

高等学校化学学报 ›› 2021, Vol. 42 ›› Issue (8): 2412.doi: 10.7503/cjcu20210048
收稿日期:2021-01-22
出版日期:2021-08-10
发布日期:2021-08-05
通讯作者:
肖琦
E-mail:qi.xiao@nnnu.edu.cn
基金资助:
HUANG Shan1, YAO Jiandong1, NING Gan1, XIAO Qi1( ), LIU Yi1,2
), LIU Yi1,2
Received:2021-01-22
Online:2021-08-10
Published:2021-08-05
Contact:
XIAO Qi
E-mail:qi.xiao@nnnu.edu.cn
Supported by:摘要:
基于苯醌类物质静态猝灭石墨烯量子点(GQDs)荧光的特性, 构建了一种利用GQDs荧光探针实时、 高效检测碱性磷酸酶(ALP)活性的新方法. 过氧化氢在辣根过氧化物酶催化作用下产生羟基自由基并将邻苯二酚氧化成邻苯醌, 导致GQDs的荧光猝灭. ALP催化抗坏血酸-2-磷酸反应生成抗坏血酸, 具有较强还原性的抗坏血酸能清除溶液中的过氧化氢和羟基自由基, 抑制邻苯醌的产生, 使GQDs的荧光猝灭效果减弱. 随着ALP活性的增大, GQDs在440 nm处的荧光强度不断增强, 由此建立了一种高效检测ALP活性的新方法. 在最佳实验条件下, 该GQDs荧光探针对ALP活性的检出限为0.084 U/L. 将此方法成功用于人血清中ALP活性的检测, 为与ALP相关疾病的诊断与治疗提供了理论基础.
中图分类号:
TrendMD:
黄珊, 姚建东, 宁淦, 肖琦, 刘义. 石墨烯量子点荧光探针对碱性磷酸酶活性的高效检测. 高等学校化学学报, 2021, 42(8): 2412.
HUANG Shan, YAO Jiandong, NING Gan, XIAO Qi, LIU Yi. Efficient Determination of Alkaline Phosphatase Activity Based on Graphene Quantum Dots Fluorescent Probes. Chem. J. Chinese Universities, 2021, 42(8): 2412.
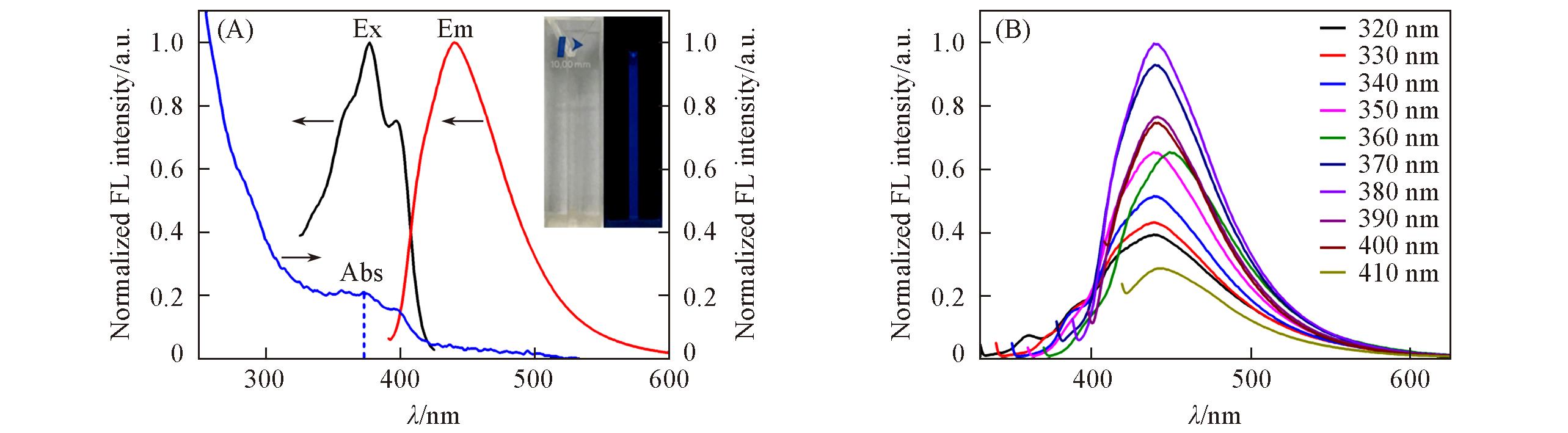
Fig.3 UV?Vis absorption and fluorescence spectra of GQDs(A) and emission spectra of GQDs under different excitation wavelengths(B)Insets in (A) are photographs of GQDs under the irradiation of white light(left) and 365 nm UV lamp(right).
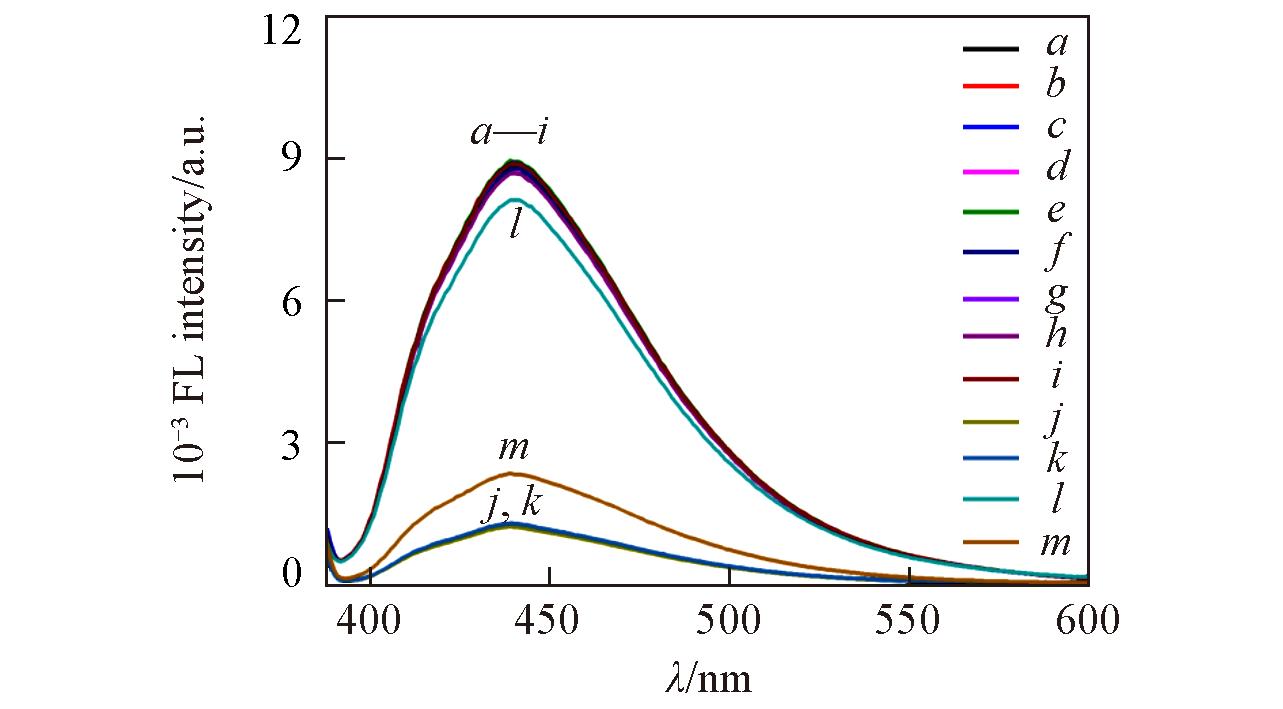
Fig.5 Fluorescence spectra of GQDs(a), GQDs/HRP(b), GQDs/o?DHB(c), GQDs/H2O2(d), GQDs/AAP(e), GQDs/ALP(f), GQDs/AA(g), GQDs/Na3VO4(h),GQDs/HRP/o?DHB(i),GQDs/HRP/H2O2/o?DHB(j), GQDs/AAP/HRP/H2O2/o?DHB(k), GQDs/ALP/AAP/HRP/H2O2/o?DHB(l) and GQDs/Na3VO4/ALP/AAP/HRP/H2O2/o?DHB(m)
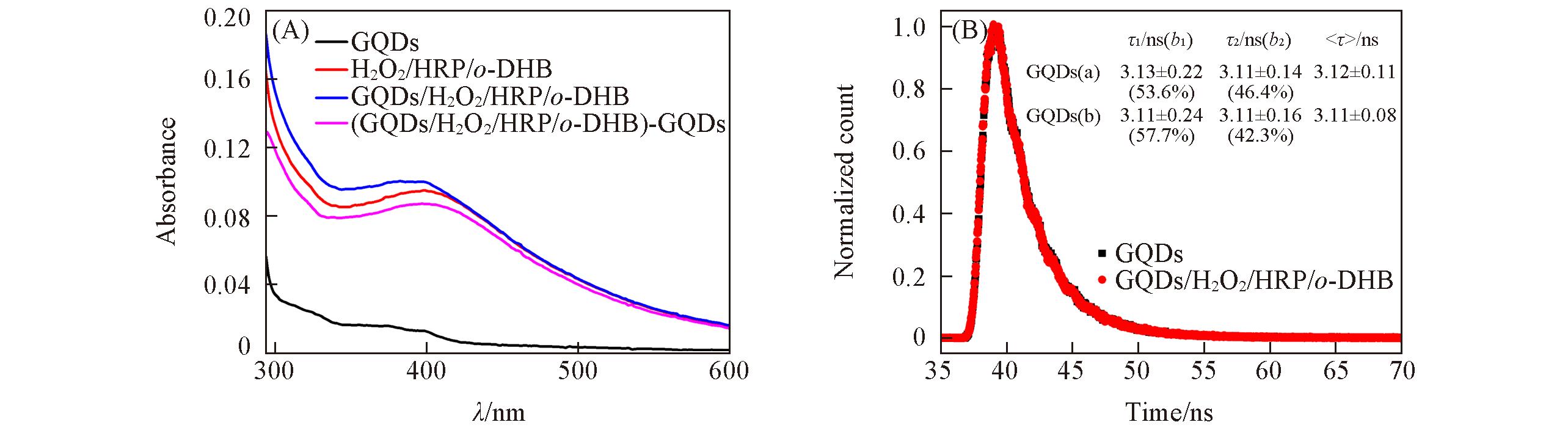
Fig.6 UV?Vis absorption spectra of GQDs, H2O2/HRP/o?DHB, GQDs/H2O2/HRP/o?DHB and (GQDs/H2O2/HRP/o?DHB)?GQDs systems(A), fluorescence decay traces of GQDs and GQDs/H2O2/HRP/o?DHB system(B)Insets in (B) are fluorescence lifetimes data of GQDs(a) and GQDs in GQDs/H2O2/HRP/o-DHB system(b). τ is the fluorescent lifetime of trypsin and b is the normalized pre-exponential factor. The average fluorescence lifetime(<τ>) is calculated according to the equation of <τ> = τibi.
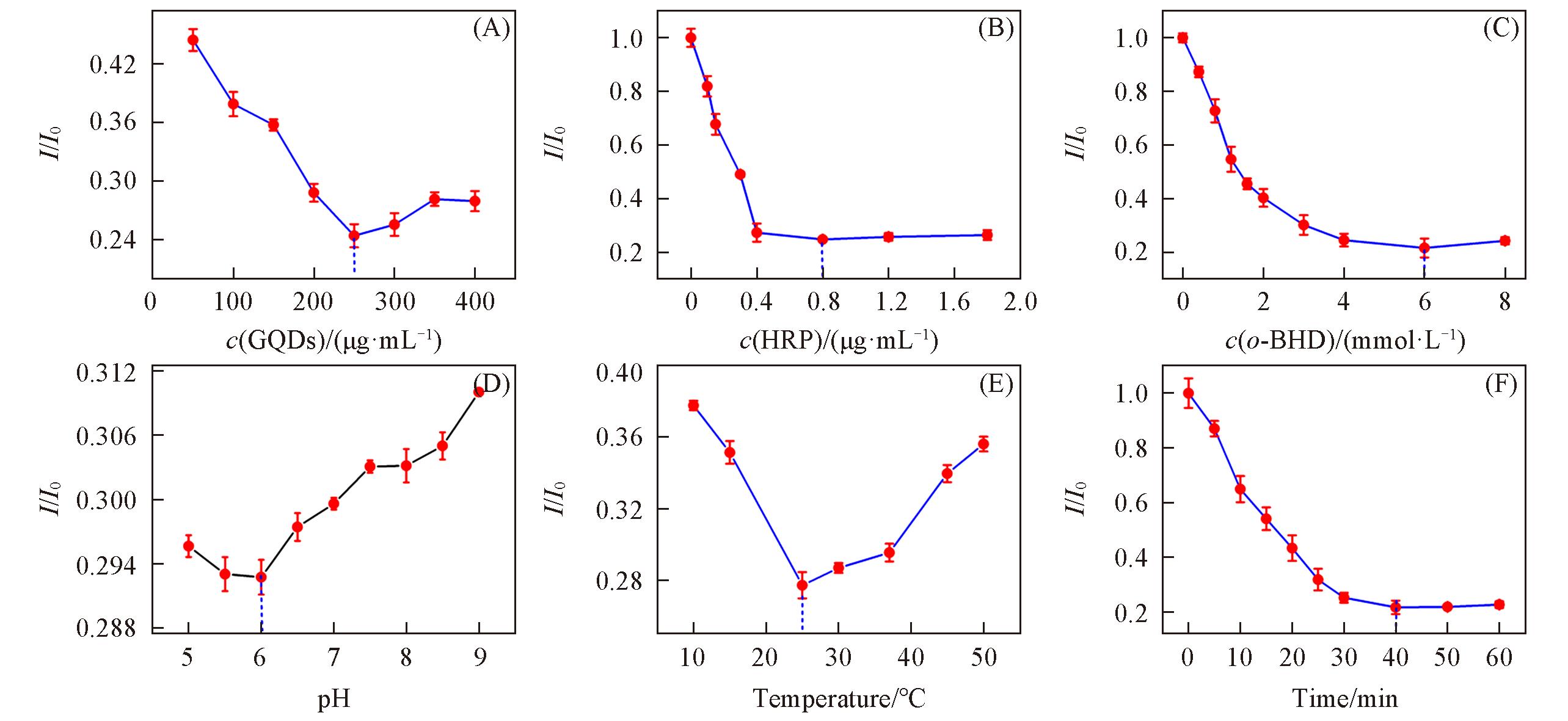
Fig.7 Effects of GQDs amount(A), HRP amount(B), o?DHB concentration(C), pH value(D), reaction temperature(E) and reaction time(F) on the relative fluorescence intensity of GQDs?based fluorescent probe
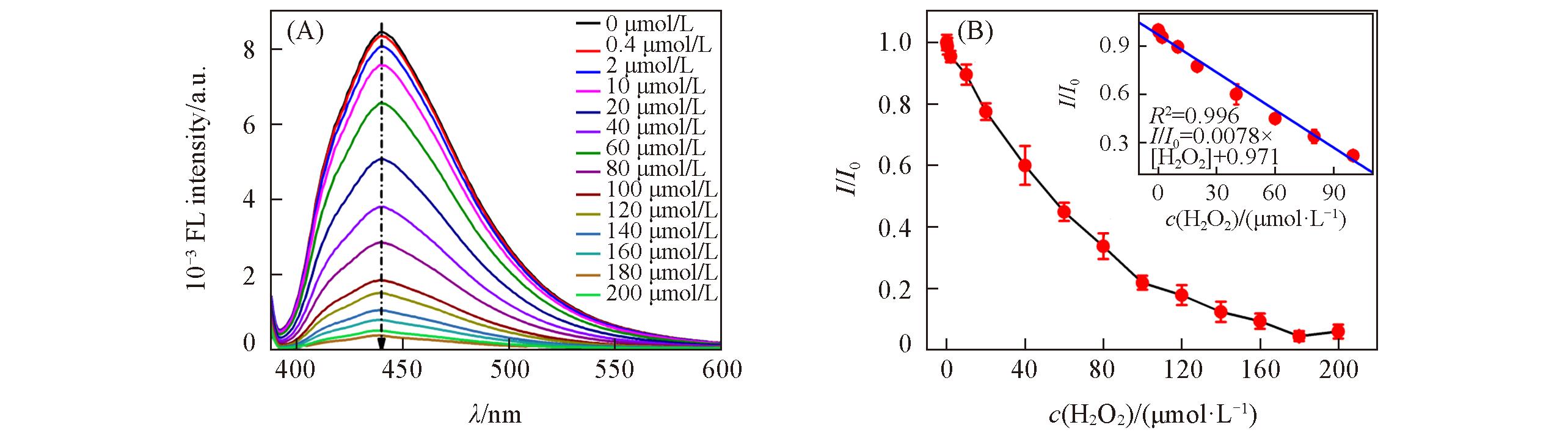
Fig.8 Fluorescence spectra of GQDs?based fluorescent probe with increasing concentration of H2O2(A) and relationship between relative fluorescence intensity ratio I/I0 and concentration of H2O2(B)Inset of (B) is the linear relationship between the relative fluorescence intensity ratio I/I0 and concentration of H2O2 from 0.4 μmol/L to 100 μmol/L.
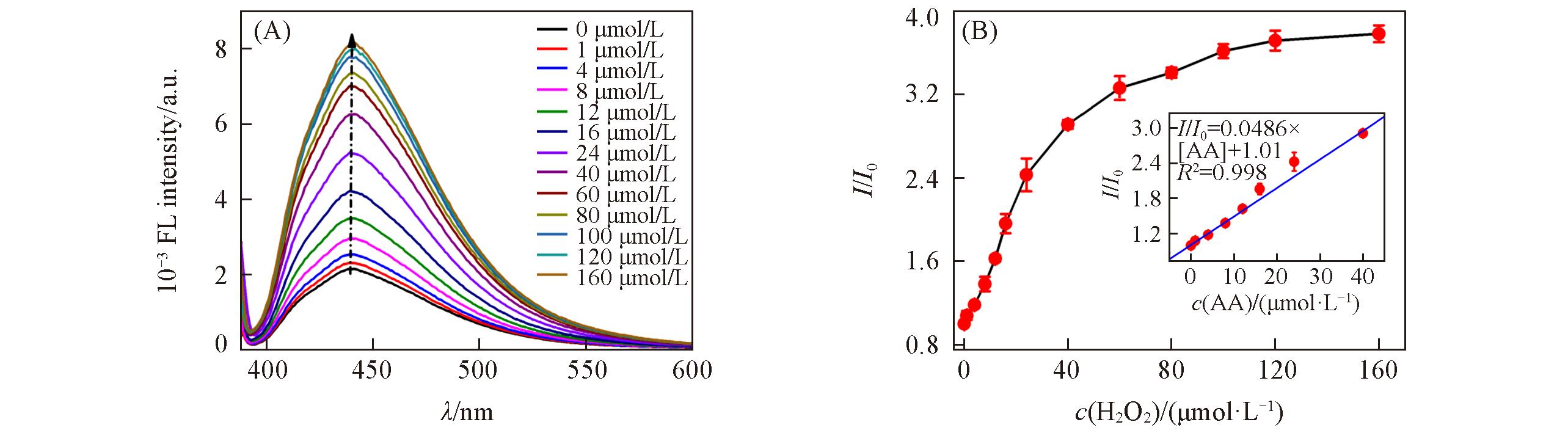
Fig.9 Fluorescence spectra of GQDs?based fluorescent probe with increasing concentration of AA(A) and relationship between relative fluorescence intensity ratio(I/I0) and concentration of AA(B)Inset of (B) is the linear relationship between the relative fluorescence intensity ratio(I/I0) and concentration of AA from 1.0 μmol/L to 40 μmol/L.
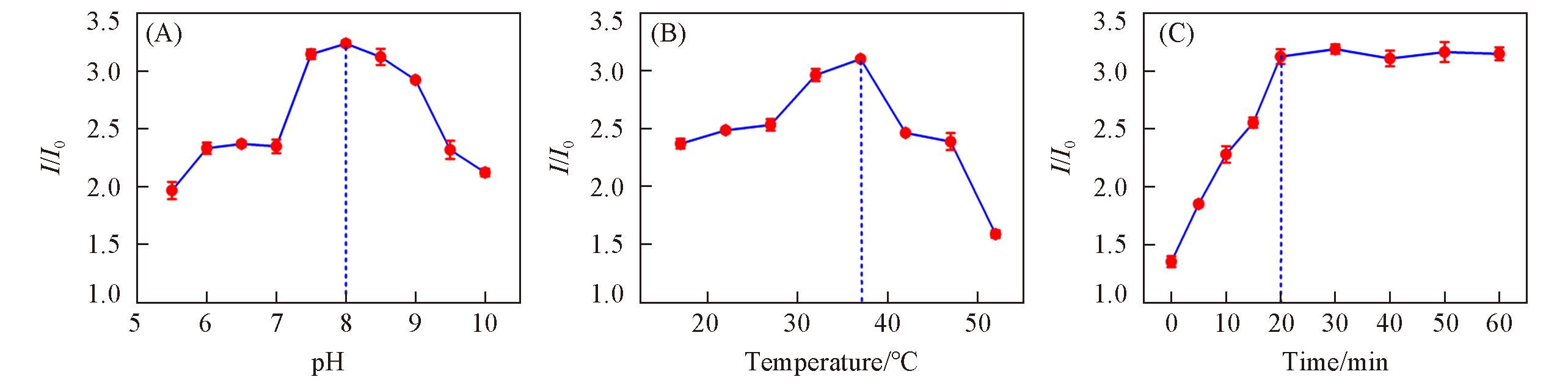
Fig.10 Effects of pH value(A), enzymatic reaction temperature(B) and enzymatic reaction time(C) on the relative fluorescence intensity of GQDs?based fluorescent probe
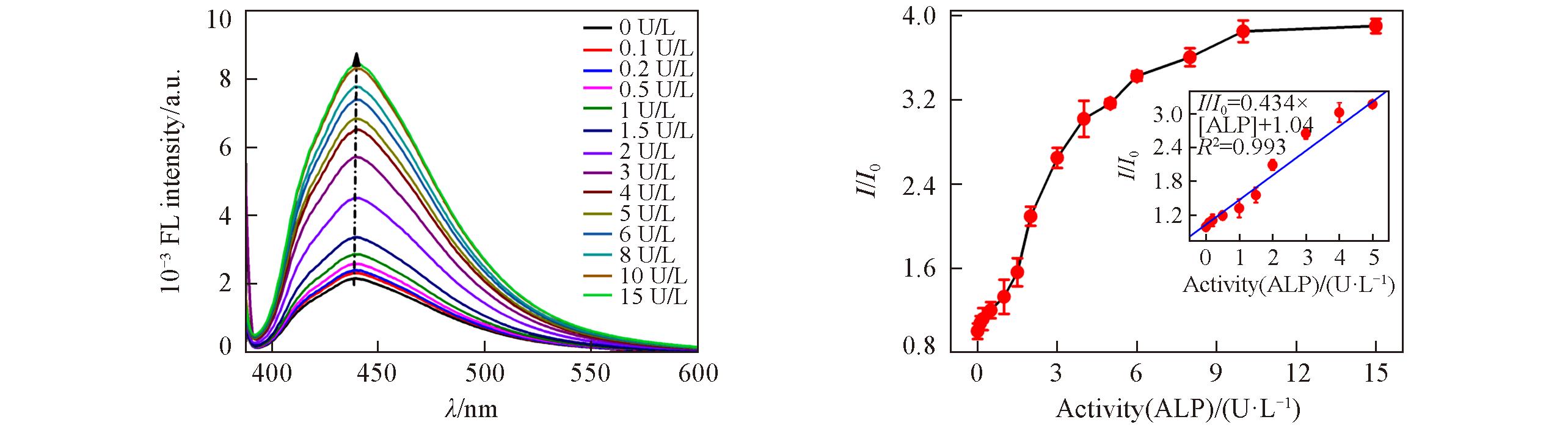
Fig.11 Fluorescence spectra of GQDs?based fluorescent probe with increasing activity of ALP(A) and relationship between relative fluorescence intensity ratio(I/I0) and activity of ALP(B)Inset of (B) is the linear relationship between the relative fluorescence intensity ratio(I/I0) and activity of ALP from 0.1 to 5 U/L.
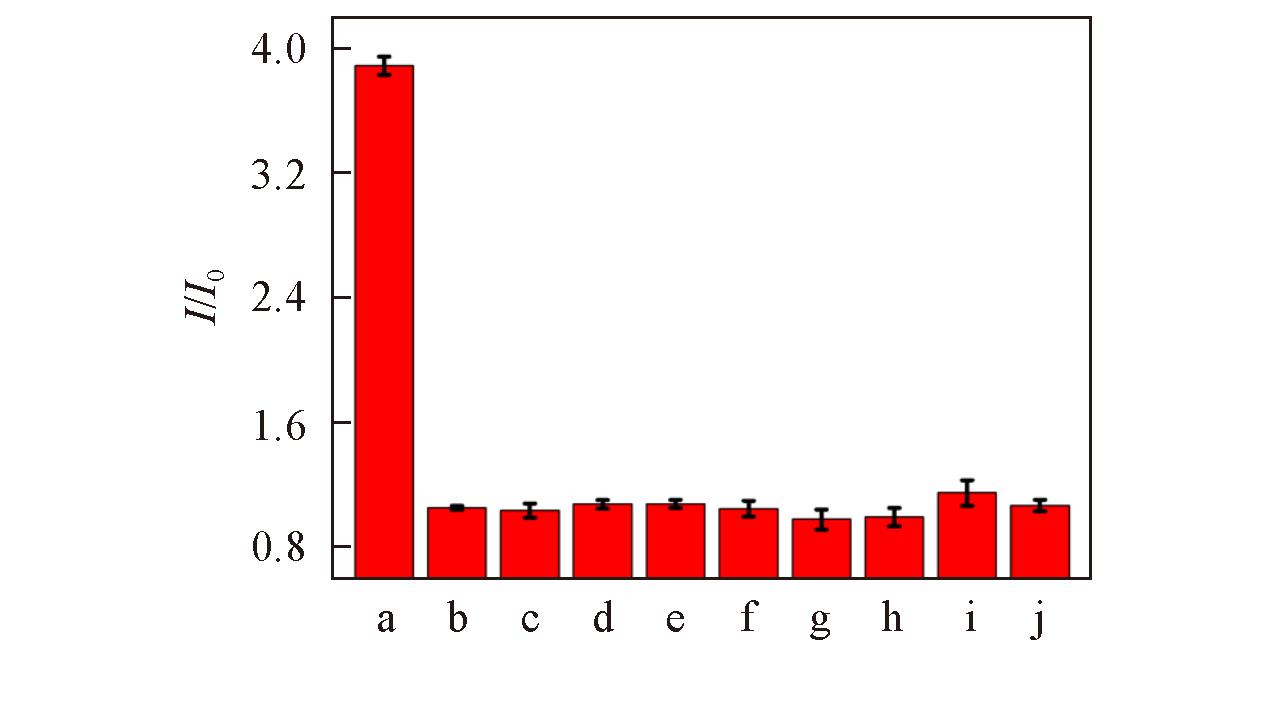
Fig.12 Effects of different enzymes on the relative fluorescence intensity of GQDs?based fluorescent probea. ALP; b. blank; c. Pep; d. TRY; e. Lys; f. Thr; g. HSA; h. BSA; i. GOx; j. AchE.
| Co?existing substance | Δ(I/I0)(%) | Co?existing substance | Δ(I/I0)(%) |
|---|---|---|---|
| L?Aspartic acid | +1.52 | Lactose | +2.74 |
| L?Serine | -3.96 | Galactose | +3.79 |
| L?Glutamate | -4.93 | Maltose | -1.13 |
| L?Lysine | +2.36 | Uric acid | -4.42 |
| L?Phenylalanine | -2.51 | Cholesterol | -0.95 |
| D?Tryptophan | +4.18 | Guanosine | +2.76 |
| L?Tryptophan | -1.61 | Adenosine | +3.59 |
| L?Glutathione oxidized | +1.65 | AlCl3 | +1.04 |
| L?Glycine | +3.32 | NiCl2 | -4.03 |
| L?Arginine | -4.55 | MnSO4 | +3.59 |
| L?Methionine | +0.47 | MgCl2 | +2.42 |
| L?Glutamine | -1.48 | CrCl3 | +1.39 |
| L?Leucine | 1.48 | CdCl2 | -4.29 |
| L?Valine | +1.36 | CaCl2 | -4.56 |
| L?α?Alanine | +4.67 | NaCl | +3.80 |
Table 1 Influences of 0.1 mmol/L co-existing substances on ALP activity evaluation
| Co?existing substance | Δ(I/I0)(%) | Co?existing substance | Δ(I/I0)(%) |
|---|---|---|---|
| L?Aspartic acid | +1.52 | Lactose | +2.74 |
| L?Serine | -3.96 | Galactose | +3.79 |
| L?Glutamate | -4.93 | Maltose | -1.13 |
| L?Lysine | +2.36 | Uric acid | -4.42 |
| L?Phenylalanine | -2.51 | Cholesterol | -0.95 |
| D?Tryptophan | +4.18 | Guanosine | +2.76 |
| L?Tryptophan | -1.61 | Adenosine | +3.59 |
| L?Glutathione oxidized | +1.65 | AlCl3 | +1.04 |
| L?Glycine | +3.32 | NiCl2 | -4.03 |
| L?Arginine | -4.55 | MnSO4 | +3.59 |
| L?Methionine | +0.47 | MgCl2 | +2.42 |
| L?Glutamine | -1.48 | CrCl3 | +1.39 |
| L?Leucine | 1.48 | CdCl2 | -4.29 |
| L?Valine | +1.36 | CaCl2 | -4.56 |
| L?α?Alanine | +4.67 | NaCl | +3.80 |
| Sample | Spiked/(U·L-1) | Found/(U·L-1) | Recovery(%) | RSD(%) |
|---|---|---|---|---|
| Human serum sample 1 | 0 | 1.39—1.45 | — | — |
| 0.5 | 1.90—1.92 | 96.5—100.4 | 2.62 | |
| 3 | 4.42—4.52 | 99.1—103.5 | 3.15 | |
| 5 | 6.31—6.36 | 97.9—98.9 | 0.02 | |
| Human serum sample 2 | 0 | 1.16—1.23 | — | — |
| 0.5 | 1.69—1.72 | 98.5—102.8 | 0.69 | |
| 3 | 4.12—4.27 | 97.5—103.4 | 2.28 | |
| 5 | 5.98—6.12 | 95.5—98.4 | 2.03 | |
| Human serum sample 3 | 0 | 0.87—0.94 | — | — |
| 0.5 | 1.39—1.42 | 97.9—103.7 | 4.09 | |
| 3 | 3.90—4.04 | 100.8—103.4 | 0.46 | |
| 5 | 5.99—6.03 | 101.7—102.4 | 0.59 |
Table 2 Evaluation of ALP activity in human serum samples
| Sample | Spiked/(U·L-1) | Found/(U·L-1) | Recovery(%) | RSD(%) |
|---|---|---|---|---|
| Human serum sample 1 | 0 | 1.39—1.45 | — | — |
| 0.5 | 1.90—1.92 | 96.5—100.4 | 2.62 | |
| 3 | 4.42—4.52 | 99.1—103.5 | 3.15 | |
| 5 | 6.31—6.36 | 97.9—98.9 | 0.02 | |
| Human serum sample 2 | 0 | 1.16—1.23 | — | — |
| 0.5 | 1.69—1.72 | 98.5—102.8 | 0.69 | |
| 3 | 4.12—4.27 | 97.5—103.4 | 2.28 | |
| 5 | 5.98—6.12 | 95.5—98.4 | 2.03 | |
| Human serum sample 3 | 0 | 0.87—0.94 | — | — |
| 0.5 | 1.39—1.42 | 97.9—103.7 | 4.09 | |
| 3 | 3.90—4.04 | 100.8—103.4 | 0.46 | |
| 5 | 5.99—6.03 | 101.7—102.4 | 0.59 |
| Method | Probe | Linear range/(U·L-1) | Detection limit/ (U·L-1) | Samples detection | Ref. |
|---|---|---|---|---|---|
| Fluorimetry | Carbon quantum dots | 3.4—100 | 0.9 | — | [ |
| Electrochemical | CdS@graphene?CoOOH nanosheets | 10—300 | 1.5 | Human serum samples | [ |
| Fluorimetry | Polydopamine/MnO2 nanocomposites | 1—80 | 0.34 | Human serum samples | [ |
| Fluorimetry | Hemicyanine?based near?infrared probe | 10—2000 | 3 | Cell, tissue and living animal | [ |
| Colorimetric | Cu(II)?phenanthroline complex | 0—200 | 1.25 | Human serum samples | [ |
| Fluorimetry | Silver nanocluster | 30—3000 | 5 | Bovine serum samples | [ |
| Fluorimetry | WS2 quantum dots | 0.5—10 | 0.2 | Human serum samples | [ |
| Fluorimetry | Polydopamine nanodots | 1—50 | 0.94 | Human serum samples | [ |
| Fluorimetry | Protein stabilized gold nanocubes | — | 1.616 | Human serum samples | [ |
| Fluorimetry | Electrospun fibrous strips | 0—80 | 1.5 | Human serum samples | [ |
| Fluorimetry | Carbon quantum dots | 4.6—383.3 | 1.4 | — | [ |
| Fluorimetry | GQDs | 0.1—5 | 0.084 | Human serum samples | This work |
Table 3 Evaluation of ALP activity with different methods and different probes
| Method | Probe | Linear range/(U·L-1) | Detection limit/ (U·L-1) | Samples detection | Ref. |
|---|---|---|---|---|---|
| Fluorimetry | Carbon quantum dots | 3.4—100 | 0.9 | — | [ |
| Electrochemical | CdS@graphene?CoOOH nanosheets | 10—300 | 1.5 | Human serum samples | [ |
| Fluorimetry | Polydopamine/MnO2 nanocomposites | 1—80 | 0.34 | Human serum samples | [ |
| Fluorimetry | Hemicyanine?based near?infrared probe | 10—2000 | 3 | Cell, tissue and living animal | [ |
| Colorimetric | Cu(II)?phenanthroline complex | 0—200 | 1.25 | Human serum samples | [ |
| Fluorimetry | Silver nanocluster | 30—3000 | 5 | Bovine serum samples | [ |
| Fluorimetry | WS2 quantum dots | 0.5—10 | 0.2 | Human serum samples | [ |
| Fluorimetry | Polydopamine nanodots | 1—50 | 0.94 | Human serum samples | [ |
| Fluorimetry | Protein stabilized gold nanocubes | — | 1.616 | Human serum samples | [ |
| Fluorimetry | Electrospun fibrous strips | 0—80 | 1.5 | Human serum samples | [ |
| Fluorimetry | Carbon quantum dots | 4.6—383.3 | 1.4 | — | [ |
| Fluorimetry | GQDs | 0.1—5 | 0.084 | Human serum samples | This work |
| 1 | Anas N. A. A., Fen Y. W., Yusof N. A., Omar N. A. S., Daniyal W. M. E. M. M., Ramdzan N. S. M., J. Appl. Phys., 2020, 128, 083105 |
| 2 | Du J. J., Feng B., Dong Y. Q., Zhao M., Yang X. D., Nanoscale, 2020, 12, 9219—9230 |
| 3 | Liu H., Na W. D., Liu Z. P., Chen X. Q., Su X. G., Biosens. Bioelectron., 2017, 92, 229—233 |
| 4 | Xia M., Zhao X. E., Sun J., Zheng Z. J., Sensors Actuat. B: Chem., 2020, 319, 128321 |
| 5 | Wang Q., Li L. F., Wang X. D., Dong C., Shuang S. M., Talanta,2020, 219, 121180 |
| 6 | Zhao X. E., Lei C. H., Wang Y. H., Qu F., Zhu S. Y., Wang H., You J. M., RSC Adv., 2016, 6, 72670—72675 |
| 7 | Xu S. M., Zhang F. M., Xu L. B., Liu X., Ma P. Y., Sun Y., Wang X. H., Song D. Q., Sensors Actuat. B: Chem., 2018, 273, 1015—1021 |
| 8 | Li X., Zhu S. J., Xu B., Ma K., Zhang J. H., Yang B., Tian W. J., Nanoscale, 2013, 5, 7776—7779 |
| 9 | He T., Qi L., Zhang J., Huang Y. L., Zhang Z. Q., Sensors Actuat. B: Chem., 2015, 215, 24—29 |
| 10 | Qu Z. Y., Li N., Na W. D., Su X. G., Talanta,2019, 192, 61—68 |
| 11 | Zhu Y. H., Wang G. F., Jiang H., Chen L., Zhang X. J., Chem. Commun., 2015, 51, 948—951 |
| 12 | Huang S., Yao J. D., Chu X., Ning G., Zhou Z. Q., Liu Y., Xiao Q., Microchim. Acta, 2020, 187, 271 |
| 13 | Liu J. J., Tang D. S., Chen Z. T., Yan X. M., Zhong Z., Kang L. T., Yao J. N., Biosens. Bioelectron., 2017, 94, 271—277 |
| 14 | Li G. L., Fu H. L., Chen X. J., Gong P. W., Chen G., Xia L., Wang H., You J. M., Wu Y. N., Anal. Chem., 2016, 88, 2720—2726 |
| 15 | Hayat A., Bulbul G., Andreescu S., Biosens. Bioelectron., 2014, 56, 334—339 |
| 16 | Liu Y. Q., Xiong E. H., Li X. Y., Li J. J., Zhang X. H., Chen J. H., Biosens. Bioelectron., 2017, 87, 970—975 |
| 17 | Kong W. S., Tan Q. Q., Guo H. Y., Sun H., Qin X., Qu F. L., Microchim. Acta, 2019, 186, 73 |
| 18 | Ruan C. M., Wang W., Gu B. H., Anal. Chem., 2006, 78, 3379—3384 |
| 19 | Hu Q., He M. H., Mei Y. Q., Feng W. J., Jing S., Kong J. M., Zhang X. J., Talanta, 2017, 163, 146—152 |
| 20 | Xiao T., Sun J., Zhao J. H., Wang S., Liu G. Y., Yang X. R., ACS Appl. Mater. Interfaces, 2018, 10, 6560—6569 |
| 21 | Xue Q., Cao X. Y., Zhang C. L., Xian Y. Z., Microchim. Acta, 2018, 185, 231 |
| 22 | Wang H. B., Li Y., Chen Y., Zhang Z. P., Gan T., Liu Y. M., Microchim. Acta, 2018, 185, 102 |
| 23 | Yin B. C., Ma J. L., Ye B. C., Wu X., Anal. Chem., 2016, 88, 9219—9225 |
| 24 | Sharma A. K., Pandey S., Khan M. S., Wu H. F., Sensors Actuat. B: Chem., 2018, 259, 83—89 |
| 25 | Hu L. Z., Zhang Q., Gan X. Y., Lin S. L., Han S., Zhang Z. C., Microchim. Acta, 2018, 185, 390 |
| 26 | Qian Z. S., Chai L. J., Tang C., Huang Y. Y., Chen J. R., Feng H., Anal. Chem., 2015, 87, 2966—2973 |
| 27 | Feng J., Ju Y. Y., Liu J. J., Zhang H. G., Chen X. G., Anal. Chim. Acta, 2015, 854, 153—160 |
| 28 | Huang S., Yao J. D., Chu X., Liu Y., Xiao Q., Zhang Y., J. Agric. Food Chem., 2019, 67, 11244—11255 |
| 29 | Qu F. L., Pei H. M., Kong R. M., Zhu S. Y., Xia L., Talanta, 2017, 165, 136—142 |
| 30 | Lin L. X., Zhang S. W., Chem. Commun., 2012, 48, 10177—10179 |
| 31 | Li P. J., Hong Y. Y., Feng H. T., Li S. F. Y., J. Mater. Chem. B, 2017, 5, 2979—2988 |
| 32 | Huang J. L., Liu F. J., Wang T. T., Liu C. E., Zheng F. Y., Wang Z. H., Li S. X., Chem. J. Chinese Universities, 2020, 41(7), 1513—1520(黄加玲, 刘凤娇, 王婷婷, 刘翠娥, 郑凤英, 王振红, 李顺兴.高等学校化学学报, 2020, 41(7), 1513—1520) |
| 33 | Huang S., Wang L. M., Huang C. S., Hu B. Q., Su W., Xiao Q., Microchim. Acta, 2016, 183, 1855—1864 |
| 34 | Zhan Y. J., Zeng Y. B., Li L., Luo F., Qiu B., Lin Z. Y., Guo L. H., ACS Sens., 2019, 4, 1252—1260 |
| 35 | Tang C., Qian Z. S., Huang Y. Y., Xu J. M., Ao H., Zhao M. Z., Zhou J., Chen J. R., Feng H., Biosens. Bioelectron., 2016, 83, 274—280 |
| 36 | Li S. J., Li C. Y., Li Y. F., Fei J. J., Wu P., Yang B., Ouyang J., Nie S. X., Anal. Chem., 2017, 89, 6854—6860 |
| 37 | Zhao L., Xie S. Z., Song X. J., Wei J. J., Zhang Z., Li X. H., Biosens. Bioelectron., 2017, 91, 217—224 |
| [1] | 何贝贝, 杨葵华, 吕瑞. 锰-铜双金属层状硅酸盐纳米酶的构筑及类酶活性[J]. 高等学校化学学报, 2022, 43(8): 20220150. |
| [2] | 王瑞娜, 孙瑞粉, 钟添华, 池毓务. 大尺寸石墨烯量子点组装体的制备及电化学发光性能[J]. 高等学校化学学报, 2022, 43(8): 20220161. |
| [3] | 赵永梅, 穆叶舒, 洪琛, 罗稳, 田智勇. 双萘酰亚胺衍生物用于检测水溶液中的苦味酸[J]. 高等学校化学学报, 2022, 43(3): 20210765. |
| [4] | 唐倩, 但飞君, 郭涛, 兰海闯. 喹啉酮-香豆素类Hg2+比色荧光探针的合成及应用[J]. 高等学校化学学报, 2022, 43(2): 20210660. |
| [5] | 王迪, 钟克利, 汤立军, 侯淑华, 吕春欣. 席夫碱共价有机框架的合成及对I ‒ 的识别[J]. 高等学校化学学报, 2022, 43(10): 20220115. |
| [6] | 李安然, 赵冰, 阚伟, 宋天舒, 孔祥东, 卜凡强, 孙立, 殷广明, 王丽艳. 基于菲并咪唑的ON⁃OFF⁃ON双比色荧光探针及细胞成像[J]. 高等学校化学学报, 2021, 42(8): 2403. |
| [7] | 杨新杰, 赖艳琼, 李秋旸, 张艳丽, 王红斌, 庞鹏飞, 杨文荣. 基于环状DNA-银纳米簇荧光探针对微囊藻毒素-LR的传感检测[J]. 高等学校化学学报, 2021, 42(12): 3600. |
| [8] | 谌委菊, 陈诗雅, 薛曹叶, 刘波, 郑晶. 缺氧响应荧光探针的成像及治疗应用[J]. 高等学校化学学报, 2021, 42(11): 3433. |
| [9] | 黄加玲,刘凤娇,王婷婷,刘翠娥,郑凤英,王振红,李顺兴. 氮硫共掺杂碳量子点对胃液pH值的精确检测[J]. 高等学校化学学报, 2020, 41(7): 1513. |
| [10] | 李明,崔小倩,汪璇,李在均. 双亲性石墨烯量子点的合成及对L-薄荷醇的缓释作用[J]. 高等学校化学学报, 2020, 41(2): 324. |
| [11] | 吴倩, 程丹, 吕芸, 袁林, 张晓兵. 大斯托克斯位移远红光至近红外荧光探针用于检测肝损伤过程中过氧化亚硝酸盐的动态变化[J]. 高等学校化学学报, 2020, 41(11): 2426. |
| [12] | 王金金, 戚少龙, 杜建时, 杨清彪, 宋岩, 李耀先. 苯并噻唑类荧光探针的合成及对N2H4·H2O和HS |
| [13] | 张勇,申城,幸志荣,陈归柒,卢资,侯志兵,陈雪梅. 可视化检测次氯酸的苯并咪唑类荧光增强型探针[J]. 高等学校化学学报, 2019, 40(12): 2480. |
| [14] | 张莹莹,黄译文,赵冰,王丽艳,宋波. Cr 3+比色荧光探针的合成及细胞成像应用[J]. 高等学校化学学报, 2019, 40(12): 2486. |
| [15] | 喻照川, 马文辉, 吴涛, 问婧, 张永, 王丽艳, 初红涛. B,N,S共掺杂石墨烯量子点的制备及对Fe 3+和H2P |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||