Ni strips was 50 μm in width and 11 μm in height. The interval between Ni strips was 100 μm. The thickness of ITO glass under the Ni strips was 1.1 mm. Ni strips were covered by an extremely thin PDMS layer. The fluid channel was above the Ni strips.
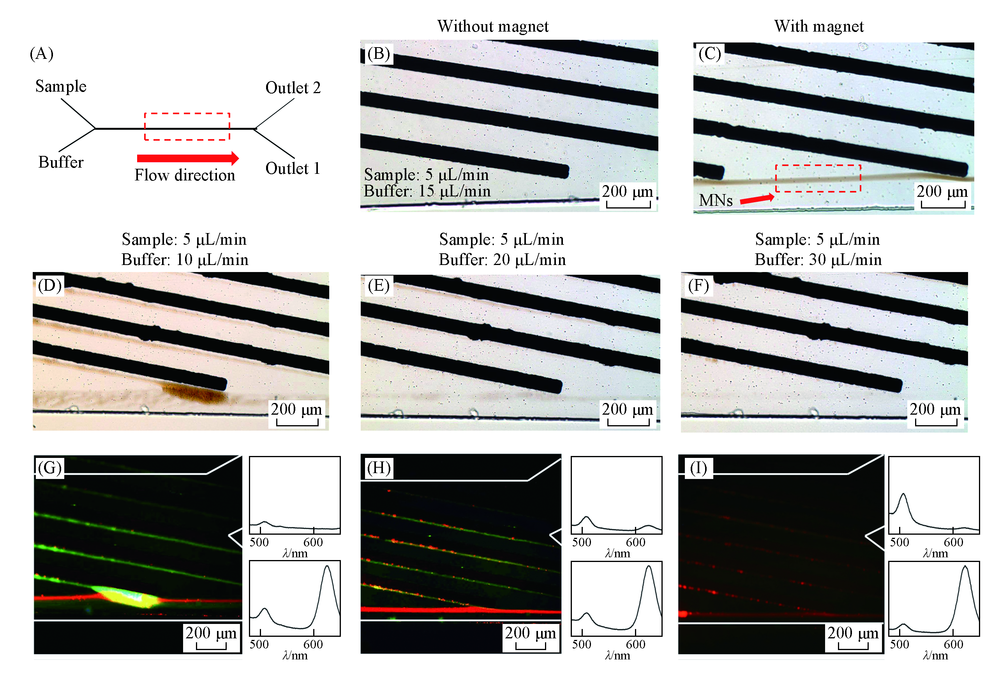
Fig.4 On-chip movement of 5-RMNs and 1-GMN at different flow rates (A) Structure of the chip; (B)—(F) images were taken in a bright field; (B) without magnetic field, MNs did not move along the Ni strips; (C) when an external magnetic field was added, the MNs became magnetized and clumped into visible micro-sized particles; (D)—(F) images of the motion of MNs at different flow rates; (G)—(I) fluorescence field images and fluorescence spectra of the fluorescent MNs collected at each outlet.
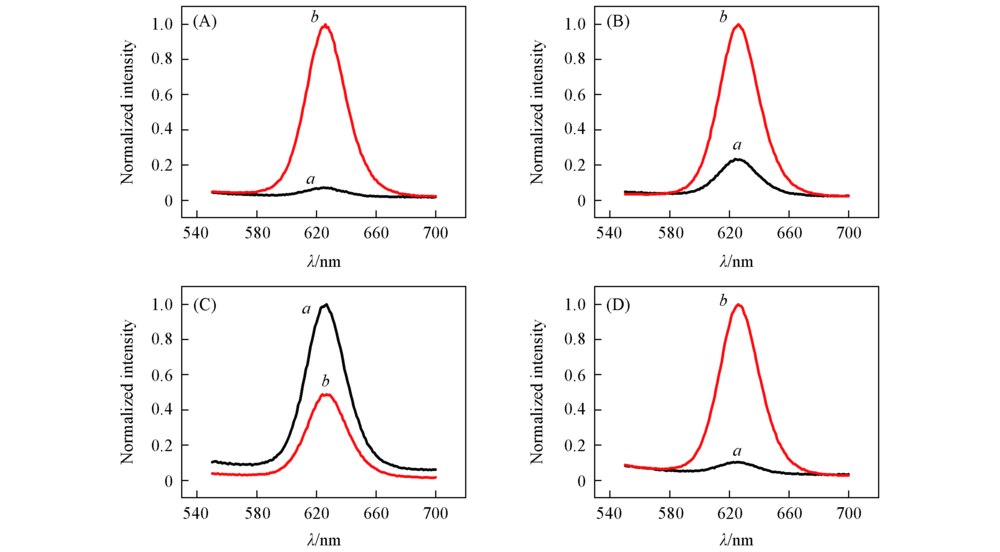
Fig.5 Fluorescence spectra of samples The flow rate of the sample was 5 μL/min and of the buffer was 30 μL/min. (A) Sample with only 1-RMNs; (B)—(D) samples with 5-MNs/1-RMNs mass ratios of 1:1, 2:1, 1:2, respectively. a. Outlet 1; b. outlet 2.
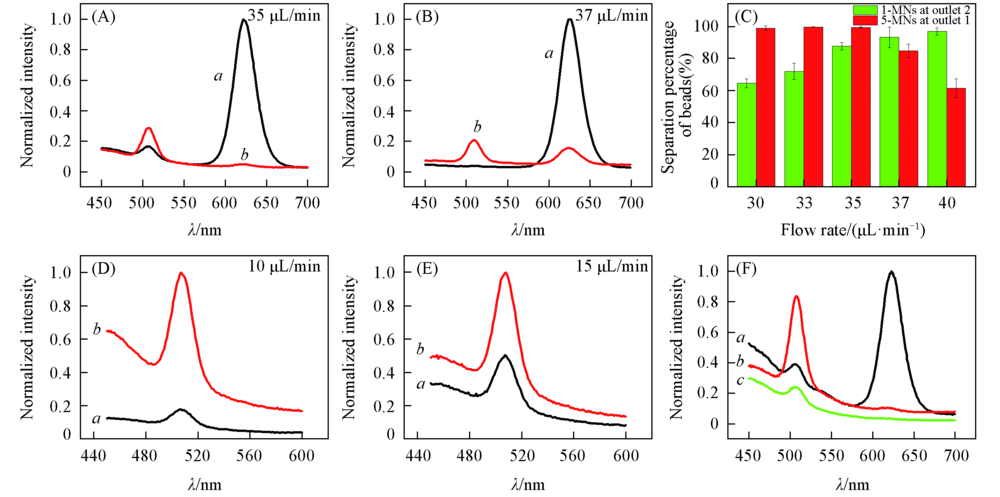
Fig.6 Separation efficiency of 5-RMNs and 1-GMNs Outlet 1 collected 5-MNs and outlet 2 collected 1-MNs. (A), (B) Separation efficiency of 5-RMNs and 1-GMNs by chip 1. (A) Flow rates: sample 5 μL/min, buffer 30 μL/min; (B) flow rates: sample 5 μL/min, buffer 32 μL/min; (C) relative separation percentages of 1-GMNs and 5-RMNs with flow rates by chip 1. Standard deviation was calculated from three experimental runs; (D), (E) separation efficiency of 1-GMNs by chip 2; (D) flow rates: sample 5 μL/min, buffer 5 μL/min; (E) flow rates: sample 5 μL/min, buffer 10 μL/min; (F) separation efficiency of 5-RMNs and 1-GMNs for the chip system. a. Outlet 1; b. outlet 2; c. waste.
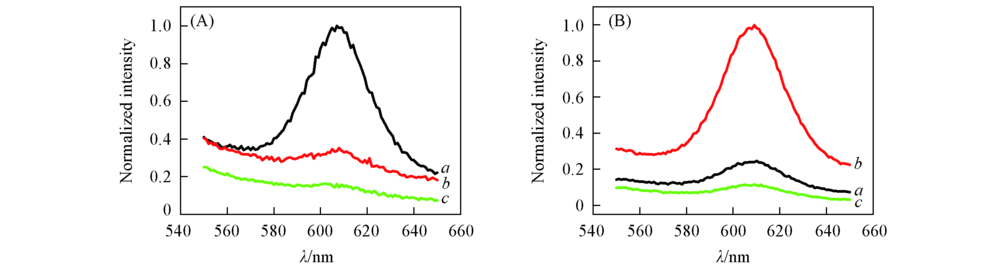
Fig.7 Separation of target DNA of HBV and HCV from the sample Distributions of HBV DNA(A) and HCV complementary DNA(B) at outlet 1(a), outlet 2(b) and the waste outlet(c).
| [1] | Xia N., Hunt T. P., Mayers B. T., Biomod. Microdev., 2006, 8, 299—308 |
| [2] |
Mizuno M., Yamada M., Mitamura R., Ike K., Toyama K., Seki M., Anal. Chem., 2013, 85, 7666—7673
doi: 10.1021/ac303336f URL |
| [3] |
Cho C. F., Lee K., Speranza M. C., Bononi F. C., Viapiano M. S., Luyt L. G., Weissleder R., Chiocca E. A., Lee H., Lawler S. E., ACS Comb. Sci., 2016, 18, 271—278
doi: 10.1021/acscombsci.5b00180 URL |
| [4] | Inglis D. W., Riehn R., Sturm J. C., Austin R. H., J. Appl. Phys., 2006, 99(8K), 101—109 |
| [5] |
Han K. H., Frazier A. B., J. Appl. Phys., 2004, 96(10), 5797—5763
doi: 10.1063/1.1803628 URL |
| [6] |
Furlani E. P., J. Phys. D, 2007, 40, 1313—1331
doi: 10.1088/0022-3727/40/5/001 URL |
| [7] |
Chung J., Issadore D., Ullal A., Lee K., Weissleder R., Lee H., Biomicrofluidics, 2013, 7, 054107
doi: 10.1063/1.4821923 URL |
| [8] |
Kim U., Soh H. T., Lab Chip, 2009, 9, 2313—2318
doi: 10.1039/b903950c URL |
| [9] |
Yassine O., Gooneratne C. P., Smara D. A., Li F., Mohammed H., Merzaban J., Kosel J., Biomicrofluidics, 2014, 8, 034114
doi: 10.1063/1.4883855 URL |
| [10] |
Lin S. J., Zhi X., Chen D., Xia F. F., Shen Y. H., Niu J. Q., Huang S. Y., Song J., Miao J. M., Cui D. X., Ding X. T., Biosens. Bioelectron., 2019, 129, 175—181
doi: 10.1016/j.bios.2018.12.058 URL |
| [11] |
Guo P. L., Tang M., Hong S. L., Yu X., Pang D. W., Zhang Z. L., Biosens. Bioelectron., 2015, 74, 628—636
doi: 10.1016/j.bios.2015.07.019 URL |
| [12] |
Lee J. J., Jeong K. J., Hashimoto M., Kwon A. H., Rwei A., Shankarappa S. A., Tsui J. H., Kohane D. S., Nano Lett., 2014, 14, 1—9
doi: 10.1021/nl3047305 URL |
| [13] |
Kwon K., Gwak H., Hyun K. A., Kwak B. S., Jung H. H., Sens. Actuators B, 2019, 294, 62—68
doi: 10.1016/j.snb.2019.05.007 URL |
| [14] |
Lee H. Y., Jung J. H., Han S. I., Han K. K., Lab Chip, 2010, 10, 2764—2770
doi: 10.1039/c005145d URL |
| [15] |
Hale C., Darabi J., Biomicrofluidics, 2014, 8, 044118
doi: 10.1063/1.4893772 URL |
| [16] |
Xu H. Y., Liao C., Zuo P., Liu Z. W., Ye B. C., Anal. Chem., 2018, 90, 13451—13459
doi: 10.1021/acs.analchem.8b03272 URL |
| [17] |
Chen W. W., Li H. J., Su W. T., Qin J. H., Biomicrofluidics, 2019, 13, 054113
doi: 10.1063/1.5110973 URL |
| [18] |
Song S. H., Lee H. L., Min Y. H., Jung H. l., Sens. Actuators B, 2009, 141, 210—216
doi: 10.1016/j.snb.2009.06.037 URL |
| [19] |
Yung C. W., Fiering J., Mueller A. J., Ingber D. E., Lab Chip, 2009, 9, 1171—1177
doi: 10.1039/b816986a URL |
| [20] |
Yu X., Feng X., Hu J., Zhang Z. L., Pang D. W., Langmuir, 2011, 27, 5147—5156
doi: 10.1021/la104400m URL |
| [21] |
Adams J. D., Kim U., Soh H. T., PNAS, 2008, 105, 18165—18170
doi: 10.1073/pnas.0809795105 URL |
| [22] |
Kang B., Cha B., Kim B., Han S., Shin M. K., Jang E., Kim H. O., Bae S. R., Jeong U., Moon I., Son H. Y., Huh Y. M., Haam S., Anal. Chem., 2016, 88, 1078—1082
doi: 10.1021/acs.analchem.5b04111 URL |
| [23] | Xie H. Y., Zuo C., Liu Y., Zhang Z. L., Pang D. W., Li X. L., Gong J. P., Dickinson C., Zhou W. Z., Small, 2005, 5, 506—509 |
| [24] | Fujii T., Microelectron. Eng., 2002, 61, 907—915 |
| [25] |
Hatch G. P., Stelter R. E., J. Magn. Magn. Mater., 2001, 225, 262—276
doi: 10.1016/S0304-8853(00)01250-6 URL |
| [26] |
Hayes M. A., Polson N. A., Garcia A. A., Langmuir, 2001, 17, 2866—2871
doi: 10.1021/la001655g URL |
| [1] | 王方圆, 张奋娴, 李毅, 高建华, 牛颜冰, 申少斐. 仿生树叶模型的制作及在琼脂糖微流控芯片中的应用[J]. 高等学校化学学报, 2022, 43(11): 20220445. |
| [2] | 彭伙, 高则航, 廖承悦, 王晓冬, 周洪波, 赵建龙. 一种用于核酸绝对定量检测的高鲁棒性液滴式数字PCR芯片[J]. 高等学校化学学报, 2020, 41(8): 1760. |
| [3] | 王洁,张宇,于敏,方瑾. 基于双向浓度梯度的微流控芯片系统对肿瘤细胞侵袭能力的多重分析[J]. 高等学校化学学报, 2019, 40(12): 2494. |
| [4] | 袁浩钧, 郜晚蕾, 景奉香, 刘松生, 周洪波, 贾春平, 金庆辉, 赵建龙. 一种用于核酸高灵敏检测的液滴式数字聚合酶链式反应芯片[J]. 高等学校化学学报, 2017, 38(7): 1140. |
| [5] | 姜健, 李盼盼, 马滢雪, 杨春光, 徐章润. 微流控液液萃取-液液波导集成化分析系统[J]. 高等学校化学学报, 2016, 37(4): 648. |
| [6] | 解飞, 张莹, 金伟, 潘婷婷, 周超, 任昊, 金钦汉, 牟颖. 基于表面等离子体子共振成像技术研究苏拉明和顺铂对肝癌细胞生长的抑制作用[J]. 高等学校化学学报, 2014, 35(1): 30. |
| [7] | 霍丹群, 方可敬, 侯长军, 杨眉, 法焕宝, 黄承洪, 罗小刚. 水凝胶微流控芯片的快速加工及在细胞培养检测中的应用[J]. 高等学校化学学报, 2013, 34(6): 1327. |
| [8] | 朱强远, 杨文秀, 高一博, 于丙文, 邱琳, 周超, 金伟, 金钦汉, 牟颖. 一种可绝对定量核酸的数字PCR微流控芯片[J]. 高等学校化学学报, 2013, 34(3): 545. |
| [9] | 杨俊, 齐莉, 马会民, 陈义. 基于尼龙微孔薄膜的微流控芯片的制作及对葡萄糖的显色响应[J]. 高等学校化学学报, 2012, 33(11): 2405. |
| [10] | 郝丽, 程和勇, 刘金华, 殷学锋. 微流动注射-等离子体质谱直接测定白酒中铅和镉[J]. 高等学校化学学报, 2012, 33(09): 1957. |
| [11] | 张小玲, 杨军, 胡宁, 侯文生, 郑小林, 谢琳, 杨忠, 陈洁. 基于薄膜电极的细胞电融合芯片[J]. 高等学校化学学报, 2012, 33(08): 1698. |
| [12] | 李栋顺, 徐溢, 彭金兰, 周桢, 甘俊, 王尧. 微流控细胞芯片LED诱导荧光检测微系统[J]. 高等学校化学学报, 2012, 33(01): 49. |
| [13] | 陈肇娜, 刘宝红, 吴会灵, 杨芃原. 微流控免疫芯片检测方法的研究进展[J]. 高等学校化学学报, 2011, 32(5): 1001. |
| [14] | 徐春秀, 殷学锋. 微流控芯片中细胞动态胞吞二氧化硅纳米颗粒的研究[J]. 高等学校化学学报, 2010, 31(9): 1712. |
| [15] | 徐章润, 李娜, 张惠丹, 樊晓峰, 方瑾. 基于热阻梯度加热系统的温度梯度芯片毛细管电泳用于DNA突变检测[J]. 高等学校化学学报, 2010, 31(4): 667. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||