

高等学校化学学报 ›› 2015, Vol. 36 ›› Issue (11): 2271.doi: 10.7503/cjcu20150651
收稿日期:2015-08-14
出版日期:2015-11-10
发布日期:2015-10-21
作者简介:联系人简介: 步宇翔, 男, 博士, 教授, 博士生导师, 主要从事理论计算与模拟化学方面的研究. E-mail:基金资助:
WANG Mei1,2, WANG Jun1, BU Yuxiang1,*( )
)
Received:2015-08-14
Online:2015-11-10
Published:2015-10-21
Contact:
BU Yuxiang
E-mail:byx@sdu.edu.cn
摘要:
利用密度泛函理论方法研究了作为空穴迁移载体的蛋白质复合的DNA三聚体(Protonated arginine…guanine…cytosine, ArgH+-GC)的氢键性质. 结果表明, 空穴迁移通过该载体单元时此类氢键表现为亚稳态, 且具有明显的负离解能. 正常情况下ArgH+基团在大小沟均能与GC碱对形成氢键, 且具有正的离解能. 然而, 当空穴转移至此将削弱氢键至亚稳态, 使之具有一定的离解势垒和负的离解能. 这种势垒抑制的负离解能现象意味着由于空穴俘获导致此三聚体结构单元在它的ArgH+…N7/O6键区储存了一定的能量(约108.78 kJ/mol). 该氢键离解通道受控于此键区两个相关组分之间的静电排斥和氢键吸引之间的平衡以及这两个相反作用随氢键距离不同的衰减速率. 基于电子密度分布的拓扑性质以及键临界点的Laplacian数值分析澄清了此类特殊的能量现象主要源自通过高能氢键(ArgH+…N7/O6)连接的授受体间的静电排斥. 进一步空穴俘获诱导的G→C质子转移可扩展负离解能区至ArgH+…N7/O6和Watson-Crick(WC) 氢键区. 另外, ArgH+ 结合到GC的大小沟增加其电离势, 因此削弱其空穴传导能力, 削弱程度取决于ArgH+与GC的距离. 推而广之, 在protonated lysine-GC和protonated histidine-GC体系也可观察到类似的现象. 显然, 此类性质可调的亚稳态氢键可调控DNA空穴迁移机理. 此工作为理解蛋白质调控的DNA空穴迁移机理提供了重要的能量学信息.
中图分类号:
TrendMD:
王梅, 王军, 步宇翔. 与蛋白质调控DNA空穴迁移相关的具有负离解能特征的亚稳态氢键. 高等学校化学学报, 2015, 36(11): 2271.
WANG Mei, WANG Jun, BU Yuxiang. Metastable Hydrogen-bonds Featuring Negative Dissociation Energies in Protein-bound DNA in Hole Migration. Chem. J. Chinese Universities, 2015, 36(11): 2271.
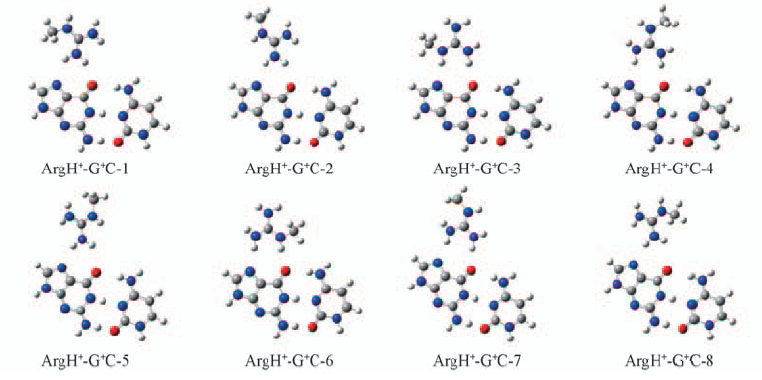
Fig.2 Optimized geometrical structures of eight complexes ArgH+-G+C-n(n=1—8) in which positive charged arginine residue resides in the major-groove face of DNA base pair
| Complex | R(N—H…N) | R(N—H…O) | RE | ΔE | ΔEBSSE | DE | AIP |
|---|---|---|---|---|---|---|---|
| ArgH+-G+C-1 | 0.2354 | 0.2123 | 11.25 | -118.95 | -120.79 | 5.56 | 911.19 |
| ArgH+-G+C-2 | 0.2203 | 0.2290 | 9.75 | -117.45 | -119.04 | 4.81 | 908.51 |
| ArgH+-G+C-3 | 0.2150 | 0.2003 | 3.01 | -110.67 | -113.22 | 5.98 | 919.22 |
| ArgH+-G+C-4 | 0.2101 | 0.2017 | 0.38 | -108.07 | -110.46 | 6.07 | 918.05 |
| ArgH+-G+C-5 | 0.2153 | 0.2429 | 9.87 | -117.53 | -119.12 | 4.56 | 920.73 |
| ArgH+-G+C-6 | 0.2101 | 0.2050 | 0 | -111.21 | -113.72 | 6.40 | 937.09 |
| ArgH+-G+C-7 | 0.2104 | 0.2011 | 3.51 | -107.70 | -110.08 | 5.94 | 934.50 |
| ArgH+-G+C-8 | 0.2117 | 0.2687 | 11.21 | -118.87 | -120.67 | 4.48 | 922.99 |
Table 1 Lengths(nm) of two hydrogen bonds(N—H…N7 H-bond and N—H…O6 H-bond) and dissociation energies(ΔE) of eight complexes ArgH+-G+C-n(n=1—8)*
| Complex | R(N—H…N) | R(N—H…O) | RE | ΔE | ΔEBSSE | DE | AIP |
|---|---|---|---|---|---|---|---|
| ArgH+-G+C-1 | 0.2354 | 0.2123 | 11.25 | -118.95 | -120.79 | 5.56 | 911.19 |
| ArgH+-G+C-2 | 0.2203 | 0.2290 | 9.75 | -117.45 | -119.04 | 4.81 | 908.51 |
| ArgH+-G+C-3 | 0.2150 | 0.2003 | 3.01 | -110.67 | -113.22 | 5.98 | 919.22 |
| ArgH+-G+C-4 | 0.2101 | 0.2017 | 0.38 | -108.07 | -110.46 | 6.07 | 918.05 |
| ArgH+-G+C-5 | 0.2153 | 0.2429 | 9.87 | -117.53 | -119.12 | 4.56 | 920.73 |
| ArgH+-G+C-6 | 0.2101 | 0.2050 | 0 | -111.21 | -113.72 | 6.40 | 937.09 |
| ArgH+-G+C-7 | 0.2104 | 0.2011 | 3.51 | -107.70 | -110.08 | 5.94 | 934.50 |
| ArgH+-G+C-8 | 0.2117 | 0.2687 | 11.21 | -118.87 | -120.67 | 4.48 | 922.99 |
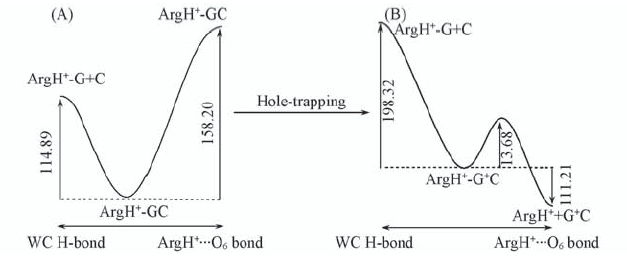
Fig.3 Schematic profiles of PES along WC and Hoogsteen H-bond dissociation coordinates of trimer complex units in their initial(ArgH+-GC) and oxidized(ArgH+-G+C) The corresponding dissociation are also shown. All the energies are expressed in kJ/mol.
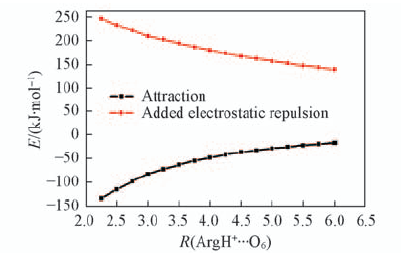
Fig.7 Effects of R(ArgH+…O6)(the distance between ArgH+ and O6 atom) on attraction interaction between ArgH+ and GC in ArgH+-GC, and added electrostatic repulsion upon one-electron oxidation The data for this figure are given in Table S4(see the Electronic Supporting Information of this paper).
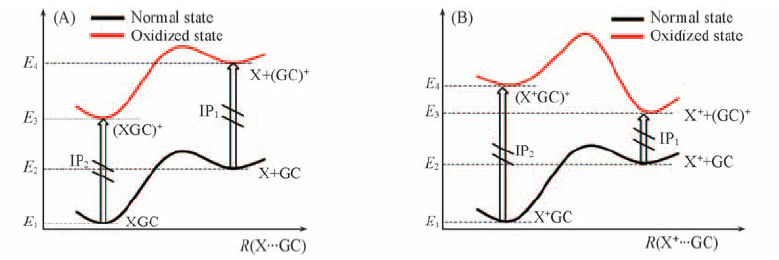
Fig.8 Potential energy surfaces of complexes XGC, X+GC(A) and their one-electron oxidized derivatives(XGC)+ and (X+GC)+(B) when the monomer X(or X+) is separated from GC and G+C, respectively En is the corresponding energy; the corresponding IPs are also shown.
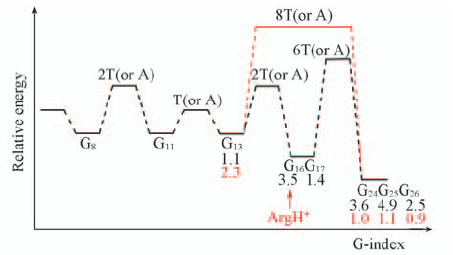
Fig.10 Schematic representation of potential energy landscapes through duplex DNA oligomers The DNA sequence is shown in Fig.S1. The “G”, “GG” and “GGG” denote an “isolated” guanine and two or three adjacent guanines, respectively. The “T” or “A” or “C” separates G or GG steps. The X-axis(“G-index”) indicates the position of guanines, GG steps, and GGG triplet along the oligomer and the energy barriers may be one base pair or some base pairs. The numbers in black and red indicate the band intensity relative to that of G24 in the protein-bound DNA duplex in the absence and presence of BamHI.
| [1] | Burrows C. J., Muller J. G., Chem. Rev., 1998, 98, 1109—1152 |
| [2] | LePage F., Guy A., Cadet J., Sarasin A., Gentil A., Nucleic Acids Res., 1998, 26, 1276—1281 |
| [3] | Burrows C. J., Muller J. G., Chem. Rev., 1998, 98, 1109—1152 |
| [4] | Hirakawa K., Ota K., Hirayama J., Oikawa S., Kawanishi S., Chem. Res. Toxicol., 2014, 27(4), 649—655 |
| [5] | Melvin T., Botchway S., Parker A. W., O’Neill P. J., Chem. Soc. Chem. Commun., 1995, 5, 653—654 |
| [6] | Steenken S., Jovanovic S. V., J. Am. Chem. Soc., 1997, 119, 617—618 |
| [7] | Jortner J., Bixon M., Langenbacher T., Michael-Beyerle M. E., Proc. Natl. Acad. Sci., 1998, 95, 12759—12765 |
| [8] | Berlin Y. A., Burin A. L., Ratner M. A., J. Am. Chem. Soc. ,2000, 122, 10903—10909 |
| [9] | Schuster G. B., Acc. Chem. Res. 2000, 33, 253—260 |
| [10] | Lewis F. D., Letsinger R. L., Wasielewski M. R., Acc. Chem. Res., 2001, 34, 159—170 |
| [11] | Wang J., Sun L.X., Bu Y. X., J. Phys. Chem. B, 2010, 114, 1144—1147 |
| [12] | Echols H., Science, 1986, 233, 1050—1056 |
| [13] | Kathuria P., Sharma P., Abendong M. N., Wetmore S. D., Biochemistry, 2015, 54(15), 2414—2428 |
| [14] | Qin P. H., Lü W. C., Qin W., Zhang W., Xie H., Chem. Res. Chinese Universities, 2014, 30(1), 125—129 |
| [15] | Corbella M., Voityuk A. A., Curutchet.C., J. Phys. Chem. Lett., 2015, 6(18), 3749—3753 |
| [16] | Paillard G., Lavery R., Structure, 2004, 12, 113—122 |
| [17] | Peters M., Rozas I., Alkorta I., Elguero J., J. Phys. Chem. B, 2003, 107, 323—330 |
| [18] | Wintjens R., Lievin J., Rooman M., Buisine E., J. Mol. Biol., 2000, 302(2), 395—410 |
| [19] | Warner D. R., Weinstein L. S., Proc. Natl. Acad. Sci., 1999, 96, 4268—4272 |
| [20] | Bond P. J., Guy A. T., Heron A. J., Bayley H., Khalid S., Biochemistry, 2011, 50(18), 3777—3783 |
| [21] | Jantz D., Berg J. M., J. Am. Chem. Soc., 2003, 125, 4960—4961 |
| [22] | Cheng A. C., Chen W. W., Fuhrmann C. N., Frankel A. D., J. Mol. Biol., 2003, 327, 781—796 |
| [23] | Allers J., Shamoo Y., J. Mol. Biol., 2001, 311, 75—86 |
| [24] | Davey C. A., Sargent D. F., Luger K., Maeder A. W., Richmond T. J., J. Mol. Biol., 2002, 319, 1097—1113 |
| [25] | Widom J., Annu. Rev. Biophys. Biomol. Struct., 1998, 27, 285—327 |
| [26] | Harp J. M., Hanson B. L., Tim D. E., Bunick G. J., Biol. Crystallogr., 2000, 56, 1513—1534 |
| [27] | Newman M., Strzelecka T., Dorner L. F., Schildkraut I., Aggarwal A. K., Science, 1995, 269, 656—663 |
| [28] | Rajski S. R., Barton J. K., Biochemistry, 2001, 40, 5556—5564 |
| [29] | Nunez M. E., Noyes K. T., Barton J. K., Chem. Biol., 2002, 9, 403—406 |
| [30] | Voityuk A. A., Davis W. B., J. Phys. Chem. B, 2007, 111, 2976—2985 |
| [31] | Bjorklund C. C., Davis W. B., Nucleic Acids Res., 2006, 34, 1836—1847 |
| [32] | Nakatani K., Dohno C., Saito I., J. Am. Chem. Soc., 2002, 124(24), 6802—6803 |
| [33] | Becke A. D., J. Chem. Phys., 1993, 98, 1372—1377 |
| [34] | Lee C., Yang W., Parr R. G., Phys. Rev. B, 1988, 37, 785—789 |
| [35] | Frisch M.J., Trucks G. W., Schlegel H. B., Scuseria G. E., Robb M. A., Cheeseman J. R., Zakrzewski V. G., Montgomery J. A., Stratmann Jr., R.E., Burant J. C., Dapprich S., Millam J. M., Daniels A. D., Kudin K. N., Strain M. C., Farkas O., Tomasi J., Barone V., Cossi M., Cammi R., Mennucci B., Pomelli C., Adamo C., Clifford S., Ochterski J., Petersson G. A., Ayala P. Y., Cui Q., Morokuma K., Rega N., Salvador P., Dannenberg J. J., Malick D. K., Rabuck A. D., Raghavachari K., Foresman J. B., Cioslowski J., Ortiz J.V., Baboul A. G., Stefanov B. B., Liu G., Liashenko A., Piskorz P., Komaromi I., Gomperts R., Martin R. L., Fox D. J., Keith T., Al-Laham M. A., Peng C. Y., Nanayakkara A., Challacombe M., Gill P. M. W., Johnson B., Chen W., Wong M. W., Andres J. L., Gonzalez C., Head-Gordon M., Replogle E.S., Pople J. A., Gaussian 03, Gaussian Inc., Pittsburgh, PA, 2003 |
| [36] | Boys S. F., Bernardi F., Mol. Phys., 1970, 19, 553—566 |
| [37] | Cioslowski J., Nanayakkara A., Challacombe M., Chem. Phys. Lett., 1993, 203, 137—142 |
| [38] | Cioslowski J., Surjan P. R., J. Mol. Struct.: Theochem., 1992, 255, 9—33 |
| [39] | Bader R. F. W., Encyclopedia of Computational Chemistry, 1998, 1, 64—86 |
| [40] | Bader R. F. W., Chem. Rev., 1991, 91, 893—928 |
| [1] | 张咪, 田亚锋, 高克利, 侯华, 王宝山. 三氟甲基磺酰氟绝缘介质理化特性的分子动力学模拟[J]. 高等学校化学学报, 2022, 43(11): 20220424. |
| [2] | 刘洋, 李旺昌, 张竹霞, 王芳, 杨文静, 郭臻, 崔鹏. Sc3C2@C80与[12]CPP纳米环之间非共价相互作用的理论研究[J]. 高等学校化学学报, 2022, 43(11): 20220457. |
| [3] | 王思佳 侯璐 李成龙 李文翠 陆安慧. 空腔型纳米炭的制备与应用[J]. 高等学校化学学报, 0, (): 20220637. |
| [4] | 武晴滢, 祝震予, 吴剑鸣, 徐昕. 泛Kennard-Stone算法的数据集代表性度量与分块采样策略[J]. 高等学校化学学报, 2022, 43(10): 20220397. |
| [5] | 王园月, 安梭梭, 郑旭明, 赵彦英. 5-巯基-1, 3, 4-噻二唑-2-硫酮微溶剂团簇的光谱和理论计算研究[J]. 高等学校化学学报, 2022, 43(10): 20220354. |
| [6] | 张伶育, 张继龙, 曲泽星. RDX分子内振动能量重分配的动力学研究[J]. 高等学校化学学报, 2022, 43(10): 20220393. |
| [7] | 沈琦 陈海瑶 高登辉 赵 熹 那日松 刘佳 黄旭日. 天然产物法卡林二醇与人类 GABAA 受体的相互作用机制研究[J]. 高等学校化学学报, 0, (): 0. |
| [8] | 陈少臣 程敏 王诗慧 吴金奎 罗磊 薛小雨 吉旭 张长春 周利. 预测金属有机骨架的甲烷和氢气输送能力的迁移学习建模[J]. 高等学校化学学报, 0, (): 20220459. |
| [9] | 彭辛哲, 葛娇阳, 王访丽, 余国静, 冉雪芹, 周栋, 杨磊, 解令海. 一种基于苯并噻吩平面格的张力与重组能的理论研究[J]. 高等学校化学学报, 0, (): 20220313. |
| [10] | 郭程, 张威, 唐云. 有序介孔材料: 历史、 现状与发展趋势[J]. 高等学校化学学报, 2022, 43(8): 20220167. |
| [11] | 汤乔伟 蔡小青 李江 诸颖 王丽华 田阳 樊春海 胡钧. 同步辐射X射线成像技术在脑成像研究中的应用[J]. 高等学校化学学报, 0, (): 20220379. |
| [12] | 杨丹, 刘旭, 戴翼虎, 祝艳, 杨艳辉. 金团簇电催化二氧化碳还原反应的研究进展[J]. 高等学校化学学报, 2022, 43(7): 20220198. |
| [13] | 戴卫, 侯华, 王宝山. 七氟异丁腈负离子结构与反应活性的理论研究[J]. 高等学校化学学报, 2022, 43(6): 20220044. |
| [14] | 施耐克, 张娅, SANSON Andrea, 王蕾, 陈骏. Zn(NCN)单轴的负热膨胀性及机理研究[J]. 高等学校化学学报, 2022, 43(6): 20220124. |
| [15] | 任娜娜, 薛洁, 王治钒, 姚晓霞, 王繁. 热力学数据对1, 3-丁二烯燃烧特性的影响[J]. 高等学校化学学报, 2022, 43(6): 20220151. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||