

高等学校化学学报 ›› 2023, Vol. 44 ›› Issue (3): 20220344.doi: 10.7503/cjcu20220344
收稿日期:2022-05-15
出版日期:2023-03-10
发布日期:2023-03-14
通讯作者:
汪铭
E-mail:mingwang@iccas.ac.cn
基金资助:
SHENG Jinhan, ZHENG Qizhen, WANG Ming( )
)
Received:2022-05-15
Online:2023-03-10
Published:2023-03-14
Contact:
WANG Ming
E-mail:mingwang@iccas.ac.cn
Supported by:摘要:
规律间隔成簇短回文重复序列及相关蛋白9(CRISPR/Cas9)系统的基因编辑技术为哺乳细胞基因组 的精准修饰与编辑研究提供了高效、 快捷的工具, 但其化学生物学应用依然面临着CRISPR基因编辑工具Cas9蛋白和gRNA的细胞及活体递送等问题. 近年来, 研究人员通过开发多种非病毒递送载体, 实现了编码CRISPR/Cas9基因编辑工具的DNA和信使RNA(mRNA)以及Cas9/gRNA核糖核蛋白(RNP)复合物的递送, 并应用于靶基因的化学修饰与编辑调控. 本文主要概述了近期CRISPR/Cas9基因编辑递送的研究进展, 并对其化学生物学应用前景进行了展望.
中图分类号:
TrendMD:
盛劲菡, 郑琪臻, 汪铭. CRISPR/Cas9基因编辑非病毒递送系统. 高等学校化学学报, 2023, 44(3): 20220344.
SHENG Jinhan, ZHENG Qizhen, WANG Ming. Non-viral Delivery of CRISPR/Cas9 Genome Editing. Chem. J. Chinese Universities, 2023, 44(3): 20220344.
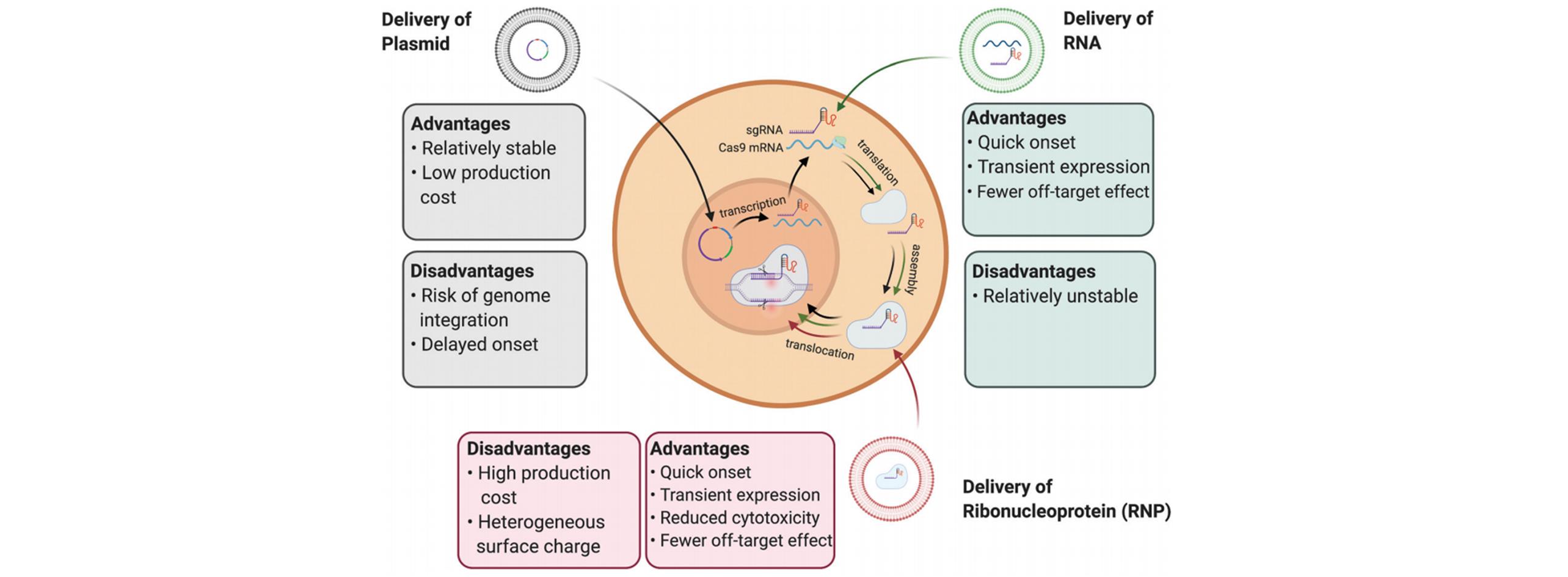
Fig.2 Intracellular delivery of CRISPR/Cas9 genome editing in the format of DNA plasmid, mRNA and RNPs[19]Copyright 2021, the Royal Society of Chemistry.
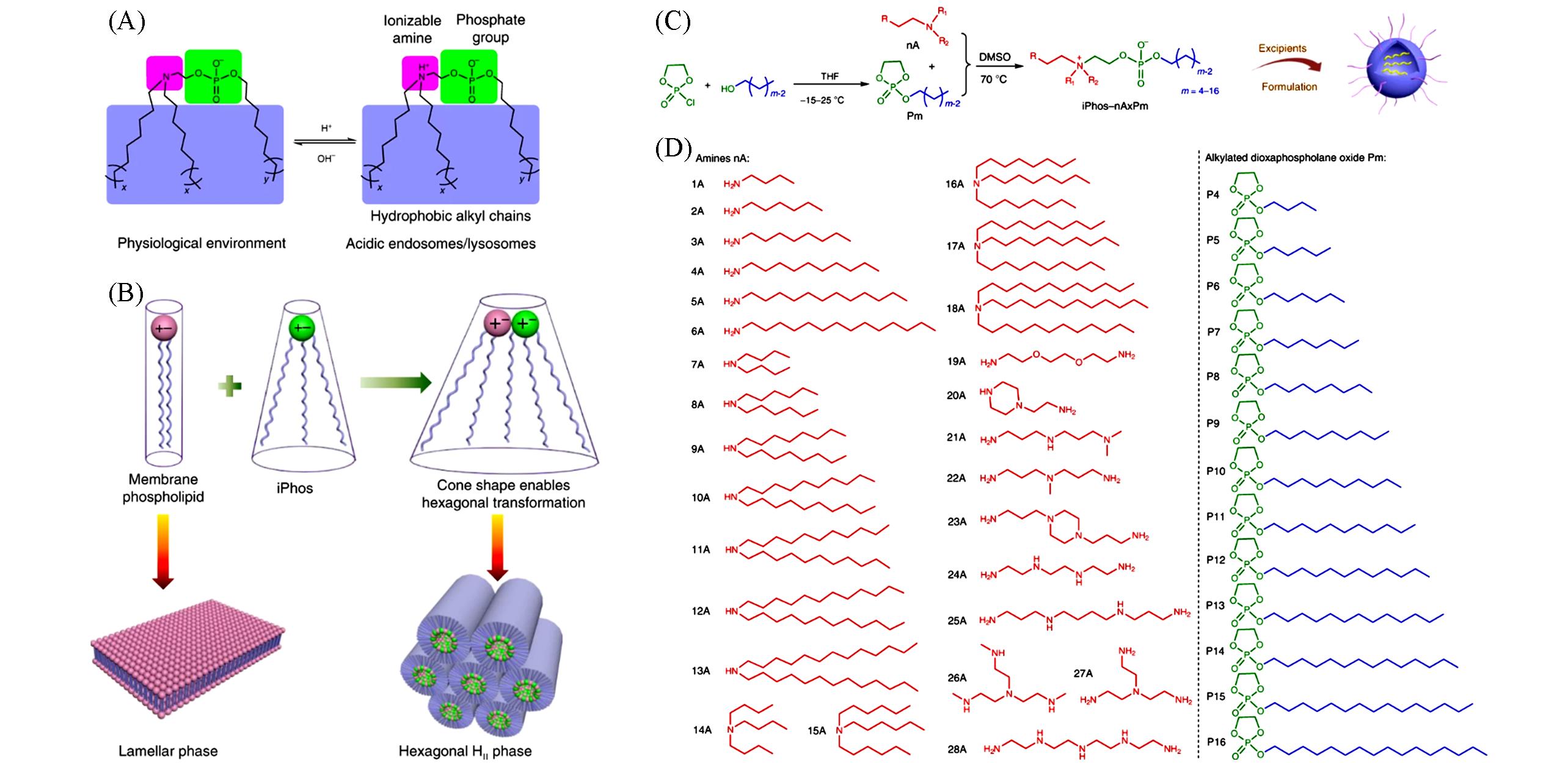
Fig.4 Membrane⁃destabilizing ionizable phospholipids for mRNA delivery[25](A) Efficacious iPhos lipids were composed of one ionizable amine, one phosphate group and three hydrophobic alkyl tails; (B) when iPhos lipids were mixed and inserted into the endosomal membranes, the formed cone shape by small ion pair head and multiple hydrophobic tails enabled hexagonal transformation; (C) synthetic routes of iPhos; (D) a list of 28 amines and 13 alkylated dioxaphospholane oxide molecules used for iPhos synthesis. Copyright 2021, Springer-Nature.
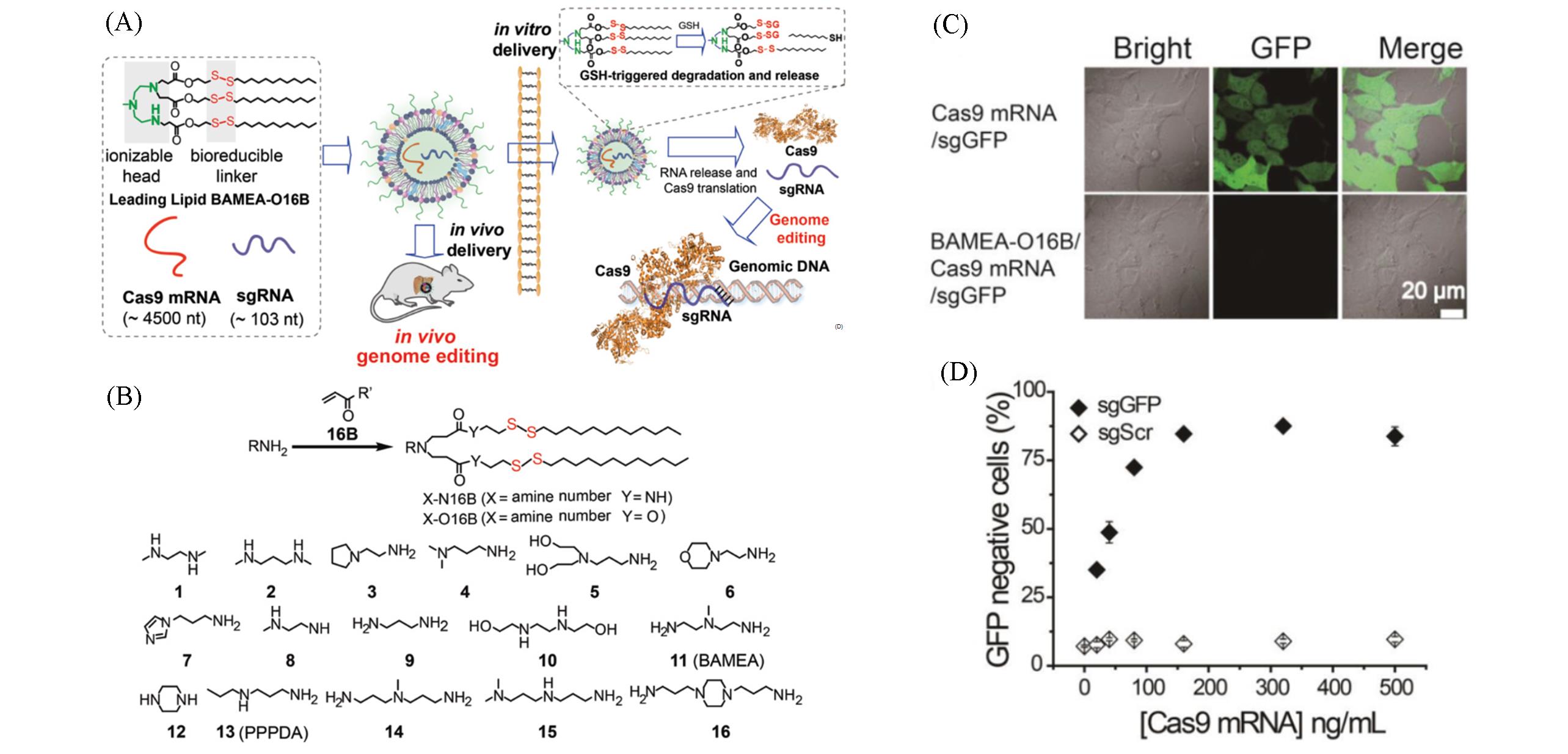
Fig.5 Bioreducible lipids BAMEA⁃O16B for mRNA delivery and genome editing[26](A) Illustration of formulating bioreducible lipid/Cas9 mRNA/sgRNA nanoparticle for CRISPR/Cas9 genome editing delivery in vitro and in vivo;(B) synthesis route, lipid nomenclature, and chemical structure of amines used for lipid synthesis; (C) CLSM images of HEK-GFP cells treated with Cas9 mRNA/sgGFP alone or BAMEA/Cas9 mRNA/sgGFP nanoparticles; (D) sgRNA sequence specific genome editing and GFP knockout using BAMEA/Cas9 mRNA/sgGFP nanoparticles. Copyright 2019, Wiley.
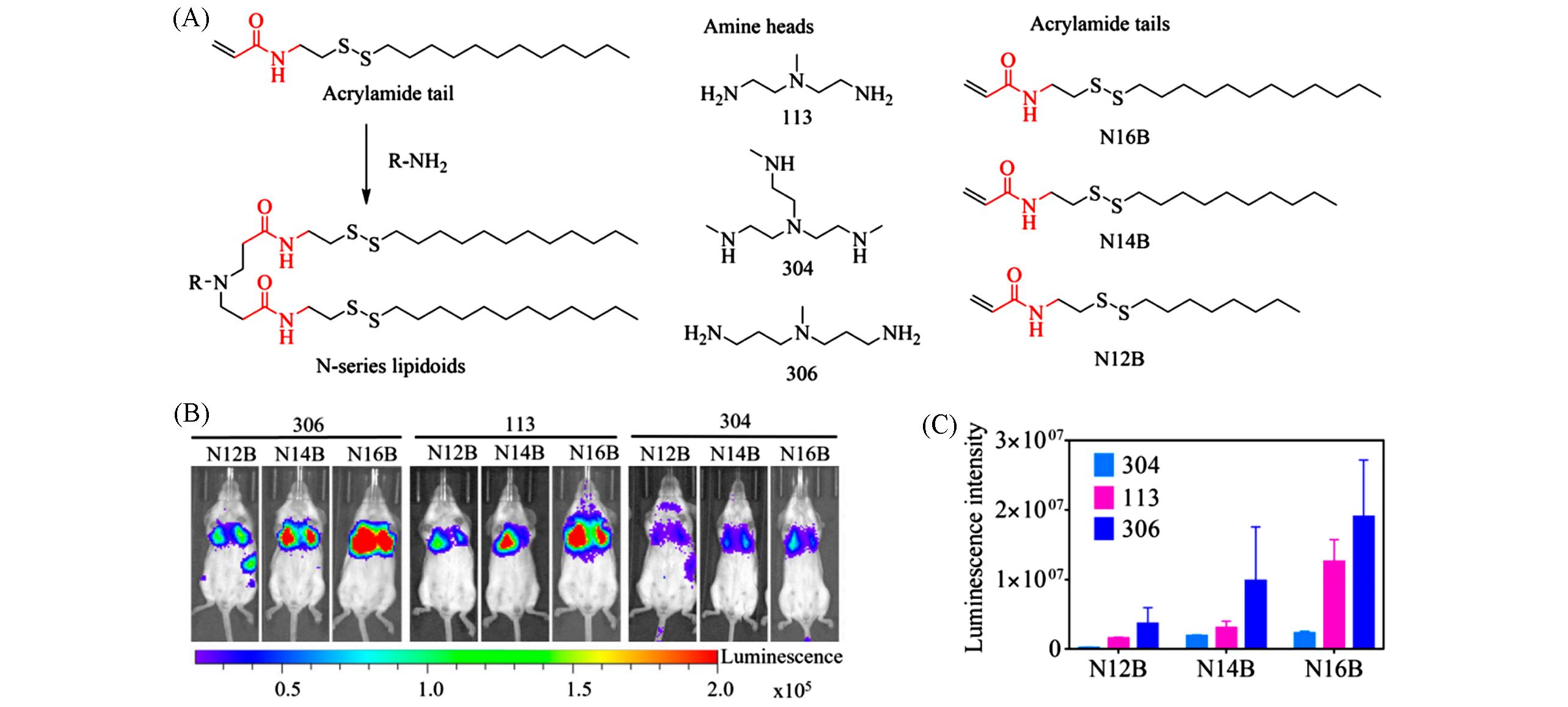
Fig.6 Synthesis and in vivo screening of N⁃series LNPs[29](A) Synthetic route and representative chemical structure of lipidoids; (B, C) representative whole body bioluminescence images of mice(B) and in vivo mRNA delivery efficacy(C) of N-series LNPs measured by the IVIS imaging system. Copyright 2022, Wiley.
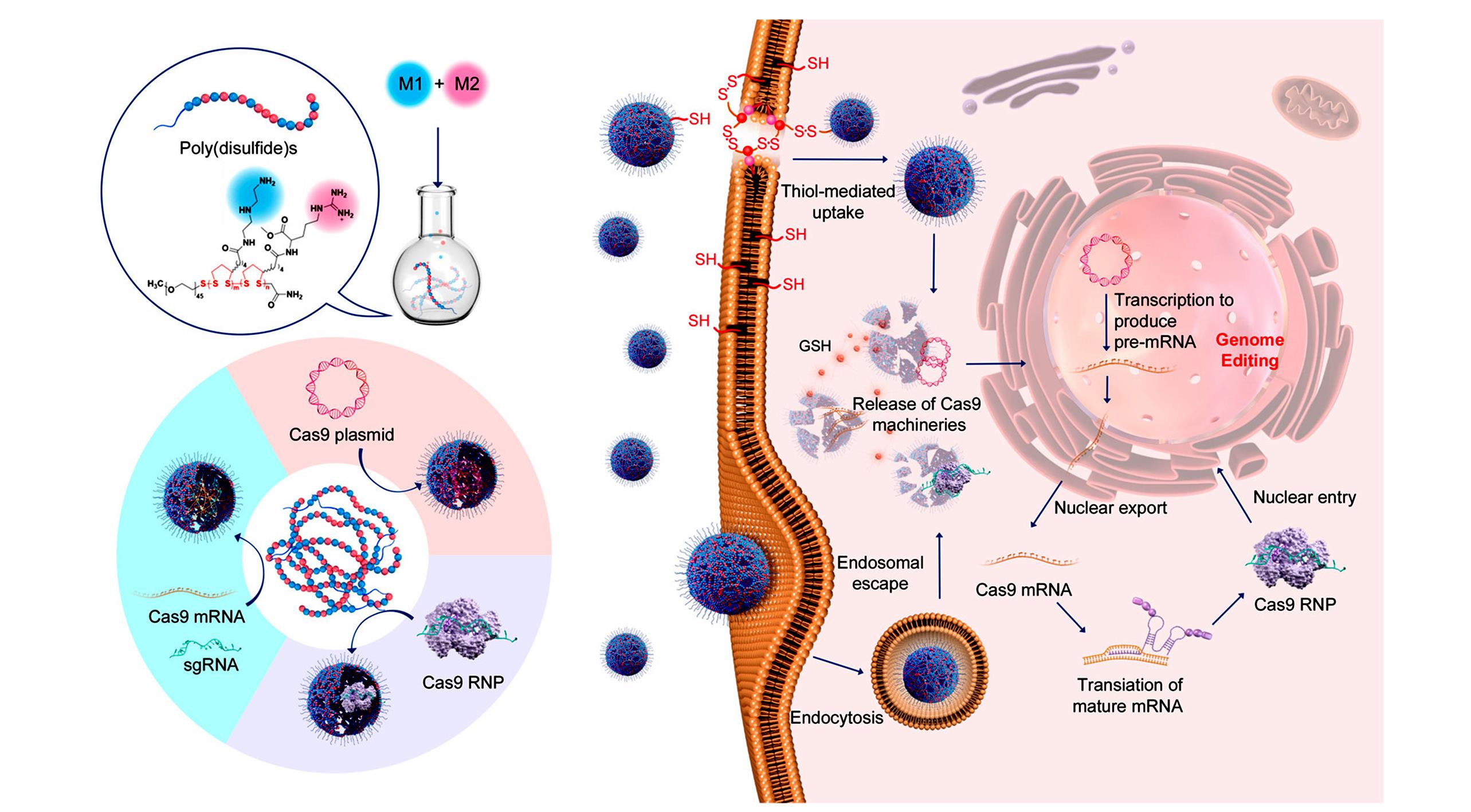
Fig.7 Polydisulfides as CRISPR mRNA/sgRNA delivery vector[34]Schematic illustration of the preparation of poly(disulfide)s, the complexation of genome-editing biomacromolecules(Cas9 plasmid, Cas9 mRNA, and Cas9 ribonucleoprotein) by poly(disulfide)s, and their intracellular delivery processes for genome editing. Copyright 2019, American Chemical Society.
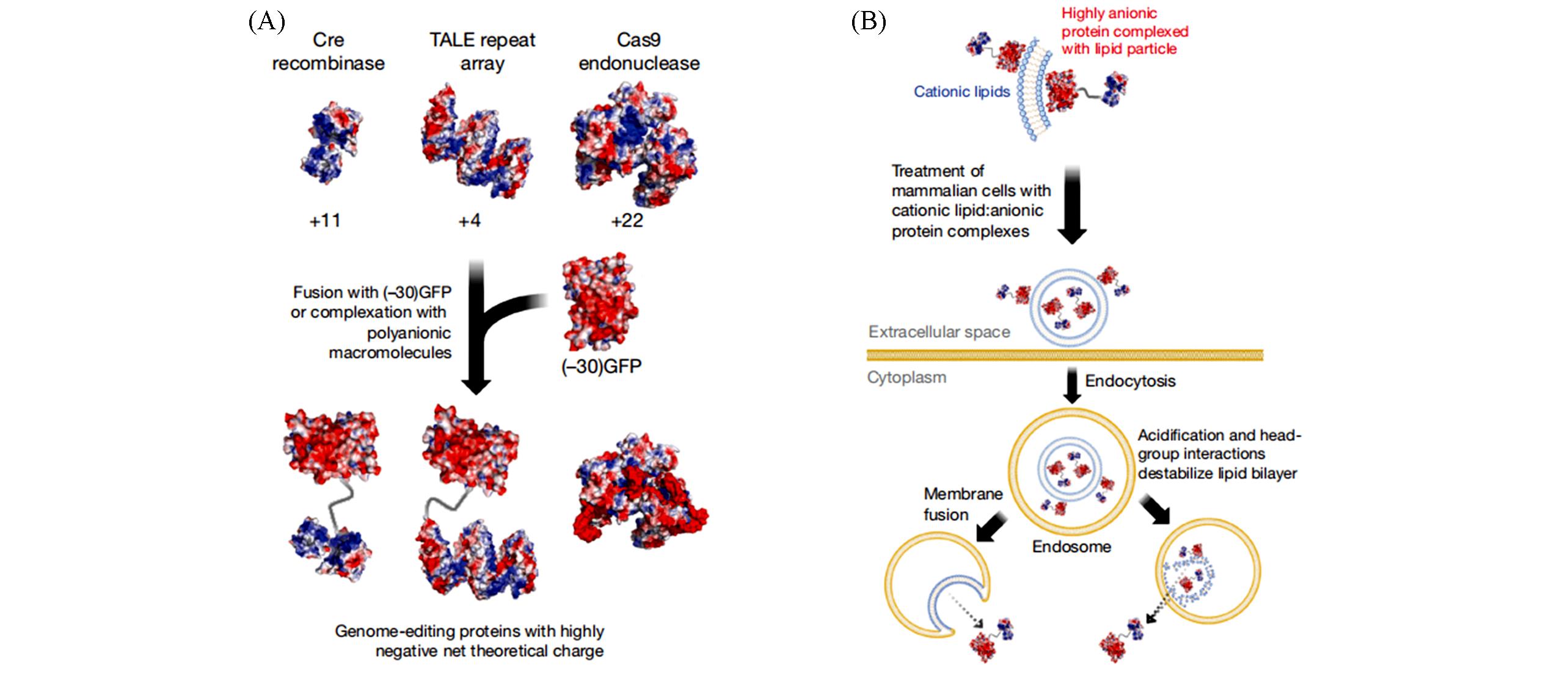
Fig.8 Strategy for delivering proteins into mammalian cells by self⁃assembling supercharged protein and cationic lipids[36]Copyright 2014, Springer⁃Nature.
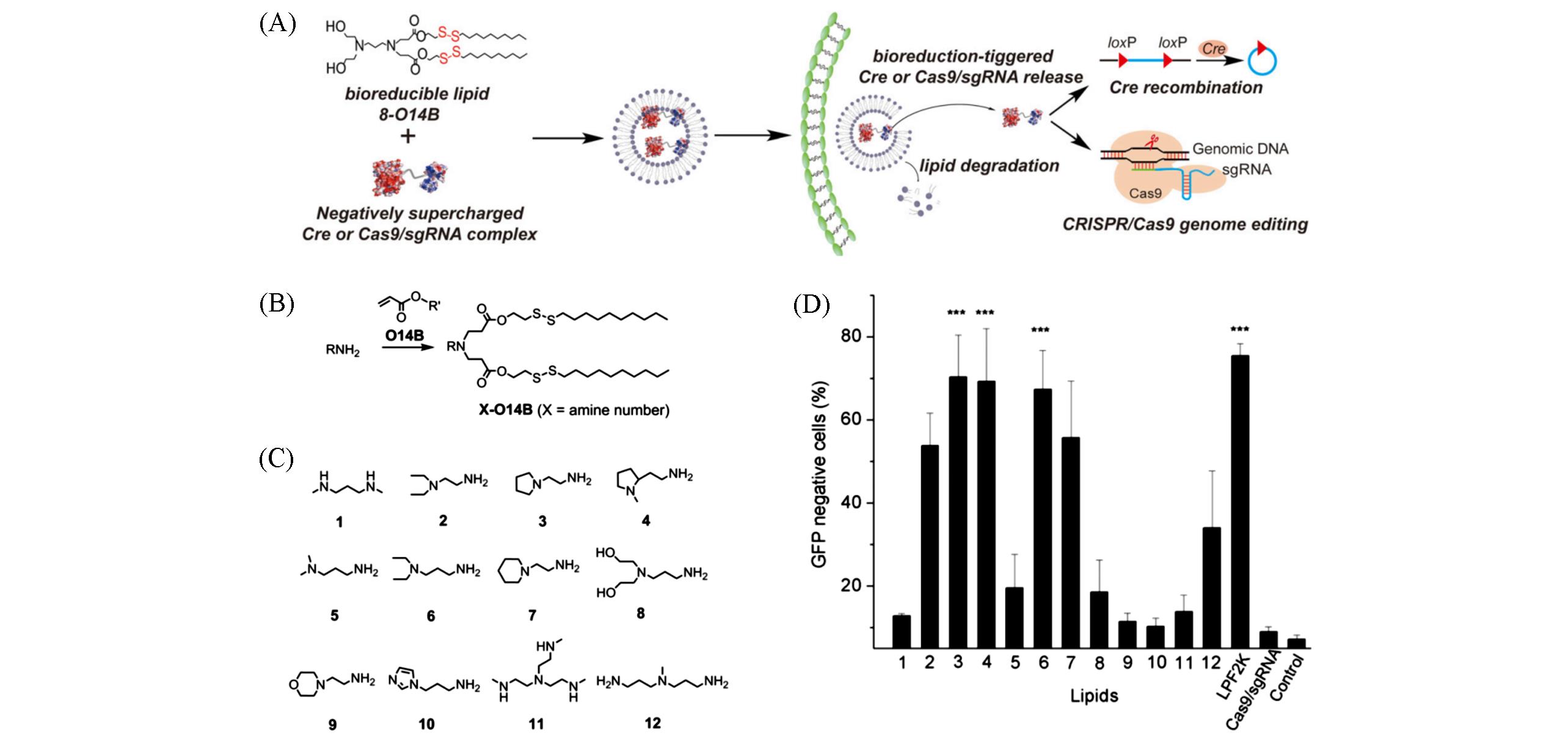
Fig.9 Bioreducible lipid⁃like materials for genome editing protein delivery[37](A) Design of bioreducible lipid-like materials and negatively supercharged protein for effective protein delivery and genome editing; (B) synthesis route and lipid nomenclature; (C) chemical structures of amines used as head groups for lipid synthesis; (D) delivery of Cas9/sgRNA complex into cultured human cells for genome editing.Copyright 2016, Wiley.
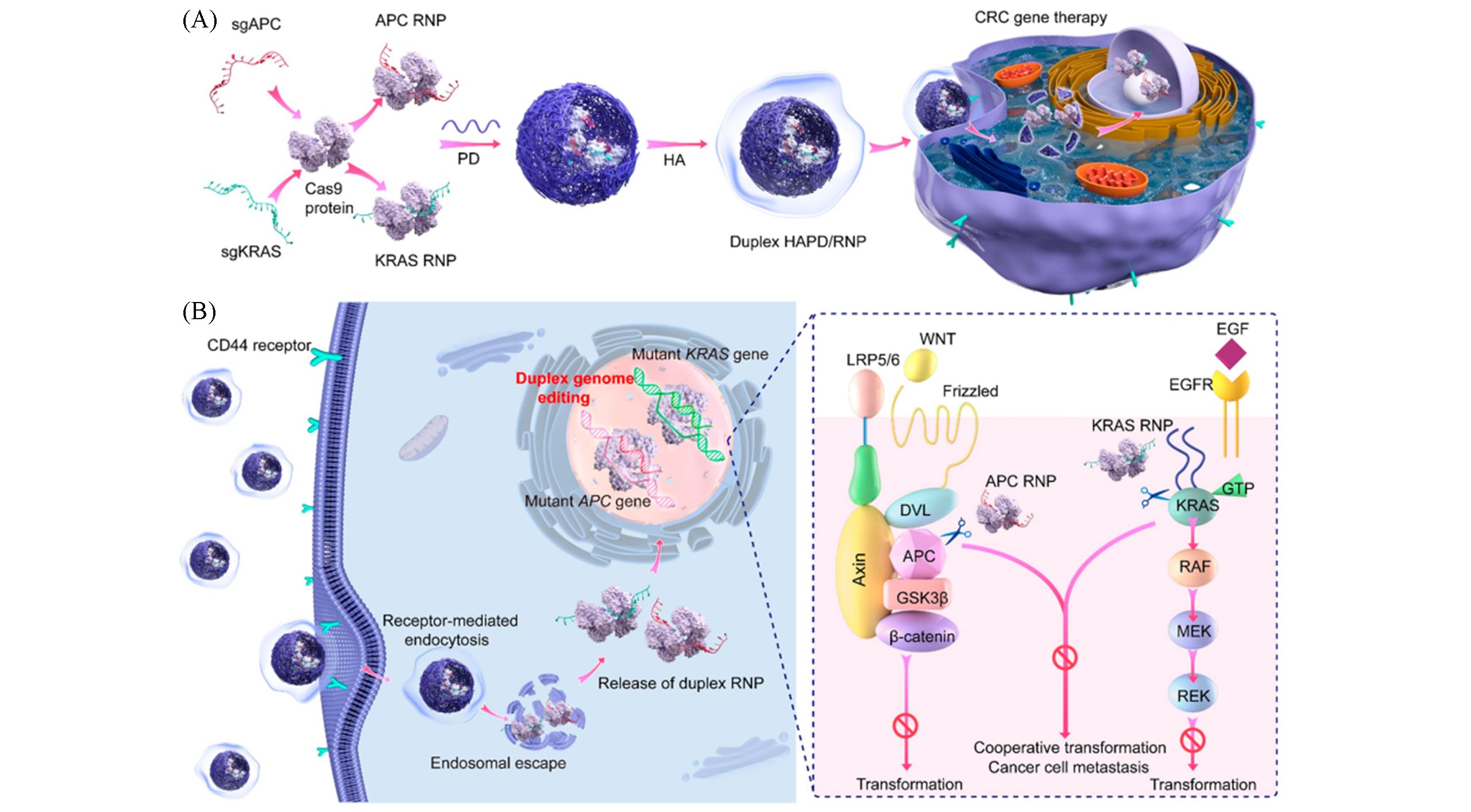
Fig.10 A duplex HAPD/CRISPR⁃Cas9 ribonucleoprotein nanomedicine[40]Schematic illustration of the preparation of duplex HAPD/RNP nanoformulationsa(A) and duplex genome editing of APC and KRAS mutations mediated by HAPD-mediated targeted delivery of RNPs through activated WNT/β-Catenin and RAS-ERK signaling pathways for the treatment of CRC(B). Copyright 2021, American Chemical Society.
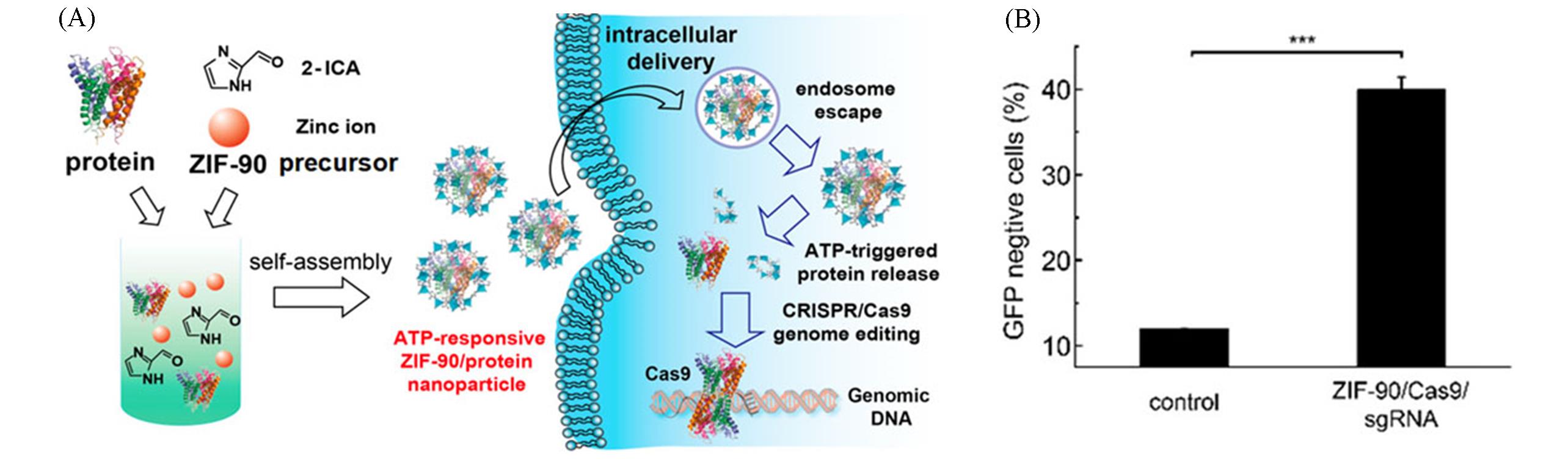
Fig.11 Nanoscale ATP⁃responsive zeolitic imidazole framework⁃90 for genome editing delivery[43](A) Schematic illustration of the self-assembly of ZIF-90/protein nanoparticle and ATP-triggered protein release from ZIF-90 nanoparticle inside cells; (B) the delivery of ZIF-90/Cas9 nanoparticles efficiently knocked out GFP expression of HeLa-GFP cells.Copyright 2021, American Chemical Society.
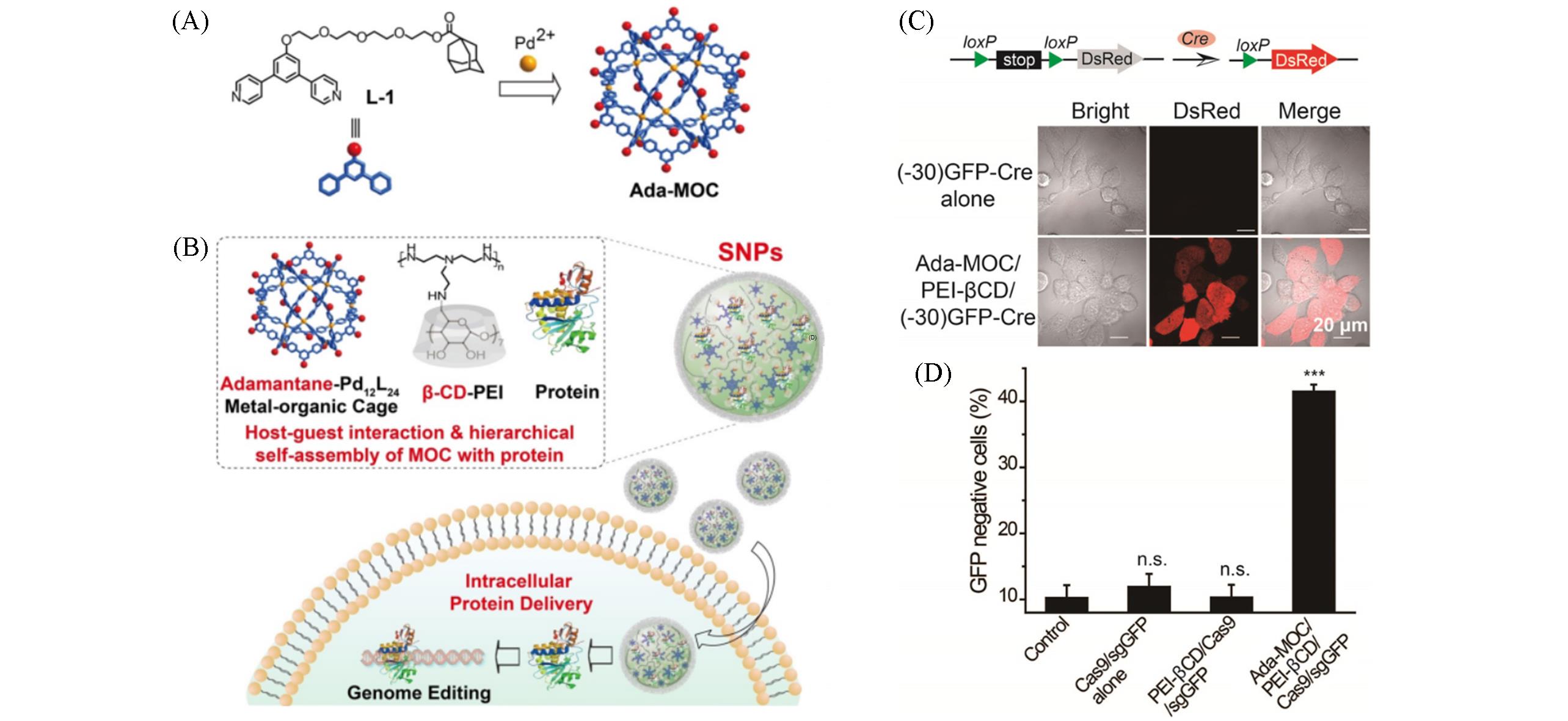
Fig.12 Hierarchical self⁃assembly discrete metal⁃organic cages into supramolecular nanoparticles for genome editing delivery[44](A, B) Illustration of the self-assembly of adamantane-functionalized M12L24 MOC and β-cyclodextrin-conjugated polyethylenimine(PEI-βCD) along with proteins into supramolecular nanoparticles for intracellular protein delivery; (C) representation shows delivery of Cre deletes the stop cassette and activates downstream DsRed protein; (D) HEK-GFP cells treated with Cas9/sgGFP alone, PEI-βCD/Cas9/sgGFP, Ada-MOC/PEI-βCD/Cas9/sgGFP nanoparticles(8 μg/ mL Cas9 and 2.5 μg/ mL sgGFP). Copyright 2021, Wiley.
| 1 | Pickar⁃Oliver A., Gersbach C. A., Nat. Rev. Mol. Cell. Biol., 2019, 20(8), 490—507 |
| 2 | Anzalone A. V., Koblan L. W., Liu D. R., Nat. Biotechnol., 2020, 38(7), 824—844 |
| 3 | Chen J. S., Ma E., Harrington L. B., Da Costa M., Tian X., Palefsky J. M., Doudna J. A., Science, 2018, 360(6387), 436—439 |
| 4 | Collias D., Beisel C. L., Nat. Commun., 2021, 12(1), 555 |
| 5 | Ran F. A., Hsu P. D., Wright J., Agarwala V., Scott D. A., Zhang F. Nat. Protoc., 2013, 8(11), 2281—2308 |
| 6 | Gaudelli N. M., Komor A. C., Rees H.A., Packer M. S., Badran A. H., Bryson D. I., Liu D. R., Nature, 2017, 551(7681), 464—471 |
| 7 | Essletzbichler P., Konopka T., Santoro F., Chen D., Gapp B. V., Kralovics R., Brummelkamp T. R., Nijman S. M., Burckstummer T., Genome Res., 2014, 24(12), 2059—2065 |
| 8 | Kraft K., Geuer S., Will A. J., Chan W. L., Paliou C., Borschiwer M., Harabula I., Wittler L., Franke M., Ibrahim D. M., Kragesteen B. K., Spielmann M., Mundlos S., Lupianez D. G., Andrey G., Cell Rep., 2015, 10(5), 833—839 |
| 9 | Xiao A., Wang Z., Hu Y., Wu Y., Luo Z., Yang Z., Zu Y., Li W., Huang P., Tong X., Zhu Z., Lin S., Zhang B., Nucleic Acids Res., 2013, 41(14), e141 |
| 10 | Suzuki K., Tsunekawa Y., Hernandez⁃Benitez R., Wu J., Zhu J., Kim E. J., Hatanaka F., Yamamoto M., Araoka T., Li Z., Kurita M., Hishida T., Li M., Aizawa E., Guo S., Chen S., Goebl A., Soligalla R. D., Qu J., Jiang T., Fu X., Jafari M., Esteban C. R., Berggren W. T., Lajara J., Nunez⁃Delicado E., Guillen P., Campistol J. M., Matsuzaki F., Liu G. H., Magistretti P., Zhang K., Callaway E. M., Zhang K., Belmonte J. C., Nature, 2016, 540(7631), 144—149 |
| 11 | Quadros R. M., Miura H., Harms D. W., Akatsuka H., Sato T., Aida T., Redder R., Richardson G. P., Inagaki Y., Sakai D., Buckley S. M., Seshacharyulu P., Batra S. K., Behlke M. A., Zeiner S. A., Jacobi A. M., Izu Y., Thoreson W. B., Urness L. D., Mansour S. L., Ohtsuka M., Gurumurthy C. B., Genome Biol., 2017, 18(1), 92 |
| 12 | Komor A. C., Kim Y. B., Packer M. S., Zuris J. A., Liu D. R., Nature, 2016, 533(7603), 420—424 |
| 13 | Thakore P. I., D'Ippolito A. M., Song L., Safi A., Shivakumar N. K., Kabadi A. M., Reddy T. E., Crawford G. E., Gersbach C. A., Nat. Methods, 2015, 12(12), 1143—1149 |
| 14 | Balboa D., Weltner J., Eurola S., Trokovic R., Wartiovaara K., Otonkoski T., Stem Cell Reports, 2015, 5(3), 448—459 |
| 15 | Jiao C., Sharma S., Dugar G., Peeck N. L., Bischler T., Wimmer F., Yu Y., Barquist L., Schoen C., Kurzai O., Sharma C. M., Beisel C. L., Science, 2021, 372(6545), 941—948 |
| 16 | Zhang Y., Qian L., Wei W., Wang Y., Wang B., Lin P., Liu W., Xu L., Li X., Liu D., Cheng S., Li J., Ye Y., Li H., Zhang X., Dong Y., Zhao X., Liu C., Zhang H. M., Ouyang Q., Lou C., ACS Synth. Biol., 2017, 6(2), 211—216 |
| 17 | Wang X. W., Hu L. F., Hao J., Liao L. Q., Chiu Y. T., Shi M., Wang Y., Nat. Cell Biol., 2019, 21(4), 522—530 |
| 18 | George J. T., Azhar M., Aich M., Sinha D., Ambi U. B., Maiti S., Chakraborty D., Srivatsan S. G., J. Am. Chem. Soc., 2020, 142(32), 13954—13965 |
| 19 | Yan J., Kang D. D., Dong Y., Biomater. Sci., 2021, 9(18), 6001—6011 |
| 20 | Cai W., Luo T., Mao L., Wang M., Angew Chem. Int. Ed., 2021, 60(16), 8596—8606 |
| 21 | Miller J. B., Zhang S., Kos P., Xiong H., Zhou K., Perelman S. S., Zhu H., Siegwart D. J., Angew. Chem. Int. Ed., 2017, 56(4), 1059—1063 |
| 22 | Lin Y., Wagner E., Lachelt U., Biomater. Sci., 2022, 10(5), 1166—1192 |
| 23 | Love K. T., Mahon K. P., Levins C. G., Whitehead K. A., Querbes W., Dorkin J. R., Qin J., Cantley W., Qin L. L., Racie T., Frank⁃Kamenetsky M., Yip K. N., Alvarez R., Sah D. W., de Fougerolles A., Fitzgerald K., Koteliansky V., Akinc A., Langer R., Anderson D. G., Proc. Natl. Acad. Sci. USA, 2010, 107(5), 1864—1869 |
| 24 | Yin H., Song C. Q., Dorkin J. R., Zhu L. J., Li Y., Wu Q., Park A., Yang J., Suresh S., Bizhanova A., Gupta A., Bolukbasi M. F., Walsh S., Bogorad R. L., Gao G., Weng Z., Dong Y., Koteliansky V., Wolfe S. A., Langer R., Xue W., Anderson D. G., Nat. Biotechnol., 2016, 34(3), 328—333 |
| 25 | Liu S., Cheng Q., Wei T., Yu X., Johnson L. T., Farbiak L., Siegwart D. J., Nat. Mater., 2021, 20(5), 701—710 |
| 26 | Liu J., Chang J., Jiang Y., Meng X., Sun T., Mao L., Xu Q., Wang M., Adv. Mater., 2019, 31(33), e1902575 |
| 27 | Tang Q., Liu J., Jiang Y., Zhang M., Mao L., Wang M., ACS Appl. Mater. Interfaces, 2019, 11(50), 46585—46590 |
| 28 | Qiu M., Glass Z., Chen J., Haas M., Jin X., Zhao X., Rui X., Ye Z., Li Y., Zhang F., Xu Q., Proc. Natl. Acad. Sci. USA, 2021, 118(10), e2020401118 |
| 29 | Qiu M., Tang Y., Chen J., Muriph R., Ye Z., Huang C., Evans J., Henske E. P., Xu Q., Proc. Natl. Acad. Sci. USA, 2022, 119(8), e2116271119 |
| 30 | Gillmore J. D., Gane E., Taubel J., Kao J., Fontana M., Maitland M. L., Seitzer J., O'Connell D., Walsh K. R., Wood K., Phillips J., Xu Y., Amaral A., Boyd A. P., Cehelsky J. E., McKee M. D., Schiermeier A., Harari O., Murphy A., Kyratsous C. A., Zambrowicz B., Soltys R., Gutstein D. E., Leonard J., Sepp⁃Lorenzino L., Lebwohl D., N. Engl. J. Med., 2021, 385(6), 493—502 |
| 31 | Rosenblum D., Gutkin A., Kedmi R., Ramishetti S., Veiga N., Jacobi A. M., Schubert M. S., Friedmann-Morvinski D., Cohen Z. R., Behlke M. A., Lieberman J., Peer D., Sci. Adv., 2020, 6(47), eabc9450 |
| 32 | Cheng Q., Wei T., Farbiak L., Johnson L.T., Dilliard S. A., Siegwart D. J., Nat. Nanotechnol., 2020, 15(4), 313—320 |
| 33 | Finn J. D., Smith A. R., Patel M. C., Shaw L., Youniss M. R., van Heteren J., Dirstine T., Ciullo C., Lescarbeau R., Seitzer J., Shah R. R., Shah A., Ling D., Growe J., Pink M., Rohde E., Wood K. M., Salomon W. E., Harrington W. F., Dombrowski C., Strapps W. R., Chang Y., Morrissey D. V., Cell Rep., 2018, 22(9), 2227—2235 |
| 34 | Guo J., Wan T., Li B., Pan Q., Xin H., Qiu Y., Ping Y., ACS Cent. Sci., 2021, 7(6), 990—1000 |
| 35 | Radis⁃Baptista G., Campelo I. S., Morlighem J. R. L., Melo L. M., Freitas V. J. F., J. Biotechnol., 2017, 252, 15—26 |
| 36 | Zuris J. A., Thompson D. B., Shu Y., Guilinger J. P., Bessen J. L., Hu J. H., Maeder M. L., Joung J. K., Chen Z. Y., Liu D. R., Nat. Biotechnol., 2015, 33(1), 73—80 |
| 37 | Wang M., Zuris J. A., Meng F., Rees H., Sun S., Deng P., Han Y., Gao X., Pouli D., Wu Q., Georgakoudi I., Liu D. R., Xu Q., Proc. Natl. Acad. Sci. USA, 2016, 113(11), 2868—2873 |
| 38 | Wei T., Cheng Q., Min Y. L., Olson E. N., Siegwart D. J., Nat. Commun., 2020, 11(1), 3232 |
| 39 | Liu C., Wan T., Wang H., Zhang S., Ping Y., Cheng Y., Sci. Adv., 2019, 5(6), eaaw8922 |
| 40 | Wan T., Pan Q., Liu C., Guo J., Li B., Yan X., Cheng Y., Ping Y., Nano Lett., 2021, 21(22), 9761—9771 |
| 41 | Alsaiari S. K., Patil S., Alyami M., Alamoudi K. O., Aleisa F. A., Merzaban J. S., Li M., Khashab N. M., J. Am. Chem. Soc., 2018, 140(1), 143—146 |
| 42 | Alyami M. Z., Alsaiari S. K., Li Y., Qutub S. S., Aleisa F. A., Sougrat R., Merzaban J. S., Khashab N. M., J. Am. Chem. Soc., 2020, 142(4), 1715—1720 |
| 43 | Yang X., Tang Q., Jiang Y., Zhang M., Wang M., Mao L., J. Am. Chem. Soc., 2019, 141(9), 3782—3786 |
| 44 | Liu J., Luo T., Xue Y., Mao L., Stang P. J., Wang M., Angew. Chem. Int. Ed., 2021, 60(10), 5429—5435 |
| 45 | Wan T., Chen Y., Pan Q., Xu X., Kang Y., Gao X., Huang F., Wu C., Ping Y., J. Control Release, 2020, 322, 236—247 |
| 46 | Sun W., Wang J., Hu Q., Zhou X., Khademhosseini A., Gu Z., Sci. Adv., 2020, 6(21), eaba2983 |
| 47 | Li F., Song N., Dong Y., Li S., Li L., Liu Y., Li Z., Yang D., Angew. Chem. Int. Ed., 2022, 61(9), e202116569 |
| 48 | Wu J., Peng H., Lu X., Lai M., Zhang H., Le X. C., Angew Chem. Int. Ed., 2021, 60(20), 11104—11109 |
| 49 | Hansen⁃Bruhn M., de Avila B. E., Beltran⁃Gastelum M., Zhao J., Ramirez⁃Herrera D. E., Angsantikul P., Vesterager Gothelf K., Zhang L., Wang J., Angew. Chem. Int. Ed., 2018, 57(10), 2657—2661 |
| 50 | Zhang L., Wang L., Xie Y., Wang P., Deng S., Qin A., Zhang J., Yu X., Zheng W., Jiang X., Angew. Chem. Int. Ed., 2019, 58(36), 12404—12408 |
| 51 | Zhai Y., Wang J., Lang T., Kong Y., Rong R., Cai Y., Ran W., Xiong F., Zheng C., Wang Y., Yu Y., Zhu H. H., Zhang P., Li Y., Nat. Nanotechnol., 2021, 16(11), 1271—1280 |
| [1] | 常丽颖, 凌鑫宇, 陈和祺, 王雪, 刘涛. 基因编辑在线粒体疾病中的应用[J]. 高等学校化学学报, 2023, 44(3): 20220363. |
| [2] | 孔好, 徐菲洋, 王依香, 张艳. 基于小分子化学反应工具构建CRISPR-Cas9功能调控体系的研究进展[J]. 高等学校化学学报, 2023, 44(3): 20220346. |
| [3] | 肖珩, 李永奎, 邢曦雯. 化学调控CRISPR/Cas9基因编辑技术的研究进展[J]. 高等学校化学学报, 2023, 44(3): 20220410. |
| [4] | 胡玉灿, 曹朝辉, 郑灵刚, 沈俊涛, 赵维, 戴磊. CRISPR⁃Cas基因编辑技术在微生物组工程中的应用[J]. 高等学校化学学报, 2023, 44(3): 20220362. |
| [5] | 王慧, 赵德偲, 杨乃亮, 王丹. 智能中空药物载体的门控设计[J]. 高等学校化学学报, 2023, 44(1): 20220237. |
| [6] | 杨霁野, 孙大吟, 王妍, 谷安祺, 叶一兰, 丁书江, 杨振忠. 若干典型中空结构材料的模板合成与应用进展[J]. 高等学校化学学报, 2023, 44(1): 20220665. |
| [7] | 仵宇帅, 尚颖旭, 蒋乔, 丁宝全. 可控自组装DNA折纸结构作为药物载体的研究进展[J]. 高等学校化学学报, 2022, 43(8): 20220179. |
| [8] | 孙雪峰, 热娜古丽·阿不都热合曼, 杨通胜, 杨倩婷. Cr,In共掺杂MgGa2O4小尺寸近红外长余辉纳米颗粒的制备及发光性能[J]. 高等学校化学学报, 2022, 43(4): 20210850. |
| [9] | 储彬彬, 何耀. 硅基纳米探针用于眼部疾病的成像检测与治疗[J]. 高等学校化学学报, 2022, 43(12): 20220546. |
| [10] | 汪诗琪, 罗博文, 俞计成, 顾臻. 近红外二区活体荧光成像在肿瘤诊疗中的应用[J]. 高等学校化学学报, 2022, 43(12): 20220577. |
| [11] | 赵恒智, 余方志, 李翔菲, 李乐乐. 基于DNA与上转换纳米颗粒相结合的生物传感与成像研究进展[J]. 高等学校化学学报, 2022, 43(12): 20220626. |
| [12] | 杨兆华, 成鸿静, 杨弋, 刘辉, 杜飞鹏, 张云飞. 聚乙烯醇载银海绵的制备及界面光热驱动水蒸发性能[J]. 高等学校化学学报, 2022, 43(10): 20220181. |
| [13] | 刘志刚, 李家宝, 杨剑, 马浩, 王赪胤, 郭鑫, 汪国秀. 新型石墨化氮化碳/锡/氮掺杂碳复合物的制备及储钠性能[J]. 高等学校化学学报, 2021, 42(2): 633. |
| [14] | 李梅, 夏晓娟, 陈志雄, 杨梦, 李紫滢, 杨通, 孟爽, 杨云慧, 胡蓉. 基于铂纳米颗粒@金属有机骨架纳米模拟酶的无标记电化学赭曲霉毒素适体传感器的构建[J]. 高等学校化学学报, 2021, 42(12): 3615. |
| [15] | 刘红, 江敬红, 段志娟, 徐仕军, 黄福建, 夏帆. 光控CRISPR技术的研究进展[J]. 高等学校化学学报, 2021, 42(11): 3321. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||