

高等学校化学学报 ›› 2021, Vol. 42 ›› Issue (11): 3321.doi: 10.7503/cjcu20210420
刘红1, 江敬红2, 段志娟1, 徐仕军1, 黄福建1( ), 夏帆1
), 夏帆1
收稿日期:2021-06-21
出版日期:2021-11-10
发布日期:2021-08-18
通讯作者:
黄福建
E-mail:huangfj@cug.edu.cn
基金资助:
LIU Hong1, JIANG Jinghong2, DUAN Zhijuan1, XU Shijun1, HUANG Fujian1( ), XIA Fan1
), XIA Fan1
Received:2021-06-21
Online:2021-11-10
Published:2021-08-18
Contact:
HUANG Fujian
E-mail:huangfj@cug.edu.cn
Supported by:摘要:
成簇规则间隔短回文重复序列和成簇规则间隔短回文重复序列相关(CRISPR-Cas)系统提供了用于可编程基因组编辑的多功能工具. CRISPR与光遗传学及光化学生物学技术的结合产生了很多新的成果. 光激活的CRISPR-Cas系统能够在空间和时间上更好地调控RNA引导的核酸酶的活性. 近年来, 科学家结合CRISPR和多种光学技术, 开发出了一系列光激活的CRISPR工具. 这些工具让研究人员能够在空间、 时间和基因组坐标上进行高分辨率的生命活动研究. 本文概述了CRISPR系统、 基因编辑技术、 光遗传学和光化学生物学的研究进展, 并对光诱导的CRISPR技术的发展进行了展望.
中图分类号:
TrendMD:
刘红, 江敬红, 段志娟, 徐仕军, 黄福建, 夏帆. 光控CRISPR技术的研究进展. 高等学校化学学报, 2021, 42(11): 3321.
LIU Hong, JIANG Jinghong, DUAN Zhijuan, XU Shijun, HUANG Fujian, XIA Fan. Recent Advance in Light-controlled CRISPR Technology. Chem. J. Chinese Universities, 2021, 42(11): 3321.
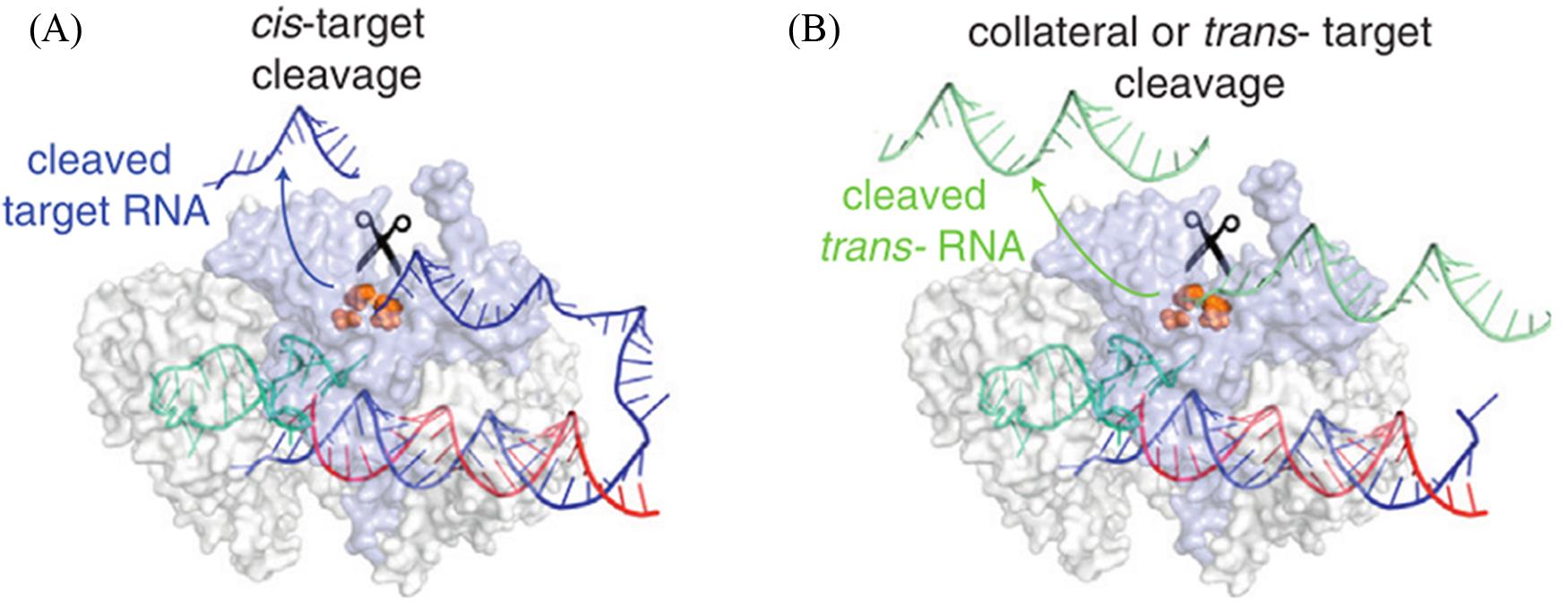
Fig.4 Molecular mechanisms of RNA?targeting by Cas13[42]A schematic highlights that Cas 13's HEPN domains once activated can cleave target?RNAs in both cis?(A) and trans?cleavage (B). Copyright 2018, Elsevier.
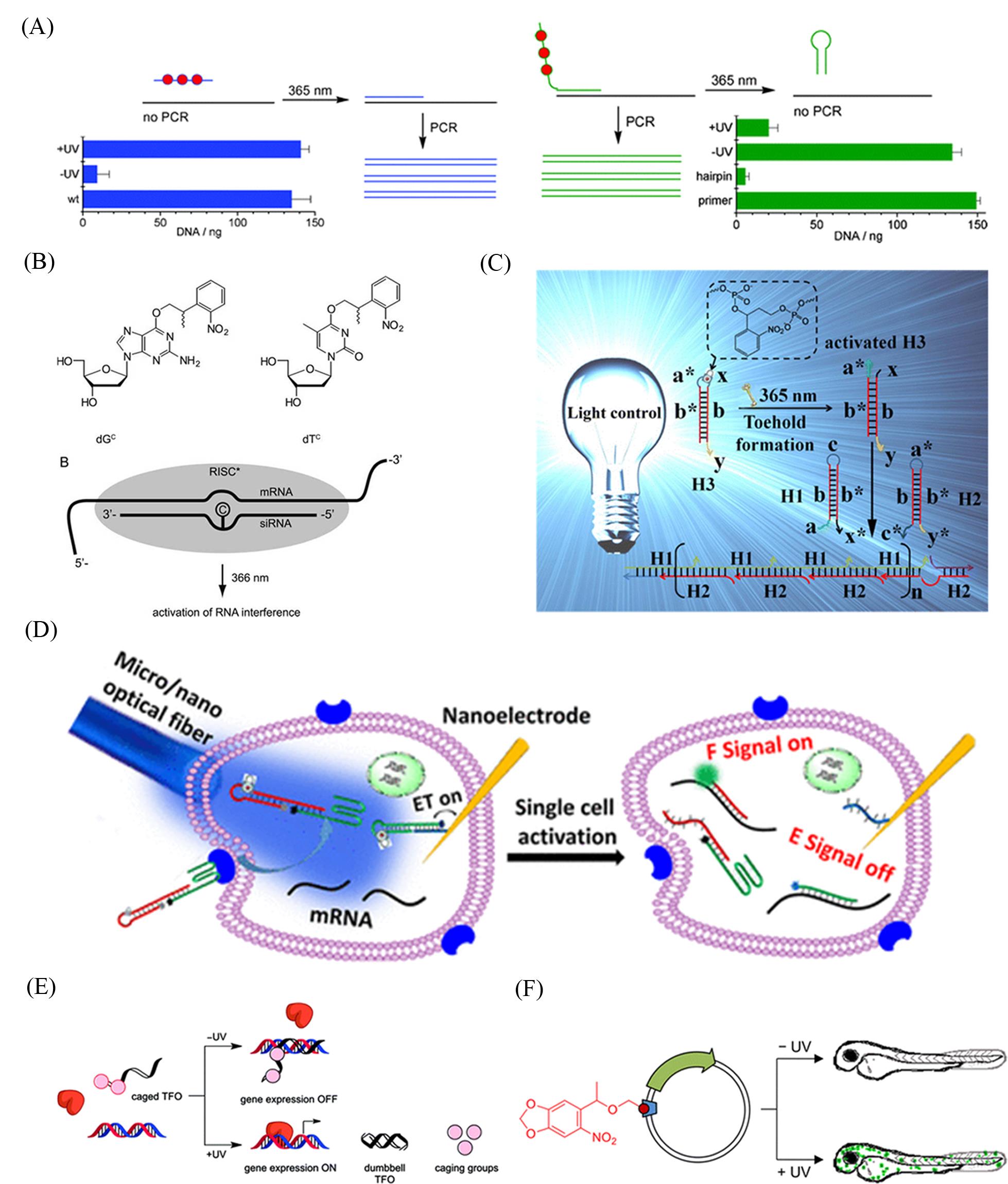
Fig.6 Photoswitchable oligonucleotides and nucleic acids(A) PCR activation (left) or deactivation (right) by light[59]. Copyright 2008, the Royal Society of Chemistry. (B) Light?dependent RNA interference with nucleobase?caged siRNAs[60]. Copyright 2007, RNA Society. (C) Photocontrolled toehold formation for toehold?mediated DNA branch migration reaction[62]. Copyright 2013, American Chemical Society. (D) Photoactivated toehold?mediated displacement reaction for mRNA detection in a single cell[63]. Copyright 2018, American Chemical Society. (E) The light?activation of gene expression using caged dumbbell TFOs[74]. Copyright 2012, American Chemical Society. (F) Optical control of transcription using a caged promoter[75]. Copyright 2014, American Chemical Society.
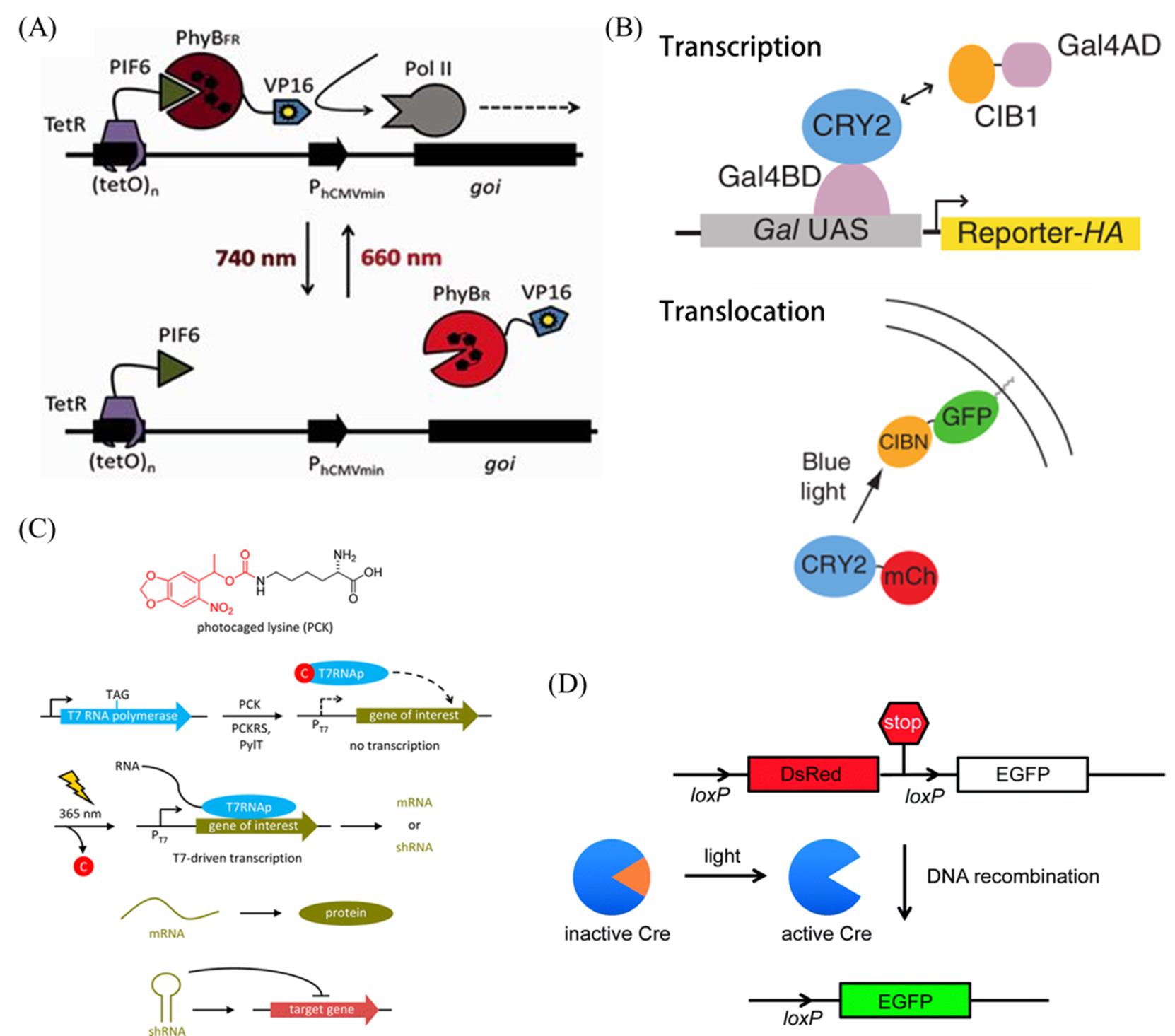
Fig.7 Photoswitchable Peptides and Proteins(A) Light?inducible expression system based on the concept of split transcription factors[81]. Copyright 2013, Oxford University Press. (B) Light?induced activation of transcription and translocation[82]. Copyright 2010, Springer Nature. (C) Photochemically control RNA polymerization in mammalian cells[85]. Copyright 2013, American Chemical Society. (D) Light?activation of caged Cre recombinase results in Cre?mediated recombination[86]. Copyright 2016, the Royal Society of Chemistry.
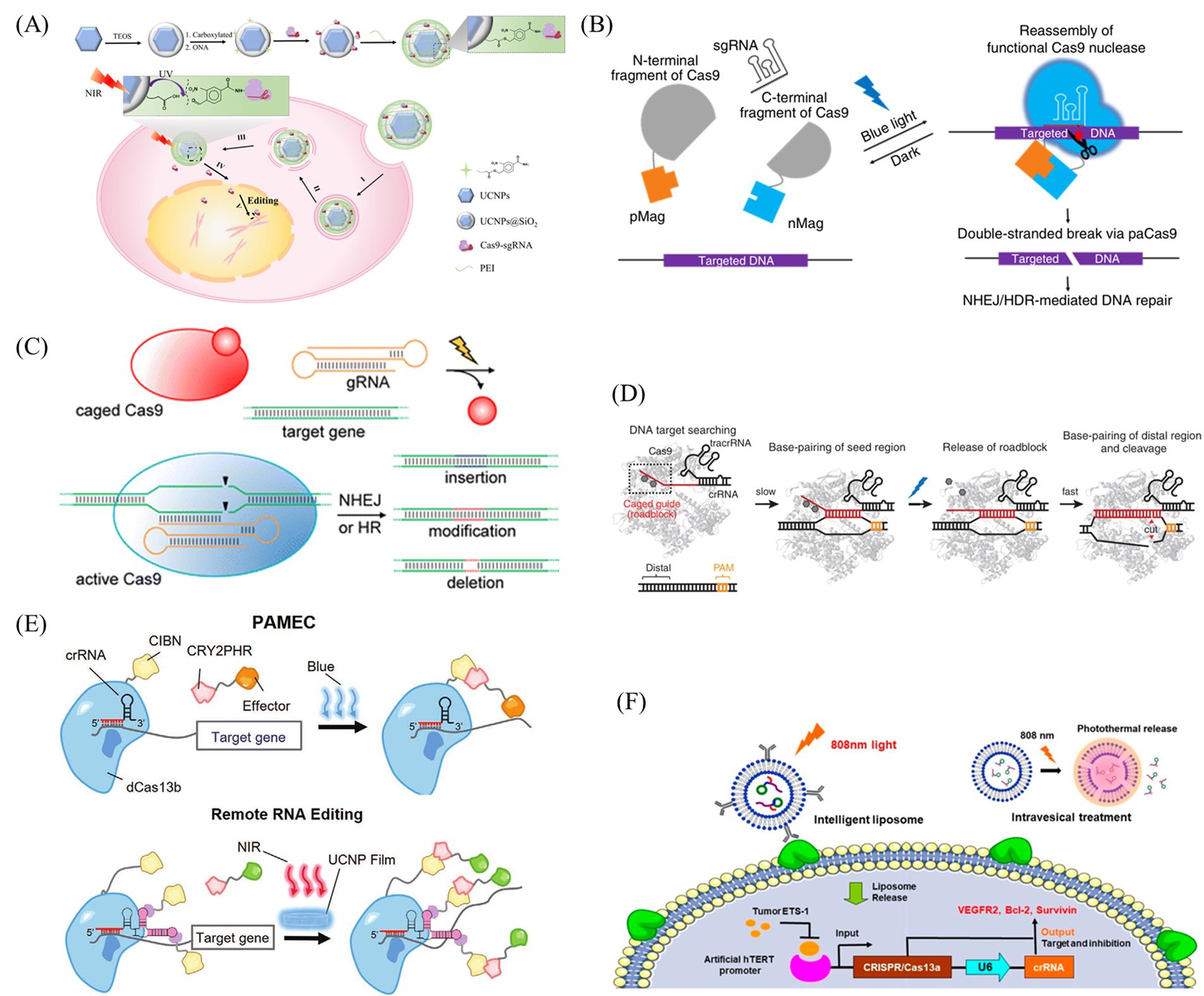
Fig.8 Light?controlled CRISPR?Cas system(A) Design of the UCNP?based CRISPR?Cas9 delivery system for NIR light?controlled gene editing[90]. Copyright 2019, American Association for the Advancement of Science. (B) Schematic of the paCas9[94]. Copyright 2015, Springer?Nature. (C) Light?activation of caged Cas9 enables optochemical control of gene editing[96]. Copyright 2015, American Chemical Society. (D) Schematic of the very fast CRISPR[97]. Copyright 2020, American Association for the Advancement of Science. (E) Schematic of the PAMEC system and NIR light activated PAMECR system[100]. Copyright 2020, Wiley?VCH. (F) Construction of the intelligent liposome delivery system[102]. Copyright 2019, American Chemical Society.
| 1 | Zhao Z. W., Ma Q., Ma L. N., Wang J., Wang X. W., Li Y. K., Journal of Ningxia University(Natural Science Edition), 2021, 42(2), 184—191(赵正伟, 马青, 马丽娜, 王锦, 王晓薇, 李颖康. 宁夏大学学报: 自然科学版, 2021, 42(2), 184—191) |
| 2 | Kang X. L., Chu D. D., Shan B., World Notes on Antibiotics,2020, 41(1), 35—41(康细林, 储丹丹, 单斌. 国外医药(抗生素分册), 2020, 41(1), 35—41) |
| 3 | Marraffini L. A., Nature., 2015, 526(7571), 55—61 |
| 4 | Amitai G., Sorek R., Nat. Rev. Microbiol., 2016, 14(2), 67—76 |
| 5 | Anzalone A. V., Koblan L. W., Liu D. R., Nat. Biotechnol., 2020, 38(7), 824—844 |
| 6 | Salsman J., Dellaire G., Biochem. Cell Biol., 2017, 95(2), 187—201 |
| 7 | Modell A. E., Siriwardena S. U., Shoba V. M., Li X., Choudhary A., Curr. Opin. Chem. Biol., 2021, 60, 113—121 |
| 8 | Javed M. R., Sadaf M., Ahmed T., Jamil A., Nawaz M., Abbas H., Ijaz A., Curr. Microbiol., 2018, 75(12), 1675—1683 |
| 9 | Zhang J., Chen L., Zhang J., Wang Y., Comput. Struct. Biotechnol. J., 2019, 17, 1171—1177 |
| 10 | Hryhorowicz M., Grześkowiak B., Mazurkiewicz N., Śledziński P., Lipiński D., Słomski R., Mol. Biotechnol., 2019, 61(3), 173—180 |
| 11 | Moroz⁃Omori E. V., Satyapertiwi D., Ramel M. C., Høgset H., Sunyovszki I. K., Liu Z., Wojciechowski J. P., Zhang Y., Grigsby C. L., Brito L., Bugeon L., Dallman M. J., Stevens M. M., ACS Cent Sci., 2020, 6(5), 695—703 |
| 12 | Zhang D., Tang B., Xie X., Xiao Y. F., Yang S. M., Zhang J. W., Cancer Biol. Ther., 2015, 16(7), 1005—1013 |
| 13 | Pawelczak K. S., Gavande N. S., VanderVere⁃Carozza P. S., Turchi J. J., ACS Chem. Biol., 2018, 13(2), 389—396 |
| 14 | Carroll D., Yale J. Biol. Med., 2017, 90(4), 653—659 |
| 15 | Gaj T., Gersbach C. A., Barbas C. F., 3rd Trends Biotechnol., 2013, 31(7), 397—405 |
| 16 | Urnov F. D., Rebar E. J., Holmes M. C., Zhang H. S., Gregory P. D., Nat. Rev. Genet., 2010, 11(9), 636—646 |
| 17 | Gupta D., Bhattacharjee O., Mandal D., Sen M. K., Dey D., Dasgupta A., Kazi T. A., Gupta R., Sinharoy S., Acharya K., Chattopadhyay D., Ravichandiran V., Roy S., Ghosh D., Life Sci., 2019, 232, 116636 |
| 18 | Koonin E. V., Makarova K. S., Zhang F., Curr. Opin. Microbiol., 2017, 37, 67—78 |
| 19 | Koonin E. V., Makarova K. S., Philos. Trans R Soc. Lond B Biol. Sci., 2019, 374(1772), 20180087 |
| 20 | Barman A., Deb B., Chakraborty S., Curr. Genet., 2020, 66(3), 447—462 |
| 21 | Ran F. A., Hsu P. D., Wright J., Agarwala V., Scott D. A., Zhang F., Nat. Protoc., 2013, 8(11), 2281—2308 |
| 22 | Jiang F., Doudna J. A., Annu. Rev. Biophys., 2017, 46, 505—529 |
| 23 | Shen B., Zhang W., Zhang J., Zhou J., Wang J., Chen L., Wang L., Hodgkins A., Iyer V., Huang X., Skarnes W. C., Nat. Methods, 2014, 11(4), 399—402 |
| 24 | Brocken D., Tark⁃Dame M., Dame R. T., Curr. Issues Mol. Biol., 2018, 26, 15—32 |
| 25 | Kleinstiver B. P., Pattanayak V., Prew M. S., Tsai S. Q., Nguyen N. T., Zheng Z., Joung J. K., Nature., 2016, 529(7587), 490—495 |
| 26 | Gao P., Lyu Q., Ghanam A. R., Lazzarotto C. R., Newby G. A., Zhang W., Choi M., Slivano O. J., Holden K., Walker J. A., 2nd Kadina A. P., Munroe R. J., Abratte C. M., Schimenti J. C., Liu D R., Tsai S. Q., Long X., Miano J. M., Genome Biol., 2021, 22(1), 83 |
| 27 | Moghadam F., LeGraw R., Velazquez J. J., Yeo N. C., Xu C., Park J., Chavez A., Ebrahimkhani M. R., Kiani S., Nat. Cell Biol., 2020, 22(9), 1143—1154 |
| 28 | Zhan T., Rindtorff N., Betge J., Ebert M. P., Boutros M., Semin. Cancer Biol., 2019, 55, 106—119 |
| 29 | Zhang Y., Ma X., Xie X., Liu Y. G., Prog. Mol. Biol. Transl. Sci., 2017, 149, 133—150 |
| 30 | Swarts D. C., van der Oost J., Jinek M., Mol. Cell., 2017, 66(2), 221—233 |
| 31 | Stella S., Alcón P., Montoya G., Nature, 2017, 546(7659), 559—563 |
| 32 | Safari F., Zare K., Negahdaripour M., Barekati⁃Mowahed M., Ghasemi Y., Cell Biosci., 2019, 9, 36 |
| 33 | Fonfara I., Richter H., Bratovič M., Le Rhun A., Charpentier E., Nature, 2016, 532(7600), 517—521 |
| 34 | Moreno⁃Mateos M. A., Fernandez J. P., Rouet R., Vejnar C. E., Lane M. A., Mis E., Khokha M. K., Doudna J. A., Giraldez A. J., Nat. Commun., 2017, 8(1), 2024 |
| 35 | Campa C. C., Weisbach N. R., Santinha A. J., Incarnato D., Platt R. J., Nat. Methods, 2019, 16(9), 887—893 |
| 36 | Li T., Zhu L., Xiao B., Gong Z., Liao Q., Guo J., Biotechnol. Adv., 2019, 37(1), 21—27 |
| 37 | Liu Q., Zhang Y., Li F., Li J., Sun W., Tian C., Biotechnol. Biofuels, 2019, 12, 293 |
| 38 | Ding X., Yin K., Li Z., Liu C., bioRxiv., 2020, 2020.03.19.998724 |
| 39 | Dai Y., Somoza R. A., Wang L., Welter J. F., Li Y., Caplan A. I., Liu C. C., Angew. Chem. Int. Ed. Engl.,2019, 58(48), 17399—17405 |
| 40 | Liu L., Li X., Wang J., Wang M., Chen P., Yin M., Li J., Sheng G., Wang Y., Cell, 2017, 168(1/2), 121—134 |
| 41 | East⁃Seletsky A., O'Connell M. R., Knight S. C., Burstein D., Cate J. H., Tjian R., Doudna, J. A., Nature, 2016, 538(7624), 270—273 |
| 42 | O'Connell M. R., J. Mol. Biol., 2018, 431(1), 66—87 |
| 43 | Abudayyeh O. O., Gootenberg J. S., Essletzbichler P., Han S., Joung J., Belanto J. J., Verdine V., Cox D., Kellner M. J., Regev A., Lander E. S., Voytas D. F., Ting A. Y., Zhang F., Nature, 2017, 550(7675), 280—284 |
| 44 | Gootenberg J. S., Abudayyeh O. O., Lee J. W., Essletzbichler P., Dy A. J., Joung J., Verdine V., Donghia N., Daringer N. M., Freije C. A., Myhrvold C., Bhattacharyya R. P., Livny J., Regev A., Koonin E. V., Hung D. T., Sabeti P. C., Collins J. J., Zhang F., Science, 2017, 356(6336), 438—442 |
| 45 | Jing X., Xie B., Chen L., Zhang N., Jiang Y., Qin H., Wang H., Hao P., Yang S., Li X., Nucleic Acids Res., 2018, 46(15), e90 |
| 46 | Zhao X., Liu L., Lang J., Cheng K., Wang Y., Li X., Shi J., Wang Y., Nie G., Cancer Lett., 2018, 431, 171—181 |
| 47 | Palaz F., Kalkan A. K., Can Ö., Demir A. N., Tozluyurt A., Özcan A., Ozsoz M., ACS Synth. Biol., 2021, 10(6), 1245—1267 |
| 48 | Silva J. M., Silva E., Reis R. L. J. Control Release, 2019, 298, 154—176 |
| 49 | Ricken J., Medda R., Wegner S. V., Adv. Biosyst., 2019, 3(3), e1800302 |
| 50 | Ahmed I., Fruk L., Mol. Biosyst., 2013, 9(4), 565—570 |
| 51 | Mayer G., Heckel A., Angew. Chem. Int. Ed. Engl., 2006, 45(30), 4900—4921 |
| 52 | Ellis-Davies G. C., Nat. Methods, 2007, 4(8), 619—628 |
| 53 | Yu H., Li J., Wu D., Qiu Z., Zhang Y., Chem. Soc. Rev., 2010, 39(2), 464—473 |
| 54 | Klán P., Šolomek T., Bochet C. G., Blanc A., Givens R., Rubina M., Popik V., Kostikov A., Wirz J., Chem. Rev., 2013, 113(1), 119—191 |
| 55 | Casey J. P., Blidner R. A., Monroe W. T., Mol. Pharm., 2009, 6(3), 669—685 |
| 56 | Brieke C., Rohrbach F., Gottschalk A., Mayer G., Heckel A., Angew. Chem. Int. Ed. Engl., 2012, 51(34), 8446—8476 |
| 57 | Hager S., Fittler F. J., Wagner E., Bros M., Cells, 2020, 9(9), 2061 |
| 58 | Chaulk S. G., MacMillan A. M., Nat. Protoc., 2007, 2(5), 1052—1058 |
| 59 | Young D. D., Edwards W. F., Lusic H., Lively M. O., Deiters A., Chem. Commun.(Camb)., 2008, (4), 462—464 |
| 60 | Mikat V., Heckel A., RNA, 2007, 13(12), 2341—2347 |
| 61 | Bardhan A., Deiters A., Ettensohn C. A., Dev. Biol., 2021, 475, 21—29 |
| 62 | Huang F., You M., Han D., Xiong X., Liang H., Tan W., J. Am. Chem. Soc., 2013, 135(21), 7967—7973 |
| 63 | Huang F., Lin M., Duan R., Lou X., Xia F., Willner I., Nano Lett., 2018, 18(8), 5116—5123 |
| 64 | Duan R., Li T., Duan Z., Huang F., Xia F., Anal. Chem., 2020, 92(8), 5846—5854 |
| 65 | Huang F., Liao W. C., Sohn Y. S., Nechushtai R., Lu C. H., Willner I., J. Am. Chem. Soc., 2016, 138(28), 8936—8945 |
| 66 | Huang F., Duan R., Zhou Z., Vázquez-González M., Xia F., Willner I.,Chem. Sci., 2020, 11(21), 5592—5600 |
| 67 | Huang F., Xu H., Tan W., Liang H., ACS Nano, 2014, 8(7), 6849—6855 |
| 68 | Huang F., Zhou X., Yao D., Xiao S., Liang H., Small, 2015, 11(43), 5800—5806 |
| 69 | Huang F., Zhang J., Li T., Duan R., Xia F., Willner I., Nano Lett., 2019, 19(1), 618—625 |
| 70 | Tallafuss A., Gibson D., Morcos P., Li Y., Seredick S., Eisen J., Washbourne P., Development, 2012, 139(9), 1691—1699 |
| 71 | Shestopalov I. A., Sinha S., Chen J. K., Nat. Chem. Biol., 2007, 3(10), 650—651 |
| 72 | Tang X., Swaminathan J., Gewirtz A. M., Dmochowski I. J., Nucleic Acids Res., 2008, 36(2), 559—569 |
| 73 | Jain P. K., Shah S., Friedman S. H., J. Am. Chem. Soc., 2011, 133(3), 440—446 |
| 74 | Govan J. M., Uprety R., Hemphill J., Lively M. O., Deiters A., ACS Chem. Biol., 2012, 7(7), 1247—1256 |
| 75 | Hemphill J., Govan J., Uprety R., Tsang M., Deiters A., J. Am. Chem. Soc., 2014, 136(19), 7152—7158 |
| 76 | Baker A. S., Deiters A., ACS Chem Biol., 2014, 9(7), 1398—1407 |
| 77 | Liu Q., Tucker C. L. Curr. Opin. Chem. Biol., 2017, 40, 17—23 |
| 78 | Baker A. S., Deiters A., ACS Chem. Biol., 2014, 9(7), 1398—1407 |
| 79 | Yazawa M., Sadaghiani A. M., Hsueh B., Dolmetsch R. E., Nat. Biotechnol., 2009, 27(10), 941—945 |
| 80 | Crefcoeur R. P., Yin R., Ulm R., Halazonetis T. D., Nat. Commun., 2013, 4, 1779 |
| 81 | Müller K., Engesser R., Metzger S., Schulz S., Kämpf M. M., Busacker M., Steinberg T., Tomakidi P., Ehrbar M., Nagy F., Timmer J., Zubriggen M. D., Weber W., Nucleic Acids Res., 2013, 41(7), e77 |
| 82 | Kennedy M. J., Hughes R. M., Peteya L. A., Schwartz J. W., Ehlers M. D., Tucker C. L., Nat. Methods, 2010,7(12), 973—975 |
| 83 | Arbely E., Torres⁃Kolbus J., Deiters A., Chin J. W., J. Am. Chem. Soc., 2012, 134(29), 11912—11915 |
| 84 | Gautier A., Nguyen D. P., Lusic H., An W., Deiters A., Chin J. W., J. Am. Chem. Soc., 2010, 132(12), 4086—4088 |
| 85 | Hemphill J., Chou C., Chin J. W., Deiters A., J. Am. Chem. Soc., 2013, 135(36), 13433—13439 |
| 86 | Luo J., Arbely E., Zhang J., Chou C., Uprety R., Chin J. W., Deiters A., Chem. Commun.(Camb)., 2016, 52(55), 8529—8532 |
| 87 | Ren W., Ji A., Ai H. W., J. Am. Chem. Soc., 2015, 137(6), 2155—2158 |
| 88 | Barman A., Deb B., Chakraborty S., Curr. Genet., 2020, 66(3), 447—462 |
| 89 | Swarts D. C., Jinek M., Wiley Interdiscip Rev. RNA, 2018, e1481 |
| 90 | Pan Y., Yang J., Luan X., Liu X., Li X., Yang J., Huang T., Sun L., Wang Y., Lin Y., Song Y., Sci. Adv., 2019, 5(4), eaav7199 |
| 91 | Moroz⁃Omori E. V., Satyapertiwi D., Ramel M. C., Høgset H., Sunyovszki I. K., Liu Z., Wojciechowski J. P., Zhang Y., Grigsby C. L., Brito L., Bugeon L., Dallman M. J., Stevens M. M., ACS Cent. Sci., 2020, 6(5), 695—703 |
| 92 | Zhang Y., Ling X., Su X., Zhang S., Wang J., Zhang P., Feng W., Zhu Y. Y., Liu T., Tang X., Angew. Chem. Int. Ed. Engl., 2020, 59(47), 20895—20899 |
| 93 | Aksoy Y. A., Yang B., Chen W., Hung T., Kuchel R. P., Zammit N. W., Grey S. T., Goldys E. M., Deng W., ACS Appl. Mater. Interfaces, 2020, 12(47), 52433—52444 |
| 94 | Nihongaki Y., Kawano F., Nakajima T., Sato M., Nat. Biotechnol., 2015, 33(7), 755—760 |
| 95 | Yu Y., Wu X., Guan N., Shao J., Li H., Chen Y., Ping Y., Li D., Ye H., Sci. Adv., 2020, 6(28), eabb1777 |
| 96 | Hemphill J., Borchardt E. K., Brown K., Asokan A., Deiters A., J. Am. Chem. Soc., 2015, 137(17), 5642—5645 |
| 97 | Liu Y., Zou R. S., He S., Nihongaki Y., Li X., Razavi S., Wu B., Ha T., Science, 2020, 368(6496), 1265—1269 |
| 98 | Zhou W., Brown W., Bardhan A., Delaney M., Ilk A. S., Rauen R. R., Kahn S. I., Tsang M., Deiters A., Angew. Chem. Int. Ed. Engl.,2020, 59(23), 8998—9003 |
| 99 | Carlson-Stevermer J., Kelso R., Kadina A., Joshi S., Rossi N., Walker J., Stoner R., Maures T., Nat. Commun., 2020, 11(1), 5041 |
| 100 | Zhao J., Li B., Ma J., Jin W., Ma X., Small, 2020, 16(30), e1907301 |
| 101 | Qi F., Tan B., Ma F., Zhu B., Zhang L., Liu X., Li H., Yang J., Cheng B., Int. J. Biol. Sci., 2019, 15(8), 1630—1636 |
| 102 | Fan J., Liu Y., Liu L., Huang Y., Li X., Huang W., ACS Synth. Biol., 2019, 9(2), 343—355 |
| 103 | Fu Y., Foden J. A., Khayter C., Maeder M. L., Reyon D., Joung J. K., Sander J. D., Nat. Biotechnol., 2013, 31(9), 822—826 |
| 104 | Pattanayak V., Lin S., Guilinger J. P., Ma E., Doudna J. A., Liu D. R., Nat. Biotechnol., 2013, 31(9), 839—843 |
| 105 | Tsai S. Q., Zheng Z., Nguyen N. T., Liebers M., Topkar V. V., Thapar V., Wyvekens N., Khayter C., Iafrate A. J., Le L. P., Aryee M. J., Joung J. K., Nat. Biotechnol., 2015, 33(2), 187—197 |
| 106 | Slaymaker I. M., Gao L., Zetsche B., Scott D. A., Yan W. X., Zhang F., Science, 2016, 351(6268), 84—88 |
| 107 | Lin Y., Cradick T. J., Brown M. T., Deshmukh H., Ranjan P., Sarode N., Wile B. M., Vertino P. M., Stewart F. J., Bao G., Nucleic Acids Res., 2014, 42(11), 7473—7485 |
| 108 | Hsu P. D., Scott D. A., Weinstein J. A., Ran F. A., Konermann S., Agarwala V., Li Y., Fine E. J., Wu X., Shalem O., Cradick T. J., Marraffini L. A., Bao G., Zhang F., Nat. Biotechnol., 2013, 31(9), 827—832 |
| 109 | Feldbauer K., Zimmermann D., Pintschovius V., Spitz J., Bamann C., Bamberg E., Proc. Natl. Acad. Sci. USA, 2009, 106(30), 12317—12322 |
| 110 | Lu Y., Xue J., Deng T., Zhou X., Yu K., Deng L., Huang M., Yi X., Liang M., Wang Y., Shen H., Tong R., Wang W., Li L., Song J., Li J., Su X., Ding Z., Gong Y., Zhu J., Mok T., Nat. Med., 2020, 26(5), 732—740 |
| 111 | Maeder M. L., Stefanidakis M., Wilson C. J., Baral R., Barrera L. A., Bounoutas G. S., Bumcrot D., Chao H., Ciulla D. M., DaSilva J. A., Dass A., Dhanapal V., Fennell T. J., Friedland A. E., Giannoukos G., Gloskowski S. W., Glucksmann A., Gotta G. M., Jayaram H., Haskett S. J., Jiang H., Nat. Med., 2019, 25(2), 229—233 |
| [1] | 常丽颖 凌鑫宇 陈和祺 王雪 刘涛. 基因编辑在线粒体疾病中的应用[J]. 高等学校化学学报, 2022, 43(Album-4): 20220363. |
| [2] | 王斌, 王晓红, 段莉梅, 许良, 赵鹏, 菅金鹏, 刘宗瑞. 基于P2Mo18 |
| [3] | 王斌, 王晓红, 刘宗瑞, 段莉梅, 徐玲, 白锁柱. 多酸/邻菲罗啉钌LBL薄膜的制备及电化学调控荧光开关性能[J]. 高等学校化学学报, 2016, 37(5): 822. |
| [4] | 靳琳, 刘禹, 曹子谏, 孟杰, 姜振华, 张大明. 高热稳定性氟化聚酰亚胺的合成及在热光开关中的应用[J]. 高等学校化学学报, 2012, 33(07): 1591. |
| [5] | 聂晶, 韩美娇, 王科志 . [Ru(phen)2dppz]2+二聚及对DNA键合性质的影响[J]. 高等学校化学学报, 2007, 28(10): 1833. |
| [6] | 郝强; 段智明; 韩美娇; 郑帅至; 吕媛媛; 王科志. 新型DNA分子光开关钌(Ⅱ) 配合物的研究[J]. 高等学校化学学报, 2006, 27(7): 1217. |
| [7] | 周翠松,江雅新,汪俊,麻宝成,李梦龙,方晓红 . 信号核酸识体用于药物托普霉素的高灵敏度检测[J]. 高等学校化学学报, 2006, 27(5): 826. |
| [8] | 沈永淼, 王炳祥, 冯玉英, 沈珠英, 沈健, 李邨, 胡宏纹. 氨基苯基类中氮茚化合物的合成及作为质子探针的研究[J]. 高等学校化学学报, 2006, 27(4): 651. |
| [9] | 孙景志, 杨新国, 李寒莹, 黄骥, 汪茫. 酸碱控制的卟啉-苝酰亚胺分子阵列荧光开关[J]. 高等学校化学学报, 2004, 25(11): 2148. |
| [10] | 凌连生, 何治柯, 吴风武, 罗庆尧, 曾云鹗. 核酸分子“光开关”的研究进展[J]. 高等学校化学学报, 2000, 21(4): 527. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||