

Chem. J. Chinese Universities ›› 2015, Vol. 36 ›› Issue (12): 2386.doi: 10.7503/cjcu20150427
• Analytical Chemistry • Previous Articles Next Articles
Received:2015-05-26
Online:2015-12-10
Published:2015-11-17
Contact:
SHEN Hong
E-mail:shzju@zju.edu.cn
Supported by:CLC Number:
TrendMD:
WU Qiwang, SHEN Hong. NUPACK Prediction Assisted of Toehold Induced Strand Displacement Reaction and Its Application in SNPs Genotyping by DNAzyme-catalyzed Microfluidic Chemiluminescence Detection†[J]. Chem. J. Chinese Universities, 2015, 36(12): 2386.
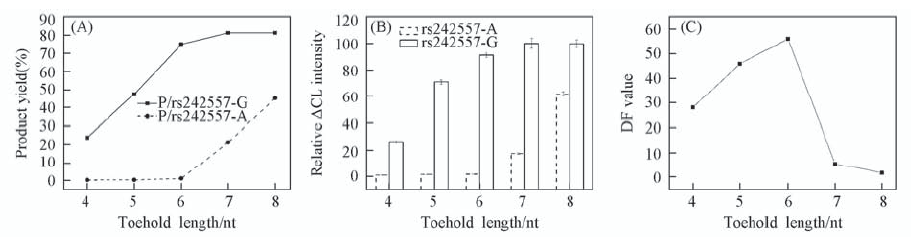
Fig.1 Impact of toehold length on SNP typing (A) Predicted by NUPACK simulations; (B) CL analysis; (C) DF calculation results. The concentrations of P strands, B strands and rs242557 are 150, 300 and 80 nmol/L, respectively.
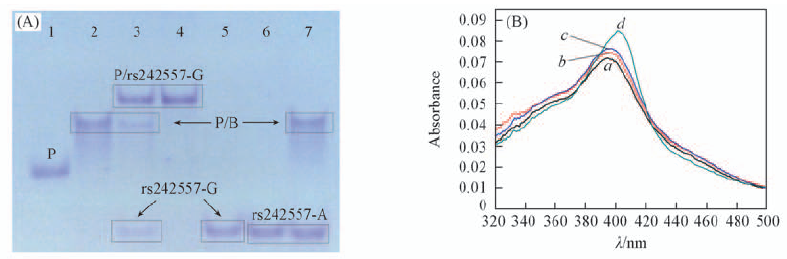
Fig.2 Native PAGE results(A) and UV-Vis absorption of hemin-oligonucleotide interaction(B) (A) Lane 1: P strand, lane 2: P strand annealed with B strand, lane 3: P/B reacted with rs242557-G, lane 4: P strand annealed with rs242557-G, lane 5: rs242557-G, lane 6: rs242557-A, lane 7: P/B reacted with rs242557-A; (B) hemin: 1.5 μmol/L, P strand: 300 nmol/L, B strand: 600 nmol/L, rs242557 strand: 200 nmol/L. a. Hemin; b. a+P/B; c. b+rs242557-A; d. b+rs242557-G.
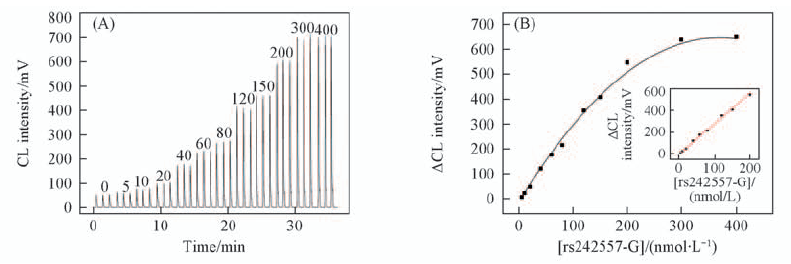
Fig.6 CL response to the different concentrations of sample rs242557-G solution(A) and corresponding calibration plot between net CL response and sample rs242557-G solution from 5 to 400 nmol/L(B) Insert of (B): magnification of the plots in the range of 5—200 nmol/L.
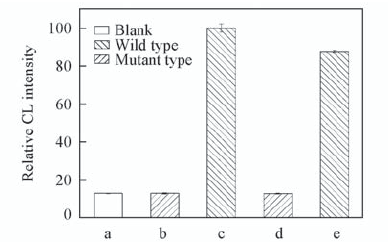
Fig.7 CL responses on control, wild type and mutant type sample solutions Target sample solutions: 80 nmol/L. a. Control; b. rs242557-A; c. rs242557-G; d. rs242557-A34; e. rs242557-G34.
| Sample | Added/(nmol·L-1) | Found/(nmol·L-1) | Recovery(%) | RSD(%, n=5) |
|---|---|---|---|---|
| 1 | 40 | 36.8 | 92.0 | 4.02 |
| 2 | 60 | 63.4 | 105.6 | 2.79 |
| 3 | 80 | 74.7 | 93.4 | 4.51 |
Table 2 DNA detection in biological samples
| Sample | Added/(nmol·L-1) | Found/(nmol·L-1) | Recovery(%) | RSD(%, n=5) |
|---|---|---|---|---|
| 1 | 40 | 36.8 | 92.0 | 4.02 |
| 2 | 60 | 63.4 | 105.6 | 2.79 |
| 3 | 80 | 74.7 | 93.4 | 4.51 |
| [1] | Stitziel N. O., Tseng Y. Y., Pervouchine D., Goddeau D., Kasif S., Liang J., J. Mol. Biol., 2003, 327, 1021—1030 |
| [2] | Myers A. J., Kaleem M., Marlowe L., Pittman A. M., Lees A., Fung H. C., Duckworth J., Leung D., Gibson A., Morris C. M., Hum. Mol. Genet., 2005, 14, 2399—2404 |
| [3] | Chen Y., Shortreed M. R., Peelen D., Lu M., Smith L. M., J. Am. Chem. Soc., 2004, 126, 3016—3017 |
| [4] | Litos I. K., Ioannou P. C., Christopoulos T. K., Traeger-Synodinos J., Kanavakis E., Anal. Chem., 2007, 79, 395—402 |
| [5] | Wang H., Li J., Wang Y., Jin J., Yang R., Wang K., Tan W., Anal. Chem., 2010, 82, 7684—7690 |
| [6] | Tyagi S., Kramer F.R.,Nat. Biotechnol., 1996, 303—308 |
| [7] | Du B. A., Li Z. P., Liu C. H., Angew. Chem. Int. Ed., 2006, 45, 8022—8025 |
| [8] | Neo J. L., AwK. D., Uttamchandani M., Analyst, 2011, 136, 1569—1572 |
| [9] | Zou J., Li N., Comput. Meth. Prog. Bio., 2013, 111, 755—762 |
| [10] | Markham N. R., Zuker M., Nucleic Acids Res., 2005, 33, W577—581 |
| [11] | Dirks R. M., Bois J. S., Schaeffer J. M., Winfree E., Pierce N. A., SIAM Rev., 2007, 49, 65—88 |
| [12] | Narcisi V., Mascini M., Perez G., Del Carlo,M., Tiscar P. G., Yamanaka H., Compagnone D., Talanta, 2011, 85, 1927—1932 |
| [13] | Zhang D. Y., Winfree E., J. Am. Chem. Soc., 2009, 131, 17303—17314 |
| [14] | Genot A.J., Zhang D. Y., Bath J., Turberfield A. J., J. Am. Chem. Soc., 2011, 133, 2177—2182 |
| [15] | Kosman J., Juskowiak B., Anal. Chim. Acta,2011, 707, 7—17 |
| [16] | Rao X. Y., Zhang J. J., Cui J., Hu Y., Liu T., Chai J. F., Cheng G. F., He P. G., Fang Y. Z., Chem. Res. Chinese Universities,2013, 29(5), 868—873 |
| [17] | Fan H. L., Zhang T., Jin W., Jin Q. H., Chem. J. Chinese Universities,2015, 36(4), 625—630 |
| (范宏亮, 张涛, 金伟, 金钦汉. 高等学校化学学报,2015, 36(4), 625—630) | |
| [18] | Song L., Pan X., Shen H., Yu Y., Luminescence, 2014, 29, 1066—1073 |
| [19] | Wang D., Tang W., Wu X., Wang X., Chen G., Chen Q., Li N., Liu F., Anal. Chem., 2012, 84, 7008—7014 |
| [20] | Zhang D. Y., Chen S. X., Yin P., Nat. Chem., 2012, 4, 208—214 |
| [21] | Travascio P., Li Y., Sen D., Chem.Biol., 1998, 5, 505—517 |
| [22] | Chen W., Li B., Xu C., Wang L., Biosens. Bioelectron., 2009, 24, 2534—2540 |
| [23] | Fan X., Li H., Zhao J., Lin F., Zhang L., Zhang Y., Yao S., Talanta, 2012, 89, 57—62 |
| [24] | Traore Z. S., Su X. G., Chem. Res. Chinese Universities,2012, 28(4), 604—608 |
| [25] | Sudarsan A. P., Ugaz V. M., Lab on A Chip,2006, 6, 74—82 |
| [26] | Kolpashchikov D. M., J. Am. Chem. Soc., 2008, 130, 2934—2935 |
| [27] | Zheng A. X., Li J., Wang J. R., Song X. R., Chen G. N., Yang H. H., Chem. Commun., 2012, 48, 3112—3114 |
| [1] | WANG Fangyuan, ZHANG Fenxian, LI Yi, GAO Jianhua, NIU Yanbing, SHEN Shaofei. Fabrication of Bionic Leaf Model and Its Application in Agarose Microfluidic Chip [J]. Chem. J. Chinese Universities, 2022, 43(11): 20220445. |
| [2] | MA Yanrong, JIANG Shengnan, JIN Yan. Sensitive and Electrochemical Detection of Telomerase Activity Based on the Signal Amplification of Strand Displacement Reaction [J]. Chem. J. Chinese Universities, 2021, 42(3): 745. |
| [3] | MAO Yu, QU Hao, ZHENG Lei. Research Progress on RNA⁃cleaving DNAzyme for the Detection of Pathogenic Bacteria [J]. Chem. J. Chinese Universities, 2021, 42(11): 3445. |
| [4] | LUO Cheng, PENG Yamei, SHEN Hong, FANG Qun, PAN Jianzhang. Research Progress of Multiplex Immunoassay [J]. Chem. J. Chinese Universities, 2021, 42(11): 3421. |
| [5] | PENG Huo, GAO Zehang, LIAO Chengyue, WANG Xiaodong, ZHOU Hongbo, ZHAO Jianlong. Robust Droplet Digital PCR Chip for Absolute Quantitative Detection of Nucleic Acid [J]. Chem. J. Chinese Universities, 2020, 41(8): 1760. |
| [6] | ZHANG Kaixiang, LIU Junjie, SONG Qiaoli, WANG Danyu, SHI Jinjin, ZHANG Haiyue, LI Jinghong. Multifunctional DNA Nanoflowers for Autophagy Inhibition and Enhanced Antitumor Chemotherapy† [J]. Chem. J. Chinese Universities, 2020, 41(7): 1461. |
| [7] | ZHANG Zuoran, ZHANG Li, ZHANG Zhiling. On-chip Sorting of Beads with Different Magnetic Responsiveness by Lateral Magnetophoresis † [J]. Chem. J. Chinese Universities, 2020, 41(6): 1243. |
| [8] | SHENG Bingchen, LI Cong, LIU Yingya, WANG Anjie, WANG Yao, ZHANG Jian, LIU Weixu. Microfluidic Synthesis of UiO-66 Metal-organic Frameworks Modified with Different Functional Groups† [J]. Chem. J. Chinese Universities, 2019, 40(7): 1365. |
| [9] | Jie WANG,Yu ZHANG,Min YU,Jin FANG. Multiple Analyses of Tumor Invasion Based on an Integrated Microfluidic System with Bi-directional Solute Concentration Gradient † [J]. Chem. J. Chinese Universities, 2019, 40(12): 2494. |
| [10] | YUAN Haojun, GAO Wanlei, JING Fengxiang, LIU Songsheng, ZHOU Hongbo, JIA Chunping, JIN Qinghui, ZHAO Jianlong. Novel Microfluidic Droplet Digital PCR Chip for High Sensitive Detection of Nucleic Acid† [J]. Chem. J. Chinese Universities, 2017, 38(7): 1140. |
| [11] | YANG Meiyue, WANG Wei, TIAN Zhiqing, LI Yingying, YUAN Zhi. Preparation of Glycyrrhetinic Acid-modified Sodium Alginate Microgel Spheres for 3D Cell Culture† [J]. Chem. J. Chinese Universities, 2017, 38(2): 326. |
| [12] | ZHAO Tiantian, CHEN Yuqing, ZHANG Min, WANG Yuerong, ZHANG Hongyang, HU Ping. Fabrication of Paper-based Microfluidic Chips and the Application on the Determination of Uric Acid in Serum Based on Gold Nanoparticle-assisted Catalysis† [J]. Chem. J. Chinese Universities, 2016, 37(5): 829. |
| [13] | JIANG Jian, LI Panpan, MA Yingxue, YANG Chunguang, XU Zhangrun. Microfluidic Analysis System Integrated with Liquid-liquid Extraction and Liquid-liquid Waveguide† [J]. Chem. J. Chinese Universities, 2016, 37(4): 648. |
| [14] | FAN Hongliang, ZHANG Tao, JIN Wei, JIN Qinhan. Novel Label-free Assay of Telomerase Based on DNAzyme† [J]. Chem. J. Chinese Universities, 2015, 36(4): 625. |
| [15] | ZHU Lina, ZHU Ying, FANG Qun. Recent Progress of Microfluidic Techniques for Protein Crystallization and Screening† [J]. Chem. J. Chinese Universities, 2014, 35(1): 1. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
