

高等学校化学学报 ›› 2020, Vol. 41 ›› Issue (12): 2690.doi: 10.7503/cjcu20200388
收稿日期:2020-06-24
出版日期:2020-12-10
发布日期:2020-12-09
通讯作者:
陈填烽
E-mail:tchentf@jnu.edu.cn
基金资助:
YUAN Zhongwen, HE Lizhen, CHEN Tianfeng( )
)
Received:2020-06-24
Online:2020-12-10
Published:2020-12-09
Contact:
CHEN Tianfeng
E-mail:tchentf@jnu.edu.cn
摘要:
单原子催化剂(SACs)具有100%的原子利用率及充分暴露的原子金属活性位点, 其催化活性和选择性更具优势, 已广泛应用在化学、 能源及环境等领域. 近年来, SACs在生物医学领域也引起了广泛关注. 本文综述了SACs在肿瘤治疗、 抗菌、 抗氧化和生物传感等生物医学领域的应用及研究进展, 并简要总结了SACs未来应用的挑战和机遇, 为合理设计多性能的SACs提供了可行策略.
中图分类号:
TrendMD:
袁中文, 贺利贞, 陈填烽. 单原子催化剂的生物医学应用. 高等学校化学学报, 2020, 41(12): 2690.
YUAN Zhongwen, HE Lizhen, CHEN Tianfeng. Biomedical Applications of Single-atom Catalysts. Chem. J. Chinese Universities, 2020, 41(12): 2690.
| Material | Metal atom | Supporting material | Biological application | Reference |
|---|---|---|---|---|
| OxgeMCC?r SAEs | Ru | MOF | Tumor therapy | [ |
| PSAF NCS | Fe | Amorphous carbon | Tumor therapy | [ |
| P?MOF | Fe | Porphyrin?MOF | Tumor therapy | [ |
| MitoCAT?g | Au | Carbon dots | Tumor therapy | [ |
| SeNCM | Se | Amorphous carbon | ― | [ |
| PMCS | Zn | Amorphous carbon | Antibacterial | [ |
| SAF?NCs | Fe | Amorphous carbon | Antibacterial | [ |
| FeN5 SA/CNF | Fe | Amorphous carbon | Antibacterial | [ |
| Fe?SAs/NC | Fe | Amorphous carbon | Antioxidant | [ |
| Fe?N/C SACs | Fe | Amorphous carbon | Antioxidant | [ |
| Co/PMCS | Co | Amorphous carbon | Antioxidant | [ |
| Pt/CeO2 | Pt | Ceria | Antioxidant | [ |
| Fe?N?C SAN | Fe | Amorphous carbon | Biosensor | [ |
| Fe?N/C | Fe | Amorphous carbon | Biosensor | [ |
| Fe?N?C SAzyme | Fe | Amorphous carbon | Biosensor | [ |
| CNT/FeNC | Fe | Carbon nanotube | Biosensor | [ |
| Fe?N?rGO | Fe | Graphene | Biosensor | [ |
| Fe SSN | Fe | Amorphous carbon | Biosensor | [ |
Table 1 Examples of application of SACs in biomedical field in recent years
| Material | Metal atom | Supporting material | Biological application | Reference |
|---|---|---|---|---|
| OxgeMCC?r SAEs | Ru | MOF | Tumor therapy | [ |
| PSAF NCS | Fe | Amorphous carbon | Tumor therapy | [ |
| P?MOF | Fe | Porphyrin?MOF | Tumor therapy | [ |
| MitoCAT?g | Au | Carbon dots | Tumor therapy | [ |
| SeNCM | Se | Amorphous carbon | ― | [ |
| PMCS | Zn | Amorphous carbon | Antibacterial | [ |
| SAF?NCs | Fe | Amorphous carbon | Antibacterial | [ |
| FeN5 SA/CNF | Fe | Amorphous carbon | Antibacterial | [ |
| Fe?SAs/NC | Fe | Amorphous carbon | Antioxidant | [ |
| Fe?N/C SACs | Fe | Amorphous carbon | Antioxidant | [ |
| Co/PMCS | Co | Amorphous carbon | Antioxidant | [ |
| Pt/CeO2 | Pt | Ceria | Antioxidant | [ |
| Fe?N?C SAN | Fe | Amorphous carbon | Biosensor | [ |
| Fe?N/C | Fe | Amorphous carbon | Biosensor | [ |
| Fe?N?C SAzyme | Fe | Amorphous carbon | Biosensor | [ |
| CNT/FeNC | Fe | Carbon nanotube | Biosensor | [ |
| Fe?N?rGO | Fe | Graphene | Biosensor | [ |
| Fe SSN | Fe | Amorphous carbon | Biosensor | [ |
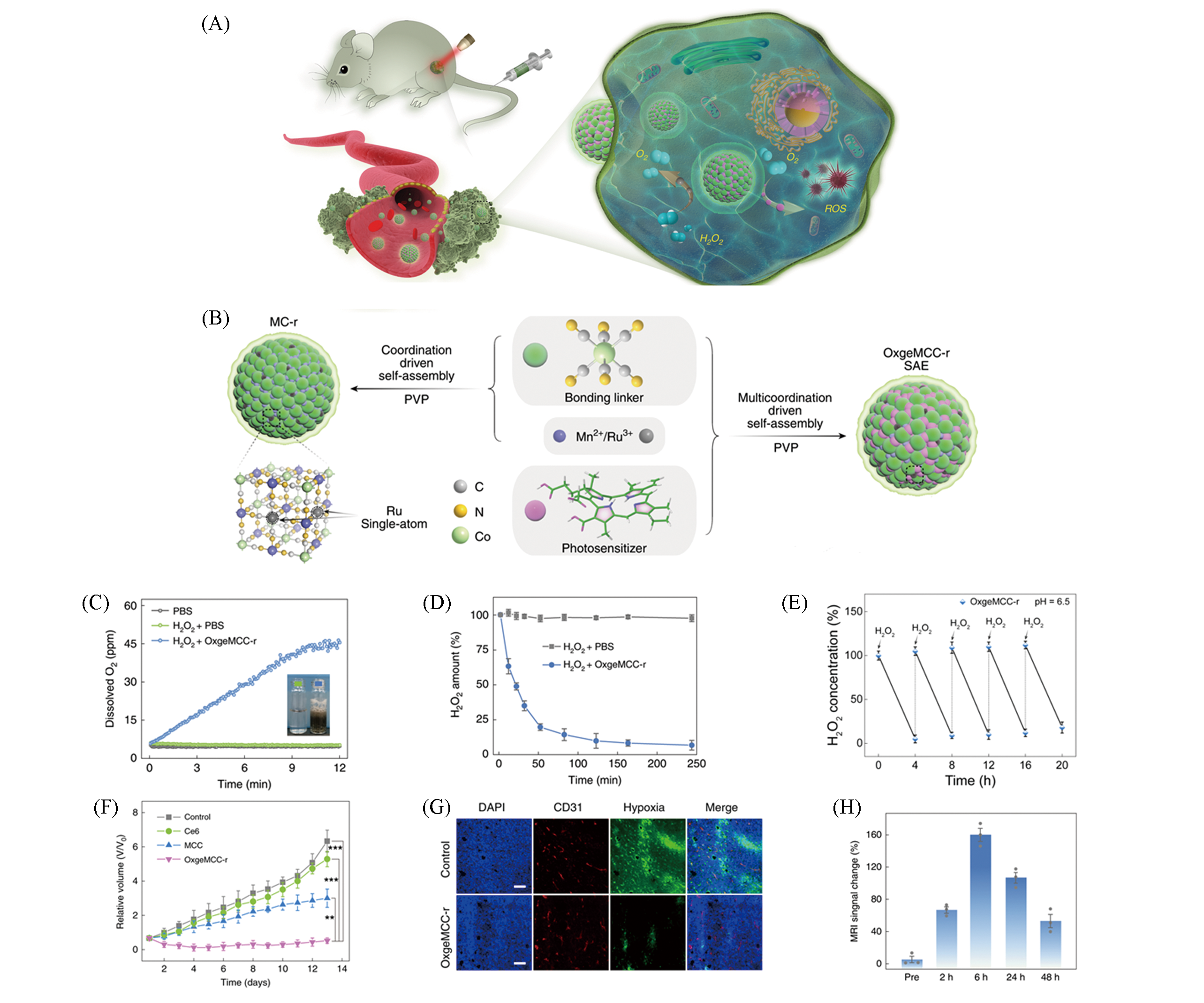
Fig.2 Self?assembled monoatomic nanozyme OxgeMCC?r SAE enhancing tumor PDT[30](A) Schematic diagram of OxgeMCC?r SAE continuous catalyzing production of O2 and ROS to enhance tumor PDT; (B) schematic diagram of OxgeMCC?r SAE synthesis; (C) O2 release curve; (D) H2O2 degradation curve; (E) OxgeMCC?r SAE catalytic stability test; (F) relative tumor volume of mice in different treatment groups, all three groups except the control group were irradiated with a laser with a power density of 100 mW/cm2 for 5 min; (G) immuno fluorescence images of tumor sections, nucleus(blue), vascular(red) and hypoxic zone(green), scale bar is 100 μm; (H) quantified map of T1?weighted magnetic resonance imaging capacity of the tumor site.Copyright 2020, Springer Nature.
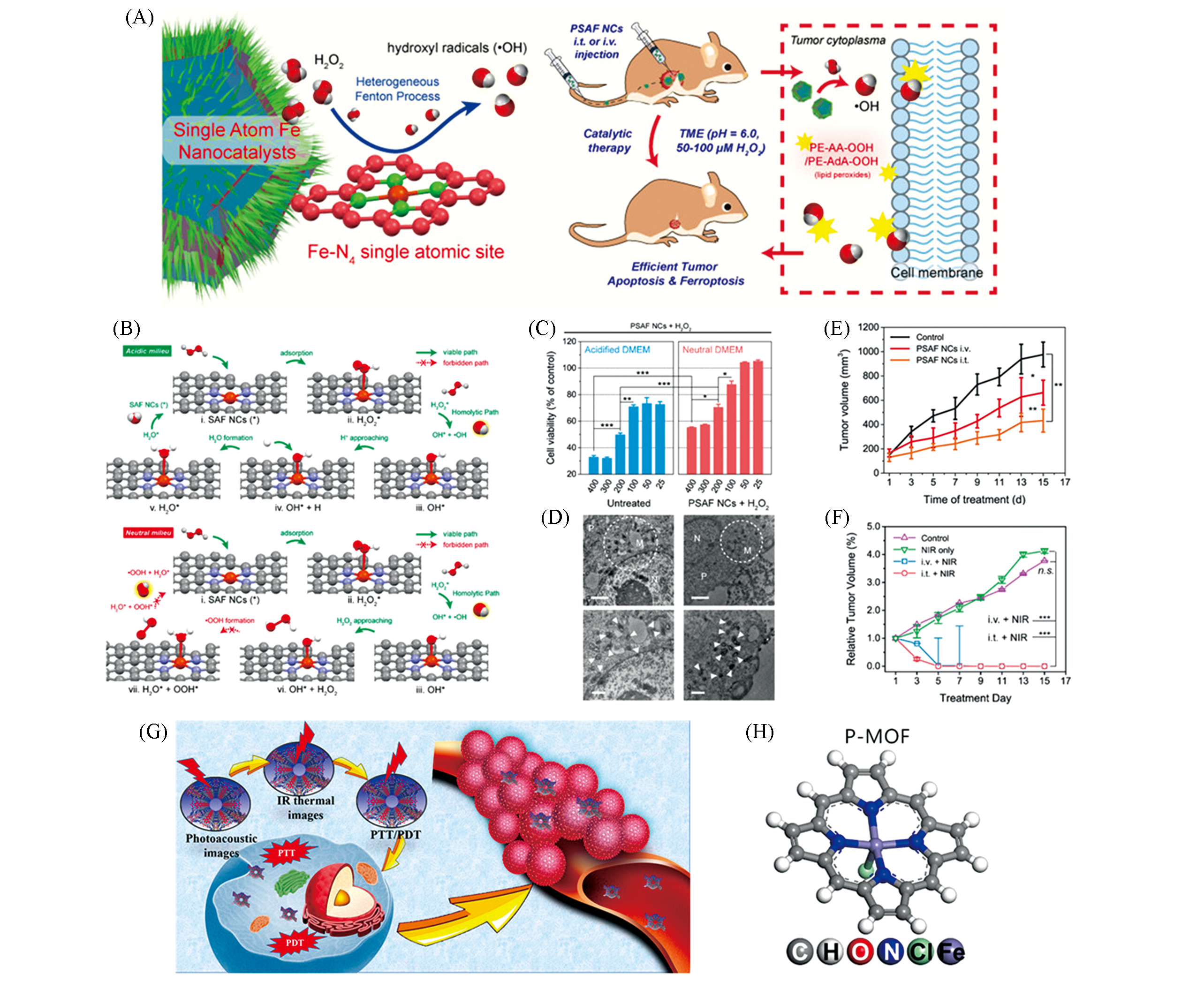
Fig.3 Tumor therapy of monoatomic iron nano catalysts PSAF NCs[31] and porphyrin?like monoatomic iron nano systems P?MOF[32](A) Schematic diagram of PSAF NCs catalytic tumor treatment; (B) schematic diagram of DFT calculation of PSAF NCs and H2O2; (C) quantitative diagram of the effects of different concentrations of PSAF NCs combined with H2O2 on 4T1 tumor cells viability in acidic and neutral media; (D) tumor cells evolutionary TEM image, the circular area and solid triangle represent the mitochondria(M) in the cell, and the cytoplasm and nucleus are denoted by P and N, respectively, scale bar is 1 μm; (E) the tumor volume change curve of mice in different treatment groups; (F) different treatment groups when combined with near?infrared light irradiation, change curve of mouse tumor volume; (G) multi?functional schematic diagram of P?MOF under near?infrared stimulation; (H) P?MOF optimized model.Copyright 2019, American Chemical Society.
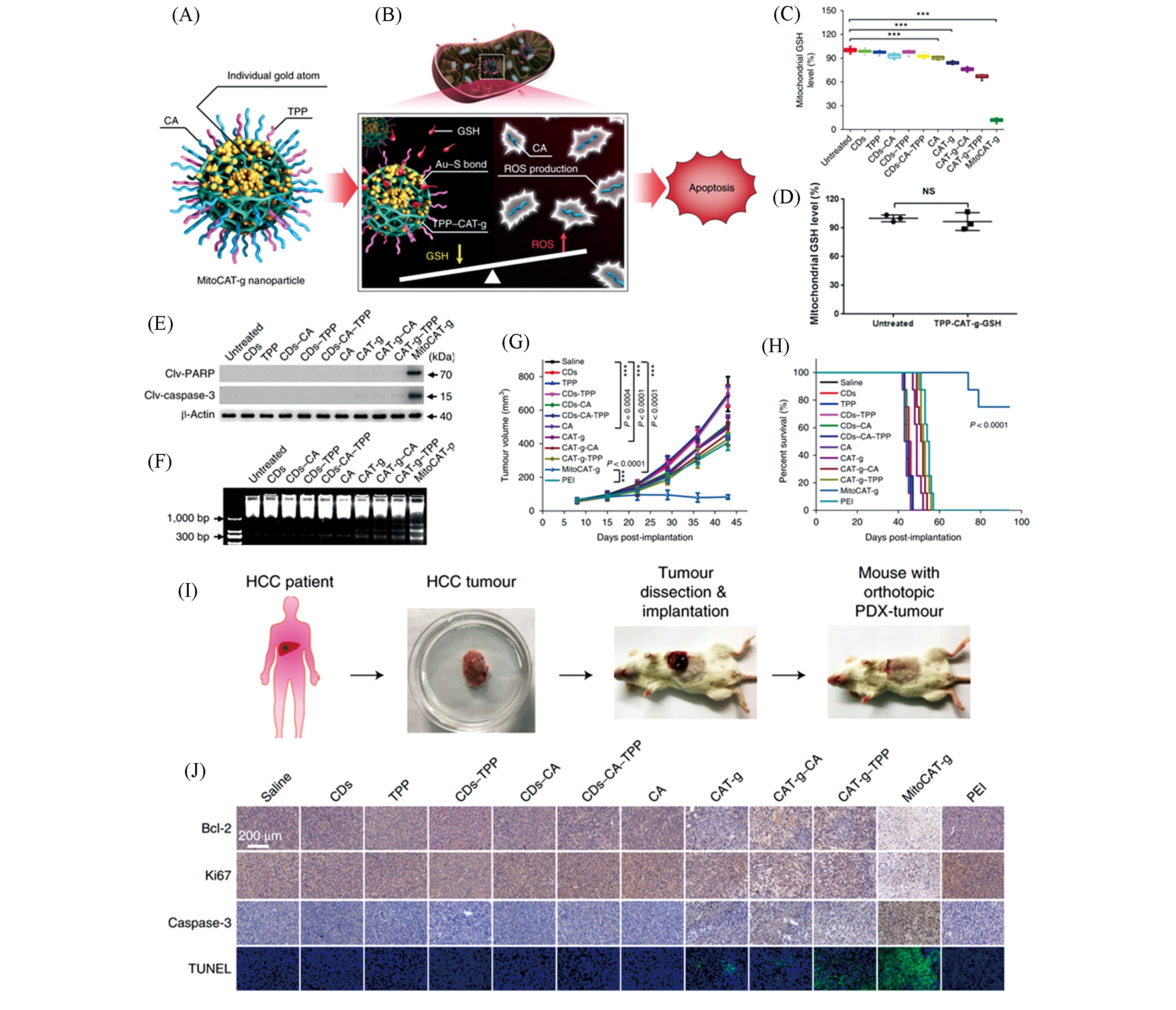
Fig.4 Oxidative stress for tumor therapy magnified by carbon dot?supported single?atom dispersed gold system MitoCAT?g[33](A) MitoCAT?g composition diagram; (B) schematic diagram of MitoCAT?g magnifying ROS levels to induce apoptosis; (C, D) mitochondrial GSH level of HepG?2 cells in different treatment groups; (E) expression of pro?apoptotic proteins; (F) HepG?2 cells were treated with different systems for 24 h, and the chromosomal DNA was broken; (G) tumor proliferation curves of mice in different treatment groups; (H) survival of mice in different treatment groups; (I) schematic diagram of establishing a PDX tumor model; (J) immunohistochemistry and TUNEL test results of different treatment groups.Copyright 2019, Springer Nature.
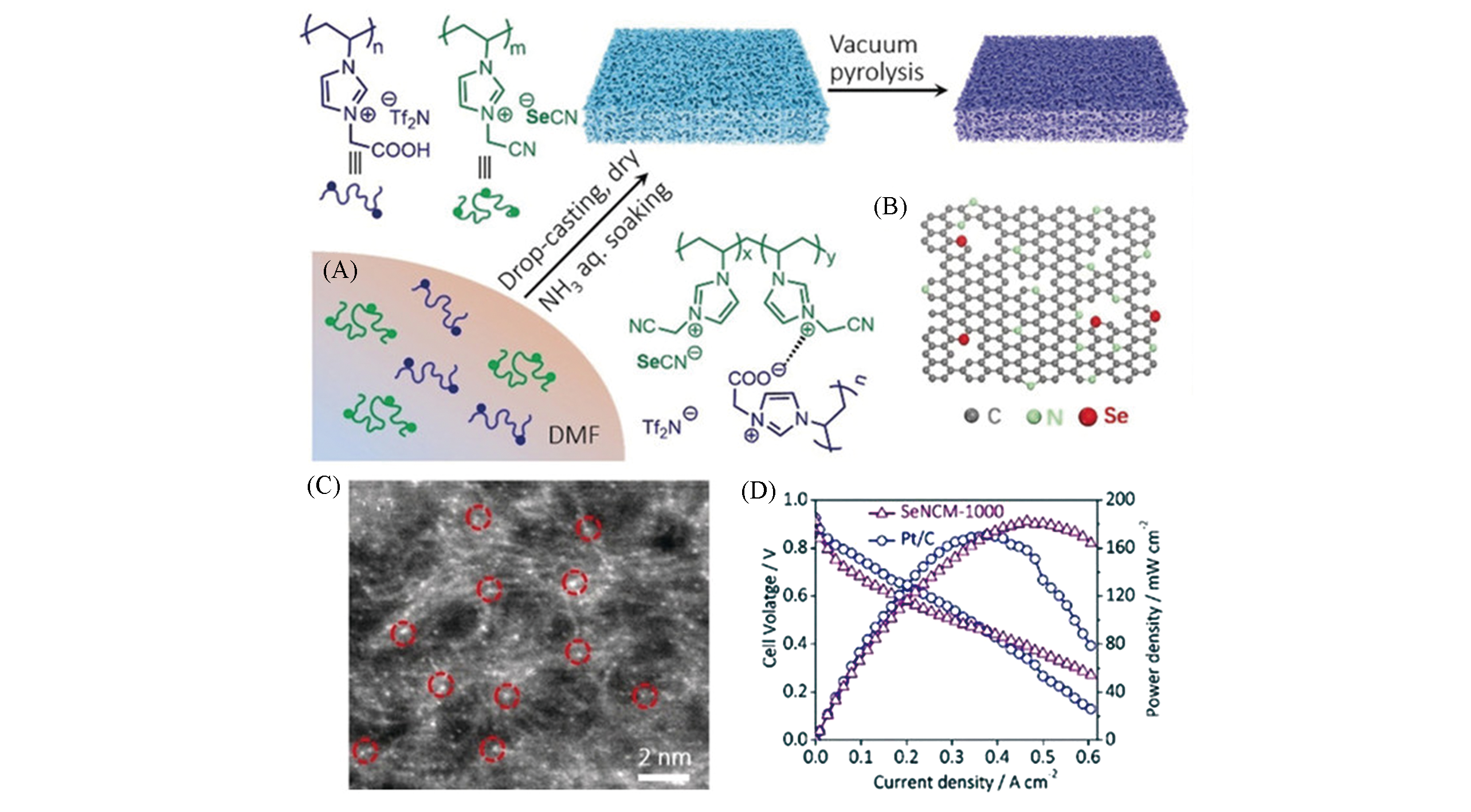
Fig.5 Synthesis and application of single atom selenium anchoring on porous carbon membrane[7](A) Preparation process for the SeNCM; (B) structural models of the SeNCM; (C) STEM image of the SeNCM?1000. SeNCM?1000: pyrolysis temperature is 1000 ℃; (D) the open?circuit potential, maximum current density and power density of SeNCM and Pt/C.Copyright 2019, Wiley?VCH.
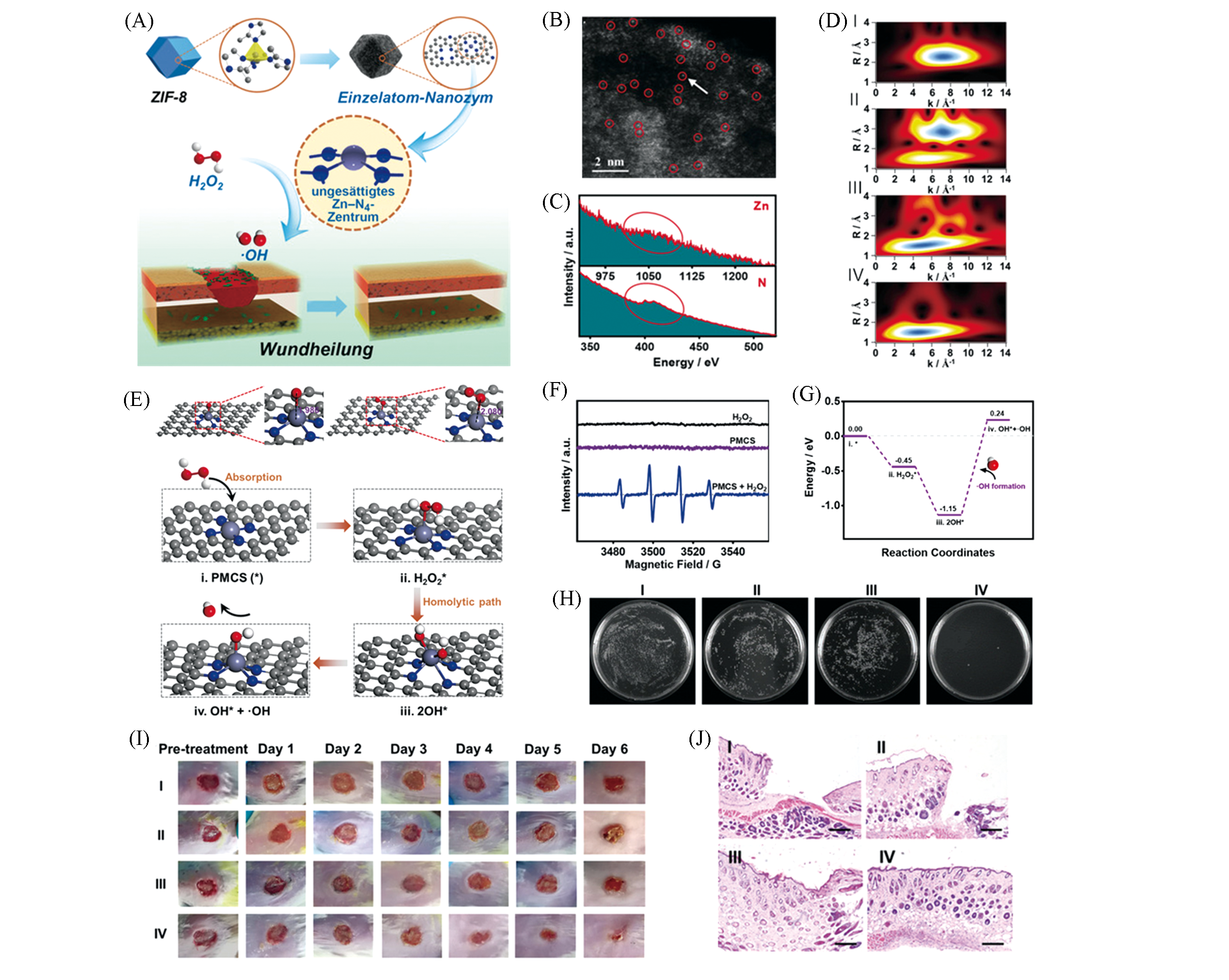
Fig.6 Monoatomic zinc nanozyme system PMCS used for antibacterial wound healing[5](A) Schematic diagram of antibacterial action of PMCS and H2O2; (B) HAADF?STEM diagram; (C) EELS diagram; (D) EXAFS diagram of PMCS; (E) catalytic mechanism of PMCS and H2O2 molecules producing ?OH; (F) ESR spectrum of ?OH; (G) calculation of free energy corresponding to the processes of graph (E); (H) pseudomonas aeruginosa exposed to NaAc buffer(Ⅰ), NaAc buffer combined with H2O2(Ⅱ), PMCS(Ⅲ) and PMCS(Ⅳ) combined with H2O2, colony formed photos; (I) pseudomonas aeruginosa infected wounds (Ⅰ―Ⅳ) photos after treatment at different times; (J) (Ⅰ―Ⅳ) histological staining of infected wounds after treating 6 days(Ⅰ―Ⅳ). Scale bar is 500 μm.Copyright 2019, Wiley?VCH.
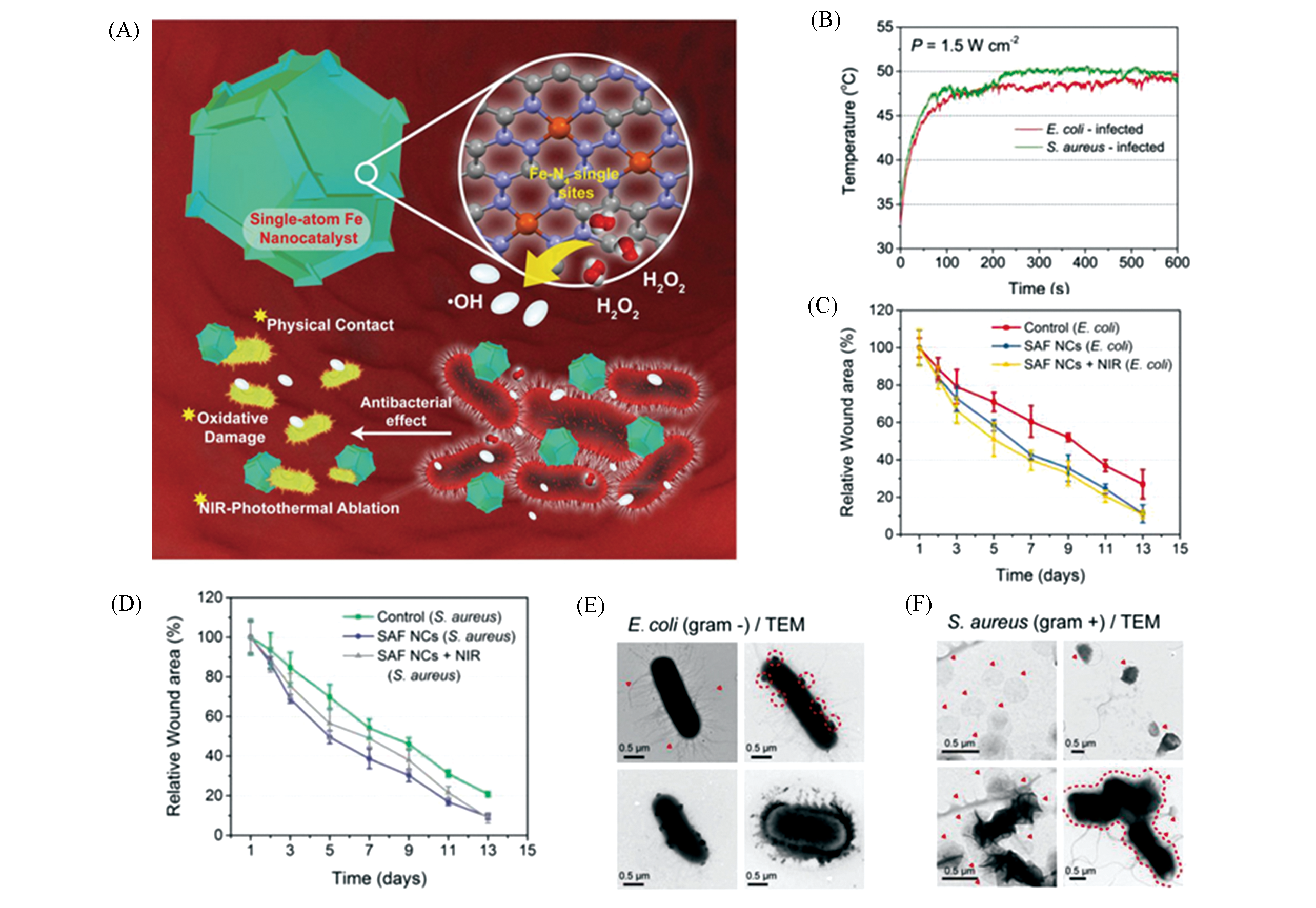
Fig.7 Monoatomic iron nano systems SAF?NCs combined with light and heat for antibacterial therapy[34](A) Schematic diagram of SAF?NCs used as antibiotics; (B) SAF?NCs and 808 nm laser irradiation for 10 min, the temperature?increasing curve of bacterial infection sites in the body; (C) relative wound area change curve of mice in E. coli; (D) S. aureus infection group after different treatments; evolution of morphology of E. coli(E)and S. aureus(F)treated by SAF?NCs combined with H2O2.Copyright 2019, Wiley?VCH.
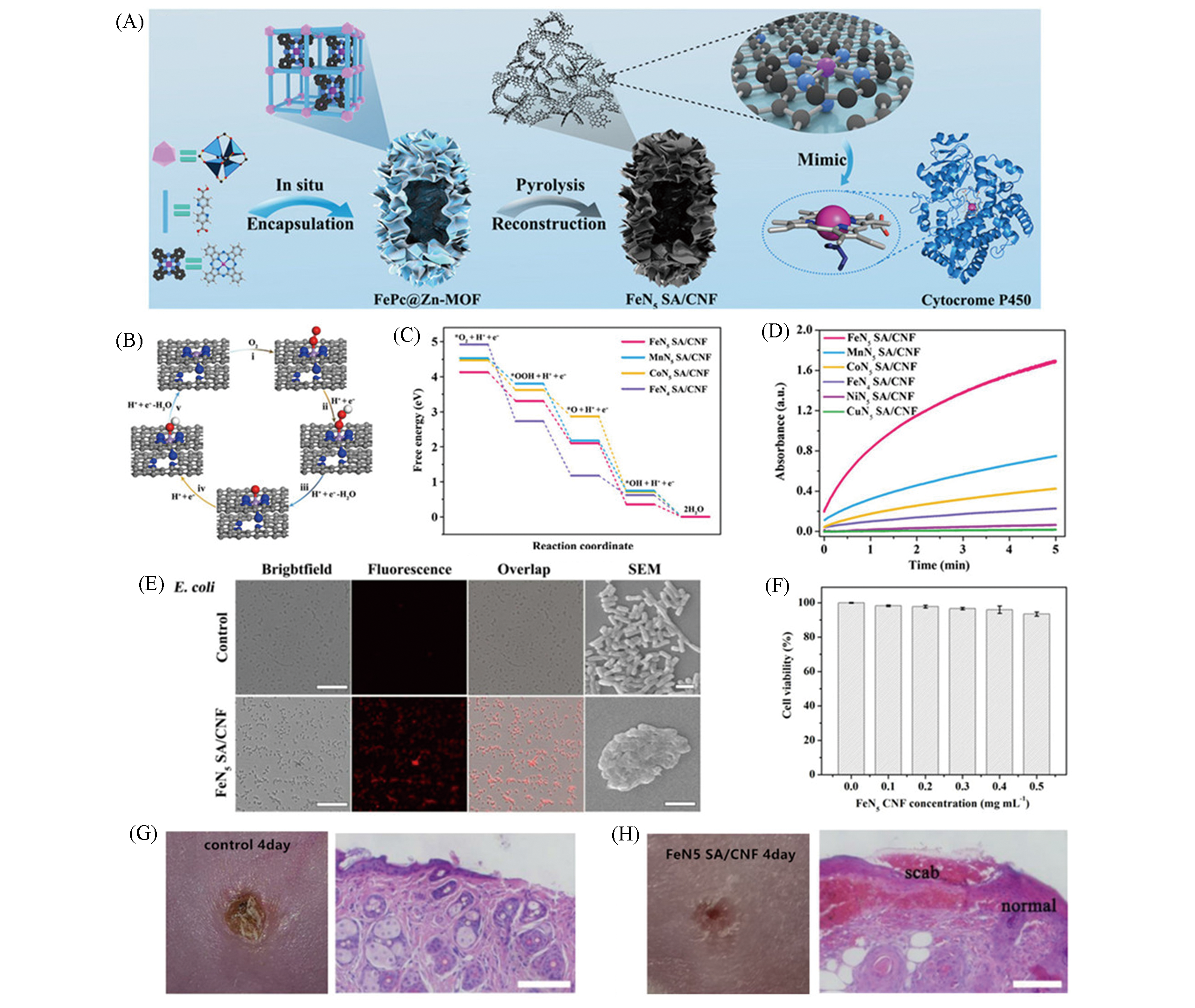
Fig.8 Monatomic iron nanozyme system FeN5 SA/CNF mimics cytochrome P450 for wound healing in mice[6](A) Schematic diagram of the process of FeN5 site formation; (B) DFT calculation of the OXD reaction pathway of FeN5 SA/CNF; (C) free energy map of each reaction process of FeN5 SA/CNF; (D) the changes curves of absorbance of different monoatomic enzymes; (E) image comparison before and after FeN5 SA/CNF treatment of E. coli, the scale bars are 40 μm for fluorescence images and 2 μm for SEM images; (F) cell viability of NCM460 cells treated with FeN5 SA/CNF at different concentrations; (G, H) photographs and H&E staining of E. coli infected wounds on the fourth day, scale bars is 100 μm.Copyright 2019, American Association for the Advancement of Science.
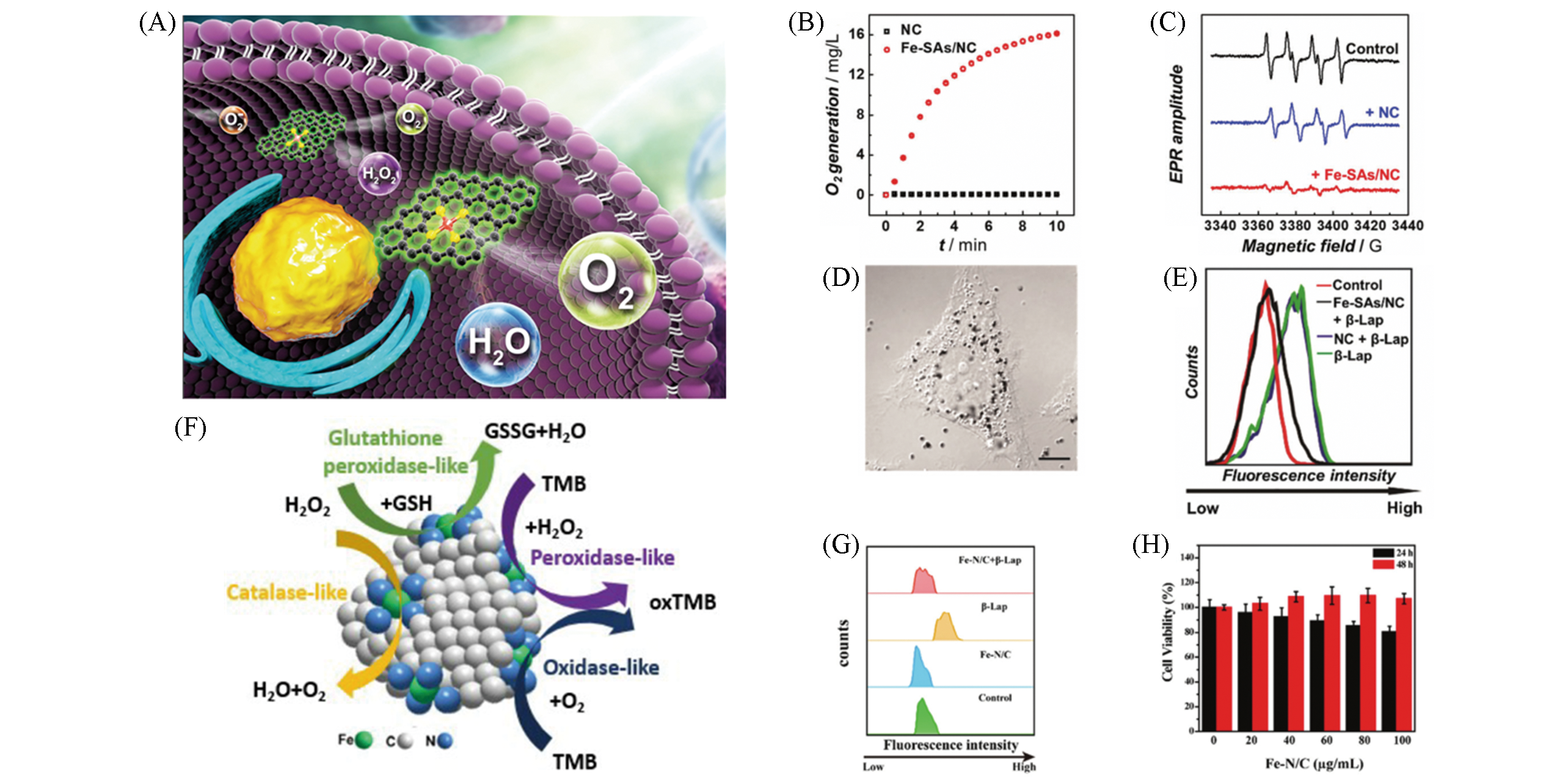
Fig.9 Multi?enzyme activities of monatomic iron nano systems Fe?SAs/NC[8] and Fe?N/C SACs[35](A) Fe?SAs/NC like?CAT, SOD schematic diagram; (B) Fe?SAs/NC oxygen production curve; (C) BMPO/*OOH ESR signal spectrum; (D) HeLa cells morphology under confocal laser scanning microscope, scale bar is 10 μm; (E) flow cytometry was used to detect the ROS clearance; (F) Fe?N/C SACs schematic diagram of multi?enzyme activity; (G) the clearance of ROS; (H) treated with different concentrations of Fe?N/C SACs HeLa cells vitality.(A—E) Copyright 2019, Royal Society of Chemistry; (F—H) Copyright 2019, Royal Society of Chemistry.
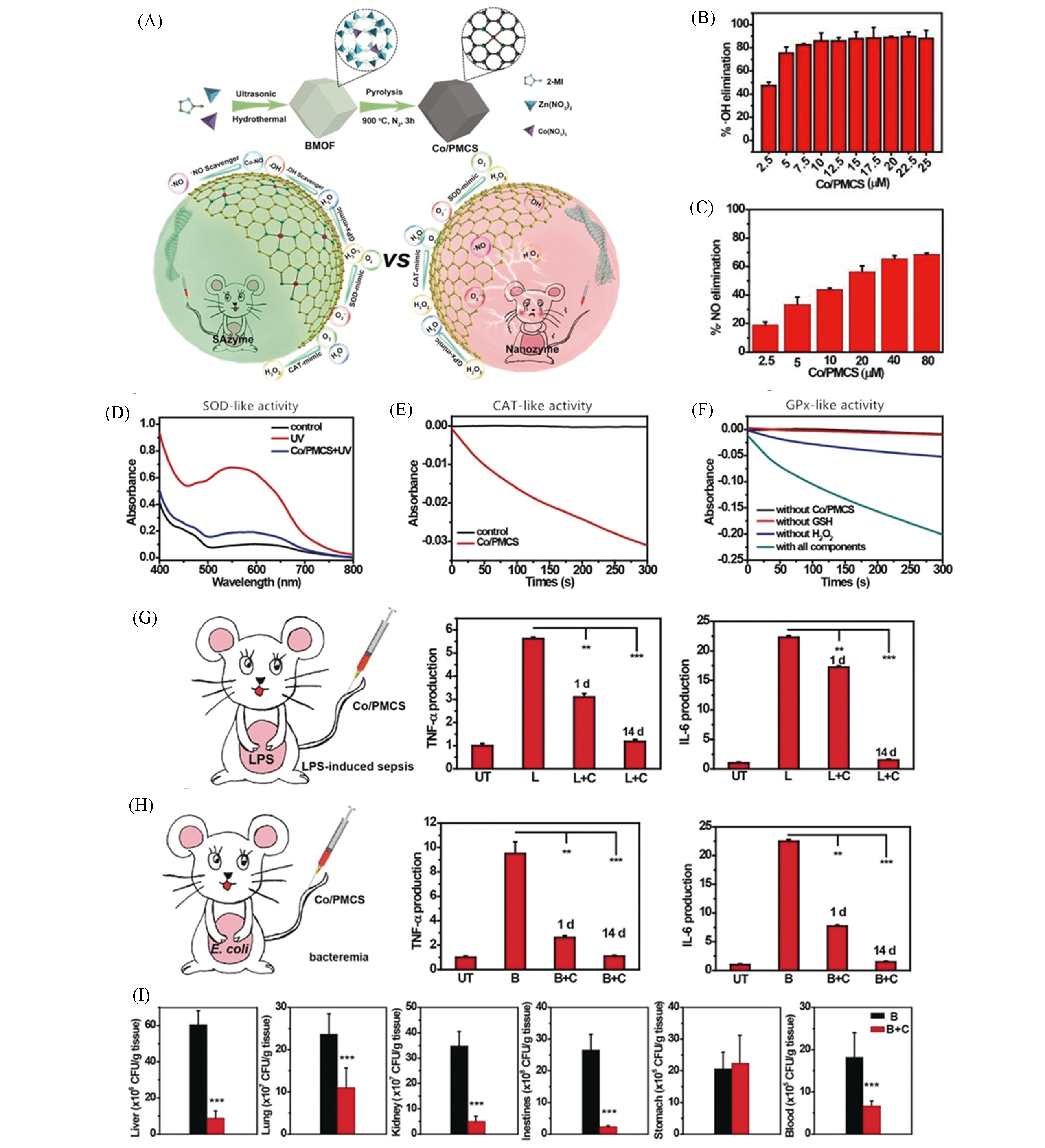
Fig.10 Monoatomic cobalt nanozyme system Co/PMCS used for the treatment of sepsis and bacteremia[36](A) Schematic diagram of Co/PMCS synthesis and as a multi?antioxidant treatment, compared with nanozymes; (B) different concentrations of Co/PMCS treatment for ?OH(B) and ?NO(C) removal rate; SOD(D), CAT(E) and GPx(F) activity; (G) TNF?α and IL?6 expression levels in sepsis mice, UT, L and C represent untreated, LPS?induced sepsis and Co/PMCS, respectively; (H) TNF?α and IL?6 expression levels in bacteremia mice, B represents bacteremia mice induced by E. coli; (I) with or without Co/PMCS treatment, the count of bacteria in liver, lung, kidney, intestine, stomach and blood of mice after 1 d.Copyright 2020, Wiley?VCH.
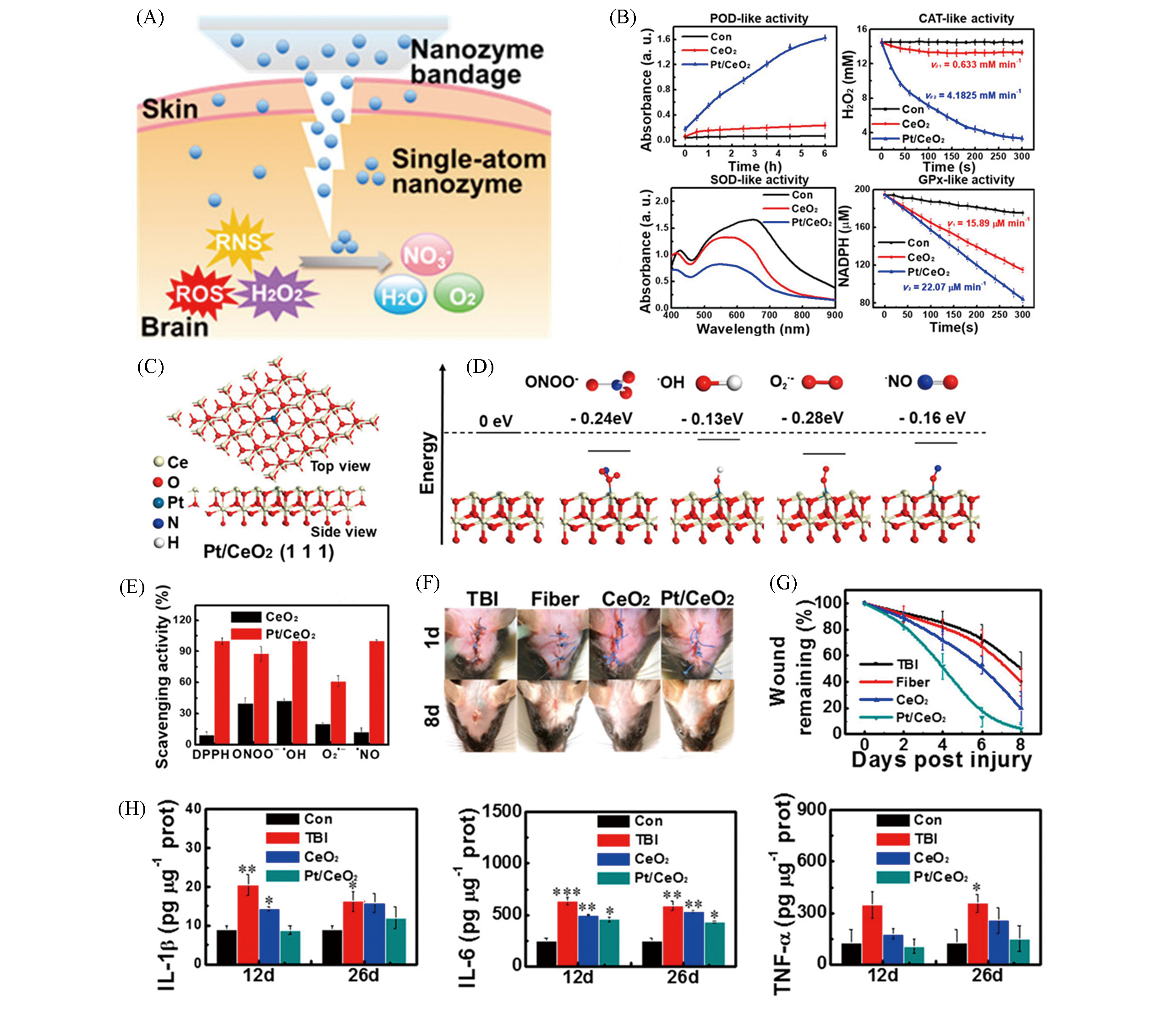
Fig.11 Monoatomic platinum nanozyme bandage Pt/CeO2 for the treatment of traumatic brain injury[37](A) Schematic diagram of Pt/CeO2 multi?enzyme activity for non?invasive TBI treatment; (B) multi?enzyme activity of Pt/CeO2 nanozyme(POD, CAT, SOD, GPx); (C) atomic structure of Pt/CeO2 (111) facets; (D) different values of the catalytic reaction energy with different free radical units; (E) the scavenging activity of Pt/CeO2 to RONS; (F) the representative photos of wound healing; (G) the corresponding time?residual wound curve after treatment in different treatment groups; (H) IL?1β, IL?6 and TNF?α level in mice.Copyright 2019, American Chemical Society.
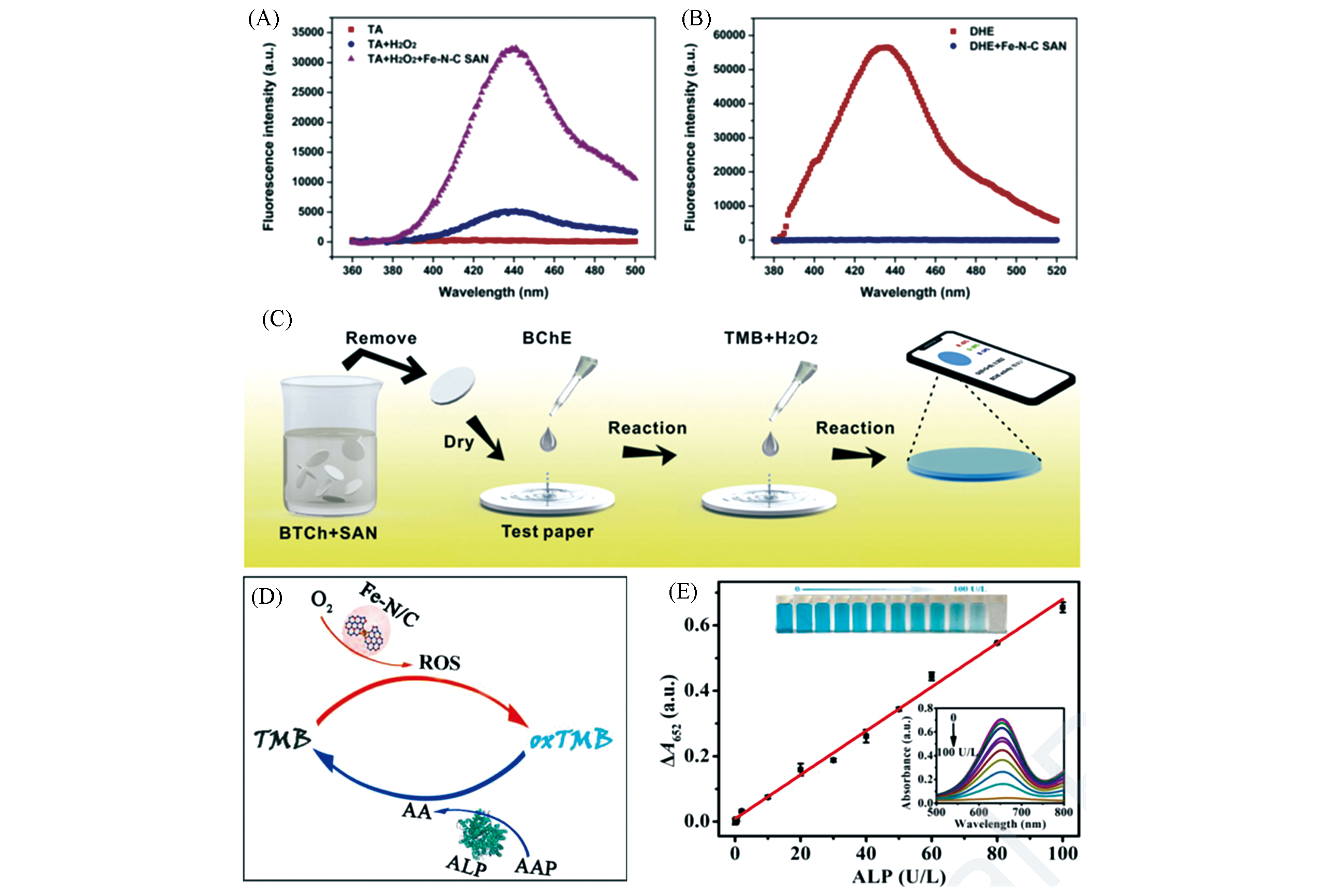
Fig.12 Monatomic iron nanozyme system Fe?N?C SAN used for butyryl cholinesterase(BChE)[38] and Fe?N/C for alkaline phosphatase(ALP)detection[39](A, B) Fluorescence spectra of different treatment groups; (C) preparation of bio?paper for BChE activity measurement; (D) schematic diagram of ALP activity measurement; (E) linear relationship diagram of different ALP concentrations and ΔA652, the illustration is Fe?N/C/TMB ultraviolet?visible absorption curve and photograph.(A―C) Copyright 2019, Elsevier; (D, E) Copyright 2020, Elsevier.
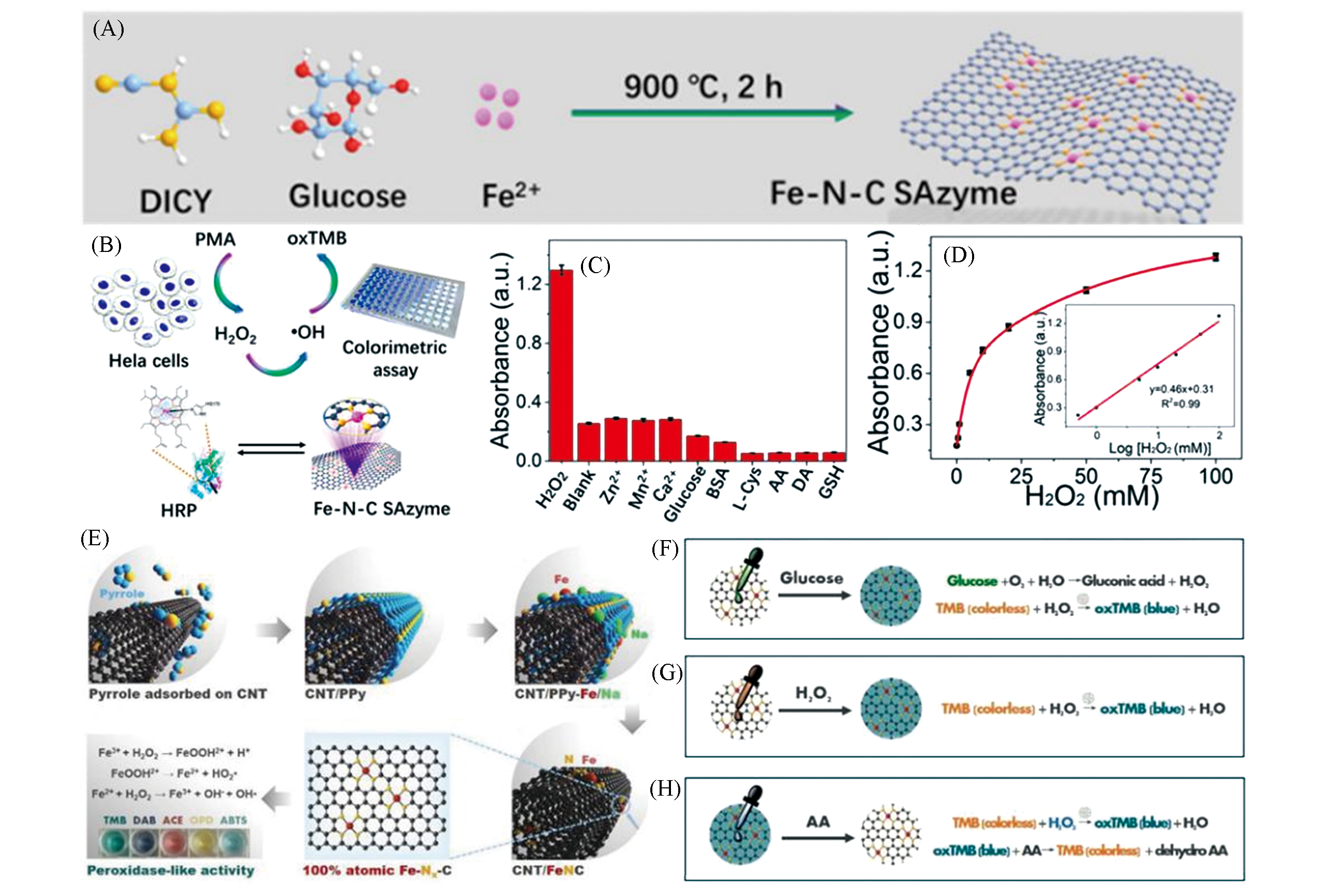
Fig.13 Single atomic iron nanozyme system Fe?N?C[40] and CNT/FeNC[41] for sensitive detection(A) Schematic diagram of Fe?N?C preparation; (B) Fe?N?C detection mechanism of H2O2 release in Hela cells; (C) Fe?N?C selective test for H2O2; (D) the calibration curve of Fe?N?C for oxidative TMB with different concentrations of H2O2; (E) synthesis and catalysis mechanism of CNT/FeNC; (F―H) are the reaction mechanisms of CNT/FeNC detection of H2O2, glucose and ascorbic acid.(A―D) Copyright 2019, American Chemical Society; (E―H) Copyright 2019, Wiley?VCH.
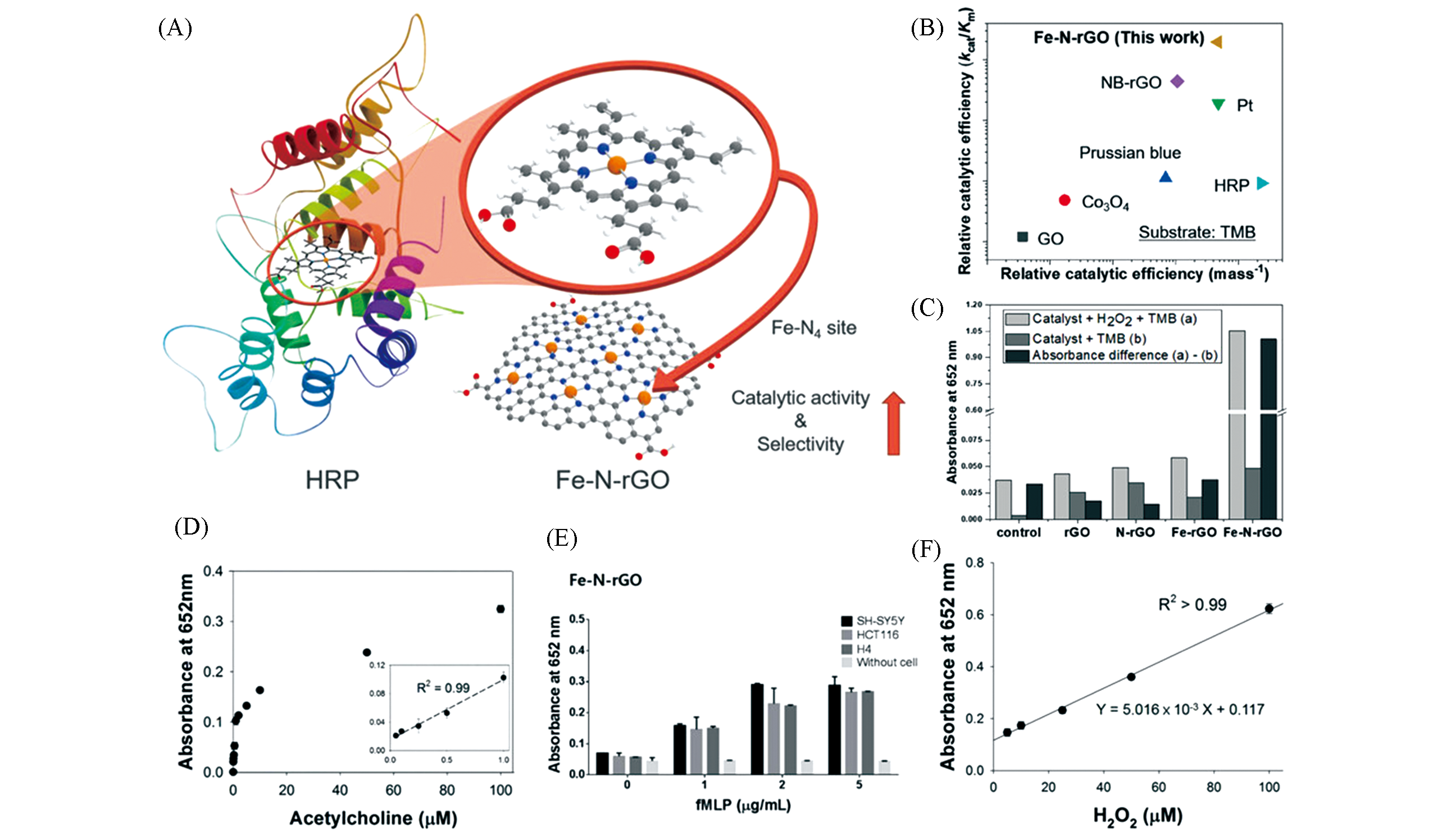
Fig.14 Monatomic iron nanozyme system Fe?N?rGO used for quantitative testing of acetylcholine(Ach) and cancer cells[42](A) Schematic diagram of Fe?N?rGO synthesis; (B) relative catalytic efficiency of different nanozymes; (C) absorption intensity at 652 nm of different treatment groups; (D) dose?response curve of Ach detected by Fe?N?rGO and its linear calibration chart(inset); (E) absorption intensity of fMLP at different concentrations; (F) calibration curve of H2O2 concentration?absorption intensity.Copyright 2019, Wiley?VCH.
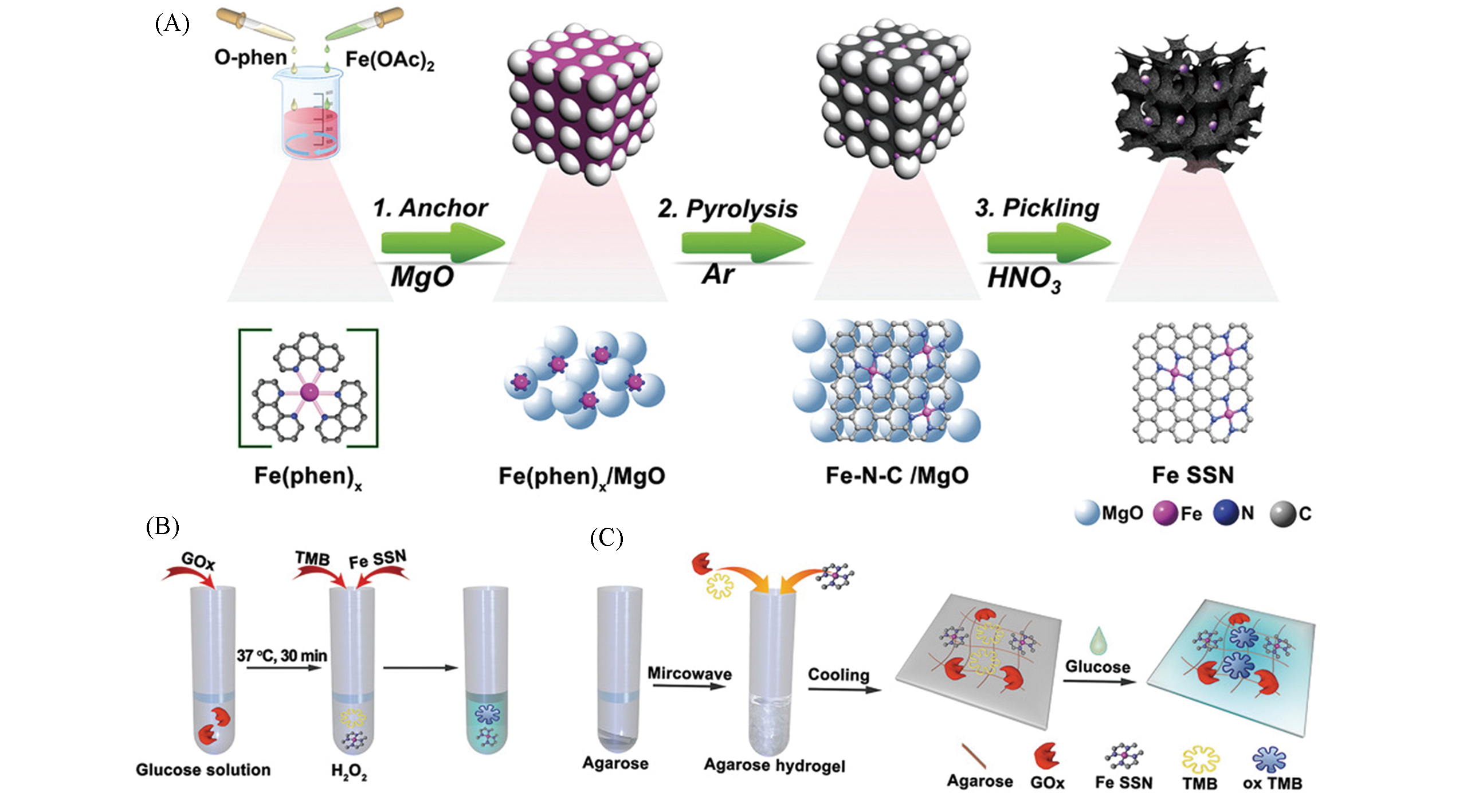
Fig.15 Single iron site nanozyme for ultrasensitive glucose detection[43](A) The schematic illustration for the preparation of Fe SSN; (B) process that cascade catalytic reaction for high sensitivity detection of glucose; (C) integrated agarose?based hydrogel is used for biosensor of glucose.Copyright 2020, Wiley?VCH.
| 1 | Mitchell S., Thomas J. M., Pérez⁃Ramírez J., Catal. Sci. Technol., 2017, 7, 4248―4249 |
| 2 | Jiao L., Yan H., Wu Y., Gu W., Zhu C., Du D., Lin Y., Angew. Chem. Int. Ed., 2020, 59, 2565―2576 |
| 3 | Qiao B., Wang A., Yang X., Allard L. F., Jiang Z., Cui Y., Liu J., Li J., Zhang T., Nat. Chem., 2011, 3, 634―641 |
| 4 | Fako E., Łodziana Z., López N., Catal. Sci. Technol., 2017, 7, 4285―4293 |
| 5 | Xu B., Wang H., Wang W., Gao L., Li S., Pan X., Wang H., Yang H., Meng X., Wu Q., Zheng L., Chen S., Shi X., Fan K., Yan X., Liu H., Angew. Chem. Int. Ed., 2019, 58, 4911―4916 |
| 6 | Huang L., Chen J., Gan L., Wang J., Dong S., Sci. Adv., 2019, 5, eaav5490 |
| 7 | Zhao C., Xiong C., Liu X., Qiao M., Li Z., Yuan T., Wang J., Qu Y., Wang X., Zhou F., Xu Q., Wang S., Chen M., Wang W., Li Y., Yao T., Wu Y., Li Y., Chem. Commun.(Camb), 2019, 552285―2288 |
| 8 | Ma W., Mao J., Yang X., Pan C., Chen W., Wang M., Yu P., Mao L., Li Y., Chem. Commun., 2019, 55, 159―162 |
| 9 | Xiang H., Feng W., Chen Y., Adv. Mater., 2020, 32, e1905994 |
| 10 | Gao L., Zhuang J., Nie L., Zhang J., Zhang Y., Gu N., Wang T., Feng J., Yang D., Perrett S., Yan X., Nat. Nanotechnol., 2007, 2, 577―583 |
| 11 | Wei H., Wang E., Chem. Soc. Rev., 2013, 42, 6060―6093 |
| 12 | Huang Y., Ren J., Qu X., Chem. Rev., 2019, 119, 4357―4412 |
| 13 | Wu J., Wang X., Wang Q., Lou Z., Li S., Zhu Y., Qin L., Wei H., Chem. Soc. Rev., 2019, 48, 1004―1076 |
| 14 | Zhang H., Lu X. F., Wu Z. P., Lou X. W. D., ACS Central. Sci., 2020 |
| 15 | Liu F., Qi X. J., Li Y. W., Xue M., Wang Y. Q., Zhao C. C., Acta Petrol. Sin. : Pet. Process. Section, 2020, 36, 428―438(刘芳, 齐学进, 李雨薇, 薛明, 王永强, 赵朝成. 石油学报: 石油加工, 2020, 36, 428―438) |
| 16 | Peng Y., Lu B., Chen S., Adv. Mater., 2018, 30, e1801995 |
| 17 | Zhang B., Fan T., Xie N., Nie G., Zhang H., Adv. Sci. (Weinh), 2019, 6, 1901787 |
| 18 | Zhao S., Chen G., Zhou G., Yin L. C., Veder J. P., Johannessen B., Saunders M., Yang S. Z., de Marco R., Liu C., Jiang S. P., Adv. Funct. Mater., 2019, 30, 1906157 |
| 19 | Liu P., Zhao Y., Qin R., Mo S., Chen G., Gu L., Chevrier D. M., Zhang P., Guo Q., Zang D., Wu B., Fu G., Zheng N., Science, 2016, 352, 797―801 |
| 20 | Nie L., Mei D., Xiong H., Peng B., Ren Z., Hernandez X. I. P., deLaRiva A., Wang M., Engelhard M. H., Kovarik L., Datye A. K., Wang Y., Science, 2017, 358, 1419―1423 |
| 21 | Li X., Yang X., Huang Y., Zhang T., Liu B., Adv. Mater., 2019, 31, e1902031 |
| 22 | Xiao M., Gao L., Wang Y., Wang X., Zhu J., Jin Z., Liu C., Chen H., Li G., Ge J., He Q., Wu Z., Chen Z., Xing W., J. Am. Chem. Soc., 2019, 141, 19800―19806 |
| 23 | Chen P., Zhou T., Xing L., Xu K., Tong Y., Xie H., Zhang L., Yan W., Chu W., Wu C., Xie Y., Angew. Chem. Int. Ed., 2017, 56, 610―614 |
| 24 | Han A., Chen W., Zhang S., Zhang M., Han Y., Zhang J., Ji S., Zheng L., Wang Y., Gu L., Chen C., Peng Q., Wang D., Li Y., Adv. Mater., 2018, 30, e1706508 |
| 25 | Li X., Bi W., Chen M., Sun Y., Ju H., Yan W., Zhu J., Wu X., Chu W., Wu C., Xie Y., J. Am. Chem. Soc., 2017, 139, 14889―14892 |
| 26 | Zhang H., Wei J., Dong J., Liu G., Shi L., An P., Zhao G., Kong J., Wang X., Meng X., Zhang J., Ye J., Angew. Chem. Int. Ed., 2016, 55, 14310―14314 |
| 27 | Wang X., Chen W., Zhang L., Yao T., Liu W., Lin Y., Ju H., Dong J., Zheng L., Yan W., Zheng X., Li Z., Wang X., Yang J., He D., Wang Y., Deng Z., Wu Y., Li Y., J. Am. Chem. Soc., 2017, 139, 9419―9422 |
| 28 | Qi J. L., Xu Q. Q., Sun J. F., Zhou D., Yin J. Z., Process. Chem., 2020, 32, 505―518(祁建磊, 徐琴琴, 孙剑飞, 周丹, 银建中. 化学进展, 2020, 32, 505―518) |
| 29 | Jin Y. Y., Hao P. P., Ren J., Li Z., ProcessChem., 2015, 27, 1689―1704(靳永勇, 郝盼盼, 任军, 李忠. 化学进展, 2015, 27, 1689―1704) |
| 30 | Wang D., Wu H., Phua S. Z. F., Yang G., Qi L. W., Gu L., Qian C., Wang H., Guo Z., Chen H., Zhao Y., Nat. Commun., 2020, 11, 357 |
| 31 | Huo M., Wang L., Wang Y., Chen Y., Shi J., ACS Nano, 2019, 13, 2643―2653 |
| 32 | Wang L., Qu X., Zhao Y., Weng Y., Waterhouse G. I. N., Yan H., Guan S., Zhou S., Biosens. Bioelectron., 2019, 11, 35228―35237 |
| 33 | Gong N., Ma X., Ye X., Zhou Q., Chen X., Tan X., Yao S., Huo S., Zhang T., Chen S., Teng X., Hu X., Yu J., Gan Y., Jiang H., Li J., Liang X. J., Nat. Nanotechnol., 2019, 14, 379―387 |
| 34 | Huo M., Wang L., Zhang H., Zhang L., Chen Y., Shi J., Small, 2019, 15, e1901834 |
| 35 | Lu M., Wang C., Ding Y., Peng M., Zhang W., Li K., Wei W., Lin Y., Chem. Commun. (Camb), 2019, 55, 14534―14537 |
| 36 | Cao F., Zhang L., You Y., Zheng L., Ren J., Qu X., Angew. Chem. Int. Ed., 2020, 59, 5108―5115 |
| 37 | Yan R., Sun S., Yang J., Long W., Wang J., Mu X., Li Q., Hao W., Zhang S., Liu H., Gao Y., Ouyang L., Chen J., Liu S., Zhang X. D., Ming D., ACS Nano, 2019, 13, 11552―11560 |
| 38 | Niu X., Shi Q., Zhu W., Liu D., Tian H., Fu S., Cheng N., Li S., Smith J. N., Du D., Lin Y., Biosens. Bioelectron., 2019, 142, 111495 |
| 39 | Chen Q., Li S., Liu Y., Zhang X., Tang Y., Chai H., Huang Y., Sensor. Actuat. B: Chem., 2020, 305, 127511 |
| 40 | Jiao L., Xu W., Yan H., Wu Y., Liu C., Du D., Lin Y., Zhu C., Anal. Chem., 2019, 91, 11994―11999 |
| 41 | Cheng N., Li J. C., Liu D., Lin Y., Du D., Small, 2019, 15, e1901485 |
| 42 | Kim M. S., Lee J., Kim H. S., Cho A., Shim K. H., Le T. N., An S. S. A., Han J. W., Kim M. I., Lee J., Adv. Funct. Mater., 2019, 30, 1905410 |
| 43 | Chen M., Zhou H., Liu X., Yuan T., Wang W., Zhao C., Zhao Y., Zhou F., Wang X., Xue Z., Yao T., Xiong C., Wu Y., Small, 2020, e2002343 |
| 44 | Bray F., Ferlay J., Soerjomataram I., Siegel R. L., Torre L. A., Jemal A., CA Cancer J. Clin., 2018, 68, 394―424 |
| 45 | Yu H. Y., Tang C. H., Song W. T., Deng M. X., Chen X. S., Chem. J. Chinese Universities, 2014, 35(5), 903―916(于海洋, 汤朝晖, 宋万通, 邓明虓, 陈学思. 高等学校化学学报, 2014, 35(5), 903―916) |
| 46 | Shields C. W. T., Wang L. L., Evans M. A., Mitragotri S., Adv. Mater., 2020, 32, e1901633 |
| 47 | Marshall H. T., Djamgoz M. B. A., Front. Oncol., 2018, 8, 315 |
| 48 | Huang X. C., Deng J. S., Chen T. F., Zhang Y. B., Luo Y., Chem. J. Chinese Universities, 2012, 33(5), 976―982(黄晓纯, 郑军生, 陈填烽, 张逸波, 罗懿, 郑文杰. 高等学校化学学报, 2012, 33(5), 976―982) |
| 49 | Lin H., Chen Y., Shi J., Chem. Soc. Rev., 2018, 47, 1938―1958 |
| 50 | Yang B., Chen Y., Shi J., Adv. Mater., 2019, 31, e1901778 |
| 51 | Wang X. L., Wang Z. X., Chem. J. Chinese Universities, 2018, 39(10), 2185―2191(王雪丽, 王振新. 高等学校化学学报, 2018, 39(10), 2185―2191) |
| 52 | He J., Liu W. H., He L. M., Chen L. Y., Chem. J. Chinese Universities, 2013, 34(10), 2308―2312(何洁, 刘文洪, 何利民, 陈丽园. 高等学校化学学报, 2013, 34(10), 2308―2312) |
| 53 | Luo Z., Zhu H., Lin X. F., Hong Y., Xiao S. W., Zhang Q., Chu T. W., Yang Z., Chem. J. Chinese Universities, 2015, 36(1), 87―92(罗政, 朱华, 林新峰, 洪业, 肖绍文, 张强, 褚泰伟, 杨志. 高等学校化学学报, 2015, 36(1), 87―92) |
| 54 | Wang L., Huo M., Chen Y., Shi J., Adv. Healthc. Mater., 2018, 7, e1701156 |
| 55 | Willems P. H., Rossignol R., Dieteren C. E., Murphy M. P., Koopman W. J., Cell Metab., 2015, 22, 207―218 |
| 56 | Weinberg S. E., Chandel N. S., Nat. Chem. Biol., 2015, 11, 9―15 |
| 57 | Hu X., Xu Z., Hu J., Dong C., Lu Y., Wu X., Wumaier M., Yao T., Shi S., Inorg. Chem. Front., 2019, 6, 2865―2872 |
| 58 | Guan B., Yan R., Li R., Zhang X., Int. J. Nanomed., 2018, 13, 7473―7490 |
| 59 | He L., Nie T., Xia X., Liu T., Huang Y., Wang X., Chen T., Adv. Funct. Mater., 2019, 29, 1901240 |
| 60 | He L., Huang G., Liu H., Sang C., Liu X., Chen T., Sci. Adv., 2020, 6, eaay9751 |
| 61 | Liu T., Lai L., Song Z., Chen T., Adv. Funct. Mater., 2016, 26, 7775―7790 |
| 62 | Huang Y., He L., Liu W., Fan C., Zheng W., Wong Y. S., Chen T., Biomaterials, 2013, 34, 7106―7116 |
| 63 | Zheng L., Li C., Huang X., Lin X., Lin W., Yang F., Chen T., Biomaterials, 2019, 216, 119220 |
| 64 | Liu H., Lin W., He L., Chen T., Biomaterials, 2020, 226, 119545 |
| 65 | Chang Y., He L., Li Z., Zeng L., Song Z., Li P., Chan L., You Y., Yu X. F., Chu P. K., Chen T., ACS Nano, 2017, 11, 4848―4858 |
| 66 | Wang Y., Chen P., Zhao G., Sun K., Li D., Wan X., Zhang J., Food Chem. Toxicol., 2015, 85 71―77 |
| 67 | Levy S. B., Marshall B., Nat. Med., 2004, 10, S122―S129 |
| 68 | Bagchi D., Rathnam V. S. S., Lemmens P., Banerjee I., Pal S. K., ACS Omega, 2018, 3, 10877―10885 |
| 69 | Yin W., Yu J., Lv F., Yan L., Zheng L. R., Gu Z., Zhao Y., ACS Nano, 2016, 10, 11000―11011 |
| 70 | Lin Y., Ren J., Qu X., Acc. Chem. Res., 2014, 47, 1097―1105 |
| 71 | Shen X., Liu W., Gao X., Lu Z., Wu X., Gao X., J. Am. Chem. Soc., 2015, 137, 15882―15891 |
| 72 | Cao R., Thapa R., Kim H., Xu X., Gyu Kim M., Li Q., Park N., Liu M., Cho J., Nat. Commun., 2013, 4, 2076 |
| 73 | Liu Y., Shi J., Nano Today, 2019, 27, 146―177 |
| 74 | Andrades M. E., Morina A., Spasic S., Spasojevic I., Crit. Care., 2011, 15, 230 |
| 75 | Singh N., Savanur M. A., Srivastava S., D’Silva P., Mugesh G., Angew. Chem. Int. Ed., 2017, 56, 14267―14271 |
| 76 | Huang Y., Liu Z., Liu C., Ju E., Zhang Y., Ren J., Qu X., Angew. Chem. Int. Ed., 2016, 55, 6646―6650 |
| 77 | Russo M. V., McGavern D. B., Science, 2016, 353, 783―785 |
| 78 | Yang T., Cui Y. N., Chen H. Y., Li W. H., Acta Chem. Sinica, 2017, 75, 339―350(杨涛, 崔亚男, 陈怀银, 李伟华. 化学学报, 2017, 75, 339―350) |
| 79 | Lin S., Wei H., Sci. China Life Sci., 2019, 62, 710―712 |
| 80 | Li S., Liu X., Chai H., Huang Y., Trac. Trend. Anal. Chem., 2018, 105, 391―403 |
| 81 | Wang J., Li Z., Wu Y., Li Y., Adv. Mater., 2018, 30, e1801649 |
| [1] | 楚宇逸, 兰畅, 罗二桂, 刘长鹏, 葛君杰, 邢巍. 单原子铈对弱芬顿效应活性位点氧还原稳定性的提升[J]. 高等学校化学学报, 2022, 43(9): 20220294. |
| [2] | 杨静怡, 李庆贺, 乔波涛. 铱单原子和纳米粒子在N2O分解反应中的协同催化[J]. 高等学校化学学报, 2022, 43(9): 20220388. |
| [3] | 林高鑫, 王家成. 单原子掺杂二硫化钼析氢催化的进展和展望[J]. 高等学校化学学报, 2022, 43(9): 20220321. |
| [4] | 任诗杰, 谯思聪, 刘崇静, 张文华, 宋礼. 铂单原子催化剂同步辐射X射线吸收谱的研究进展[J]. 高等学校化学学报, 2022, 43(9): 20220466. |
| [5] | 滕镇远, 张启涛, 苏陈良. 聚合物单原子光催化剂的载流子分离和表面反应机制[J]. 高等学校化学学报, 2022, 43(9): 20220325. |
| [6] | 王茹玥, 魏呵呵, 黄凯, 伍晖. 单原子材料的冷冻合成[J]. 高等学校化学学报, 2022, 43(9): 20220428. |
| [7] | 秦永吉, 罗俊. 单原子催化剂在CO2转化中的应用[J]. 高等学校化学学报, 2022, 43(9): 20220300. |
| [8] | 江博文, 陈敬轩, 成永华, 桑微, 寇宗魁. 单原子材料在电化学生物传感中的研究进展[J]. 高等学校化学学报, 2022, 43(9): 20220334. |
| [9] | 林治, 彭志明, 贺韦清, 沈少华. 单原子与团簇光催化: 竞争与协同[J]. 高等学校化学学报, 2022, 43(9): 20220312. |
| [10] | 赵润瑶, 纪桂鹏, 刘志敏. 吡咯氮配位单原子铜催化剂的电催化二氧化碳还原性能[J]. 高等学校化学学报, 2022, 43(7): 20220272. |
| [11] | 王君旸, 刘争, 张茜, 孙春燕, 李红霞. DNA银纳米簇在功能核酸荧光生物传感器中的应用[J]. 高等学校化学学报, 2022, 43(6): 20220010. |
| [12] | 庄嘉豪, 王定胜. 单原子催化的关键进展与未来挑战[J]. 高等学校化学学报, 2022, 43(5): 20220043. |
| [13] | 张宏伟, 陈雯, 赵美淇, 马超, 韩云虎. 单原子催化剂在电化学中的研究进展[J]. 高等学校化学学报, 2022, 43(5): 20220129. |
| [14] | 尹肖菊, 孙逊, 赵程浩, 姜波, 赵晨阳, 张乃庆. 单原子催化剂在锂硫电池中的研究进展[J]. 高等学校化学学报, 2022, 43(5): 20220076. |
| [15] | 夏天, 万家炜, 于然波. 异原子配位结构碳基单原子电催化剂结构与性能相关性的研究进展[J]. 高等学校化学学报, 2022, 43(5): 20220162. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||