

高等学校化学学报 ›› 2020, Vol. 41 ›› Issue (11): 2335.doi: 10.7503/cjcu20200323
• 庆祝《高等学校化学学报》复刊40周年专栏 • 上一篇 下一篇
赵关芳1,2, 邹天一1,2, 程思航1,2, 于洋1,2, 王慧利1, 王宏达1,2( )
)
收稿日期:2020-06-04
出版日期:2020-11-10
发布日期:2020-11-06
通讯作者:
王宏达
E-mail:hdwang@ciac.ac.cn
基金资助:
ZHAO Guanfang1,2, ZOU Tianyi1,2, CHENG Sihang1,2, YU Yang1,2, WANG Huili1, WANG Hongda1,2( )
)
Received:2020-06-04
Online:2020-11-10
Published:2020-11-06
Contact:
WANG Hongda
E-mail:hdwang@ciac.ac.cn
Supported by:摘要:
对生物大分子复合物的研究和结构分析对于全面了解其功能和生物学意义至关重要. 冷冻电子显微镜在提供生物大分子结构及大分子分布等方面起到重要的作用. 近年来, 冷冻电子显微镜的硬件和软件的发展进一步提高了冷冻电子显微镜的有效性, 使其对各种生物结构、 蛋白质结构的解析更加准确快捷. 但是, 对于生物系统来说, 蛋白质和大分子复合物等均处于复杂的生理环境中, 因此原位检测生物分子的三维结构对于生物体系和结构生物学具有重要意义. 冷冻电子断层扫描作为一种功能强大的技术, 可以无需标记直接通过冷冻样品的固有衬度识别生物大分子的结构, 并且可在原位生理环境中对生物分子进行纳米级分辨率的三维成像. 本文综述了与冷冻电子断层扫描相关的样品制备和数据处理技术, 并总结了冷冻电子断层扫描技术在分离的大分子复合物和整个细胞或组织中的生物学应用.
中图分类号:
TrendMD:
赵关芳, 邹天一, 程思航, 于洋, 王慧利, 王宏达. 原位冷冻电子断层扫描的发展及在生物研究方面的应用. 高等学校化学学报, 2020, 41(11): 2335.
ZHAO Guanfang, ZOU Tianyi, CHENG Sihang, YU Yang, WANG Huili, WANG Hongda. Developments of in situ Cryo-electron Tomography for Biological Applications. Chem. J. Chinese Universities, 2020, 41(11): 2335.
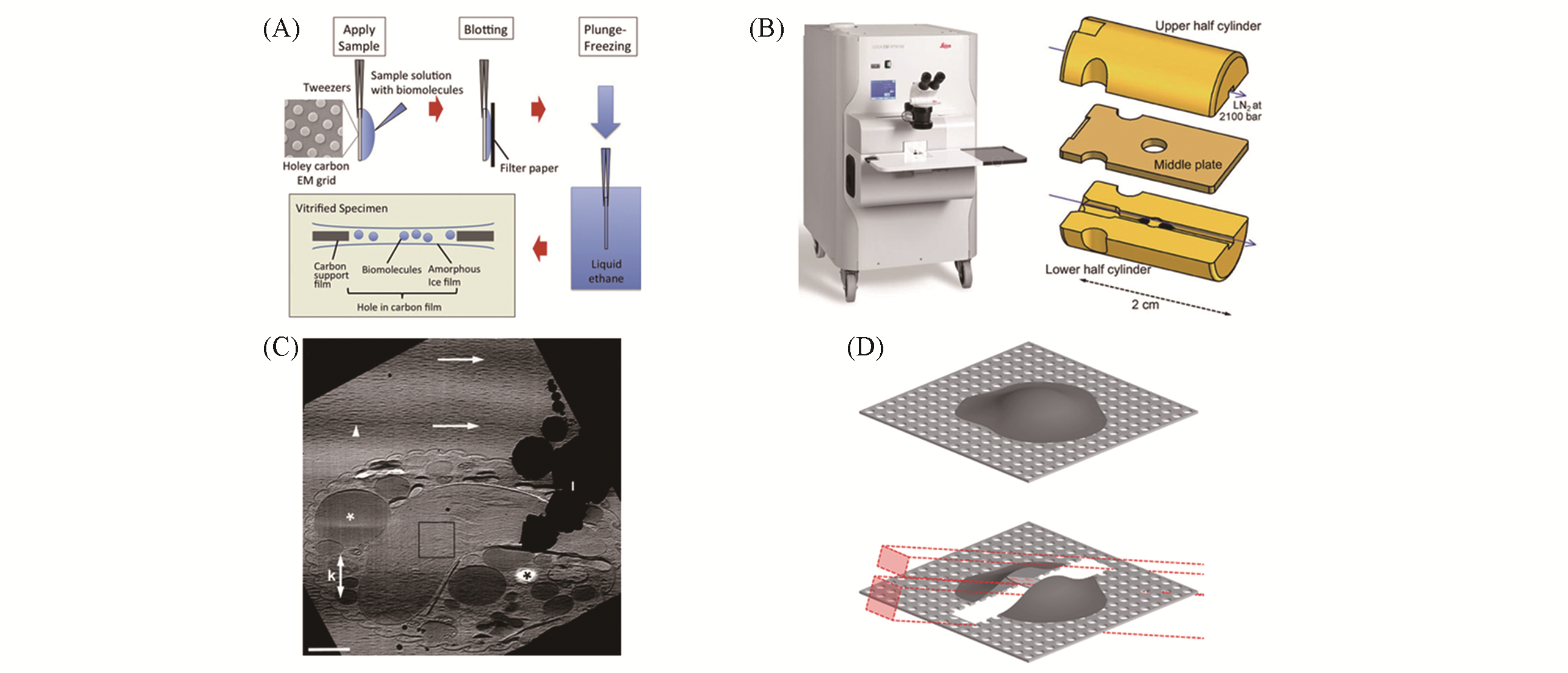
Fig.1 Methods of preparing samples suitable for cryo?ET(A) Samples were prepared by plunge freezing[32]: sample solution is added to the carbon support films and using filter paper to remove excess solution from one or both sides; grid is rapidly plunged into cryogen precooled at liquid nitrogen temperature and imaged with cryo?ET; (B) schematic diagram of high pressure freezing[48]; (C) tomographic slice of a mitotic Hela S3 cell and the oblique striations of the image intensity is the knife marks[39]; (D) vitrified sample is micromachined using FIB milling and the patterns(red boxes) are chosen depending on the milling geometry[49].(A) Copyright 2017, Elsevier; (B) Copyright 2014, Springer Science Business Media; (C) Copyright 2005, Elsevier; (D) Copyright 2013, Elsevier.
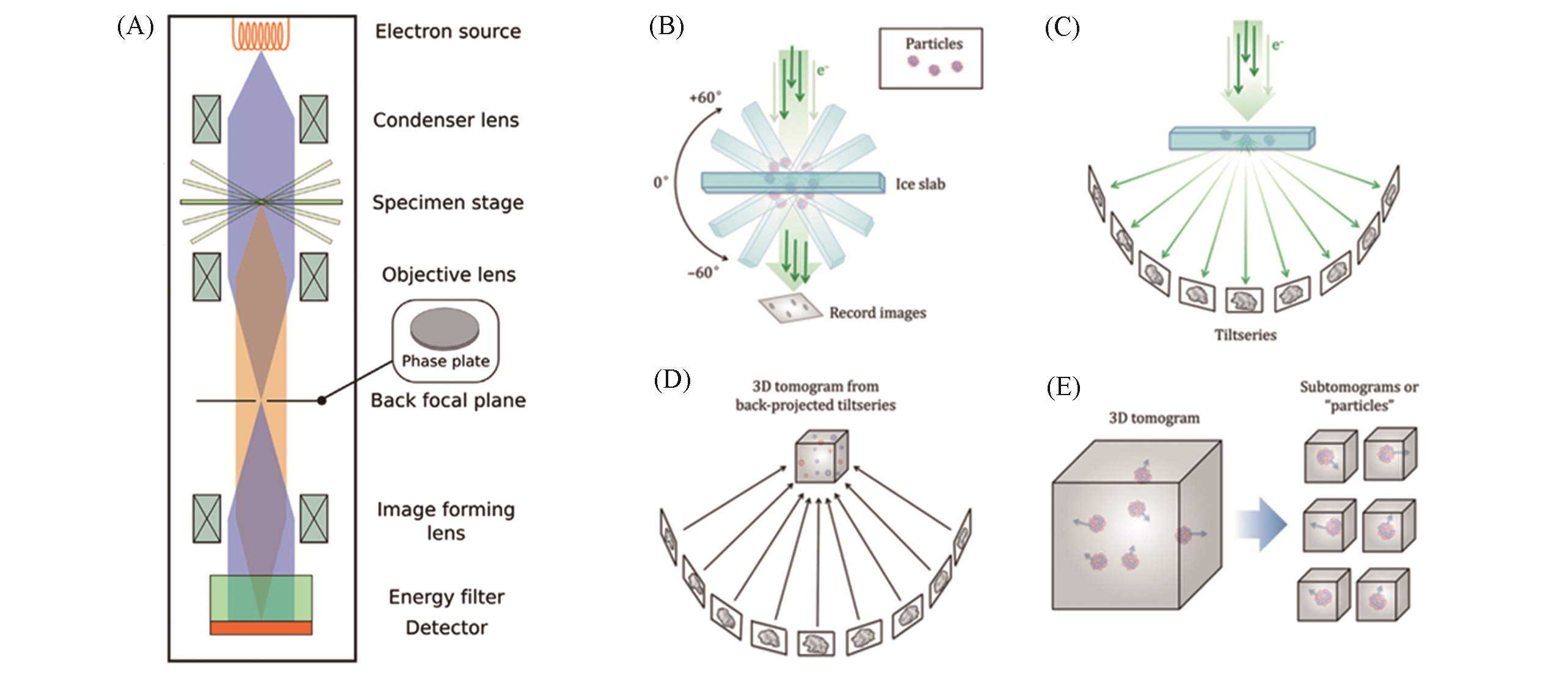
Fig.2 Principles of data collection and processing in cryo?ET[14,56](A) Schematic diagram of cryo?EM; (B) samples embedded in ice is tilted by a certain range of angles and record the projec?ted images at each angle; (C) the tilt series of projected images collected around a common axis; (D) the projected images in the tilt series are aligned with their common axis through calculation and reconstructed into a 3D tomogram by weighted projection or other method; (E) sub?tomograms representing 3D view of single macromolecules can be extracted from the reconstructed tomogram, and then aligned and averaged. (A) Copyright 2015, Elsevier; (B—E) Copyright 2017, Springer Nature.
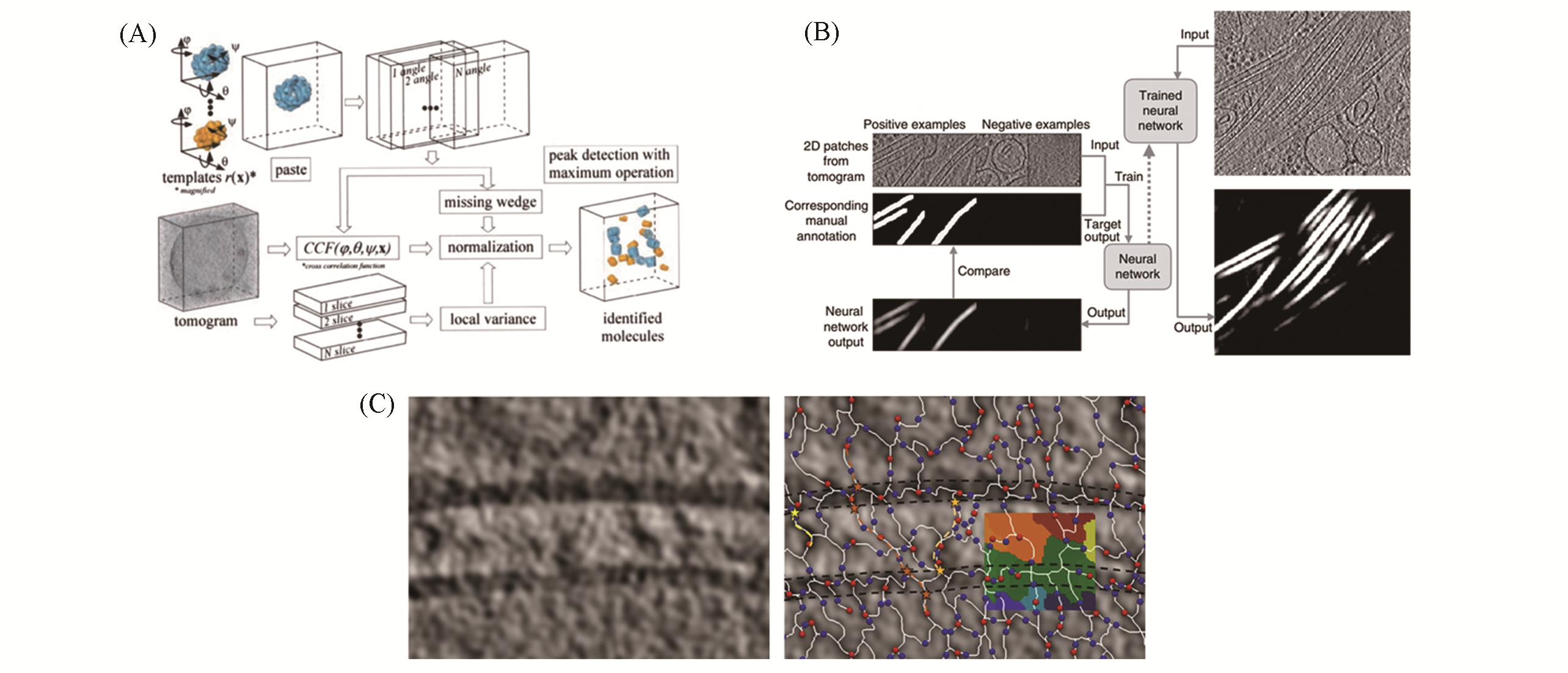
Fig.3 Procedures to identify structures of interest in tomograms(A) Schematic flow diagram of template matching showing the detection and identification strategy[62]; (B) training and automated annotation process of convolutional neural networks for identifying features of interest in tomograms[65]; (C) tracing density at a synapes from rodent cerebrocortical synaptosomal preparation through Pyseg without templates[68]: tomographic slice(left), Morce complex(right).(A) Copyright 2002, The National Academy of Sciences; (B) Copyright 2017, Springer Nature; (C) Copyright 2020, Springer Nature.
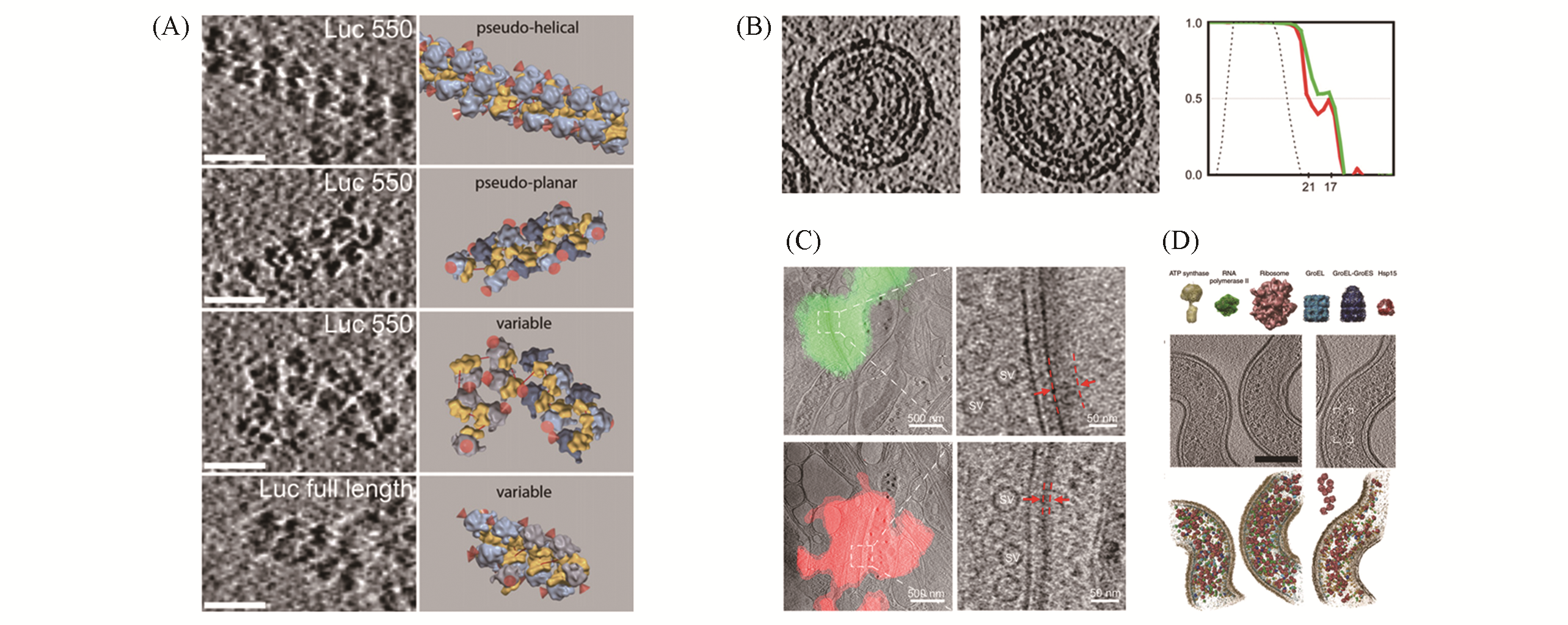
Fig.4 Some biological applications of cryo?ET(A) Tomographic slices of polysome organizations(left) and the corresponding isosurface of single ribosomes(right)[78]; (B) immature HIV virus: tomographic slices(left) and fourier shell correlation plot before (red) and after(green) CTF correction(right)[80]; (C) identifyexcitatory and inhibitory synapses by combining cryo?ET and fluorescence microscope. Tomographic slices of excita?tory(green fluorescence) and inhibitory(red fluorescence) synapses(left). Zoom?in views of the boxed area(right) show the differnce between excitatory and inhibitory synapses(red dashed lines)[84]; (D) template matching in L. interrogans cells[88](top: template library of proteins; middle: representative subvolume of cells; bottom: corresponding surface?rendered models).(A) Copyright 2009, Elsevier; (B) Copyright 2009, National Academy of Sciences; (C) Copyright 2018, Society for Neuroscience; (D) Copyright 2009, Springer Nature.
| 1 | Asano S., Engel B. D., Baumeister W., J. Mol. Biol., 2016, 428(2, Part A), 332―343 |
| 2 | Smyth M. S., Martin J. H., Molecular Pathology: MP, 2000, 53(1), 8―14 |
| 3 | Sprangers R., Velyvis A., Kay L. E., Nat. Methods, 2007, 4(9), 697―703 |
| 4 | Shashkova S., Leake M. C., Biosci. Rep., 2017, 37(4), BSR20170031 |
| 5 | Toca⁃Herrera J. L., ChemSusChem, 2019, 12(3), 603―611 |
| 6 | Blanc F., Isabet T., Benisty H., Sweeney H. L., Cecchini M., Houdusse A., Proc. Nat. Acad. Sci. USA, 2018, 115(24), 6213 |
| 7 | Latychevskaia T., Fink H. W., Opt. Express, 2018, 26(23), 30991―31017 |
| 8 | Wang H., Hao X., Shan Y., Jiang J., Cai M., Shang X., Ultramicroscopy, 2010, 110(4), 305―312 |
| 9 | Zhao W., Tian Y., Cai M., Wang F., Wu J., Gao J., Liu S., Jiang J., Jiang S., Wang H., PLoS One, 2014, 9(5), e91595 |
| 10 | Jing Y., Cai M., Xu H., Zhou L., Yan Q., Gao J., Wang H., Nanoscale, 2018, 10(16), 7457―464 |
| 11 | Yan Q., Lu Y., Zhou L., Chen J., Xu H., Cai M., Shi Y., Jiang J., Xiong W., Gao J., Wang H., Proc. Natl. Acad. Sci. USA, 2018, 115(27), 7033―7038 |
| 12 | Alberts B., Cell, 1998, 92(3), 291―294 |
| 13 | Wagner J., Schaffer M., Fernández⁃Busnadiego R., FEBS Lett., 2017, 591(17), 2520―2533 |
| 14 | Galaz⁃Montoya J. G., Ludtke S. J., Biophysics Reports, 2017, 3(1), 17―35 |
| 15 | Harapin J., Eibauer M., Medalia O., Structure, 2013, 21(9), 1522―1530 |
| 16 | Dierksen K., Typke D., Hegerl R., Baumeister W., Ultramicroscopy, 1993, 49(1), 109―120 |
| 17 | Grimm R., Bärmann M., Häckl W., Typke D., Sackmann E., Baumeister W., Biophys. J., 1997, 72(1), 482―489 |
| 18 | Schaffer M., Engel B. D., Wehmer M., Albert S., Sakata E., Rast A., Nickelsen J., Plitzko J. M., Baumeister W., Microsc. Microanal., 2016, 22(S3), 72―73 |
| 19 | von Appen A., Beck M., J. Mol. Biol., 2016, 428(10, Part A), 2001―2010 |
| 20 | Siegmund S. E., Grassucci R., Carter S. D., Barca E., Farino Z. J., Juanola⁃Falgarona M., Zhang P., Tanji K., Hirano M., Schon E. A., Frank J., Freyberg Z., iScience, 2018, 6, 83―91 |
| 21 | Vijayakrishnan S., McElwee M., Loney C., Rixon F., Bhella D., bioRxiv, 2020, 2020.02.06.936948 |
| 22 | Melia C. E., Bharat T. A. M., Biochim. Biophys. Acta, 2018, 1866(9), 973―981 |
| 23 | Maimon T., Elad N., Dahan I., Medalia O., Structure, 2012, 20(6), 998―1006 |
| 24 | Lewis A. J., Genoud C., Pont M., van de Berg W. D. J., Frank S., Stahlberg H., Shahmoradian S. H., Al⁃Amoudi A., Curr. Opin. Struct. Biol., 2019, 58, 138―148 |
| 25 | Böhm J., Frangakis A. S., Hegerl R., Nickell S., Typke D., Baumeister W., Proc. Nat. Acad. Sci. USA, 2000, 97(26), 14245 |
| 26 | Turoňová B., Schur F. K. M., Wan W., Briggs J. A. G., J. Struct. Biol., 2017, 199(3), 187―195 |
| 27 | Briggs J. A. G., Curr. Opin. Struct. Biol., 2013, 23(2), 261―267 |
| 28 | Chen Y., Pfeffer S., Fernández José J., Sorzano Carlos Oscar S., Förster F., Structure, 2014, 22(10), 1528―1537 |
| 29 | Taylor K., Glaeser R., Science(New York), 1975, 186, 1036―1037 |
| 30 | Taylor K. A., Glaeser R. M., J. Ultrastruct. Res., 1976, 55(3), 448―456 |
| 31 | Dubochet J., Adrian M., Chang J. J., Homo J. C., Lepault J., McDowall A. W., Schultz P., Q. Rev. Biophys., 1988, 21(2), 129―228 |
| 32 | Murata K., Wolf M., Biochimica et Biophysica Acta(BBA)⁃General Subjects, 2018, 1862(2), 324―334 |
| 33 | Brüggeller P., Mayer E., Nature, 1980, 288(5791), 569―571 |
| 34 | Studer D., Humbel B. M., Chiquet M., Histochem. Cell Biol., 2008, 130(5), 877―889 |
| 35 | Bellare J. R., Davis H. T., Scriven L. E., Talmon Y., J. Electron Microsc. Tech., 1988, 10(1), 87―111 |
| 36 | Iancu C. V., Tivol W. F., Schooler J. B., Dias D. P., Henderson G. P., Murphy G. E., Wright E. R., Li Z., Yu Z., Briegel A., Gan L., He Y., Jensen G. J., Nat. Protoc., 2006, 1(6), 2813―2819 |
| 37 | Kanno H., Speedy R. J., Angell C. A., Science, 1975, 189(4206), 880 |
| 38 | Kudryashev M., Castaño⁃Díez D., Stahlberg H., Computational and Structural Biotechnology Journal, 2012, 1(2), e201207002 |
| 39 | Al⁃Amoudi A., Studer D., Dubochet J., J. Struct. Biol., 2005, 150(1), 109―121 |
| 40 | Eltsov M., MacLellan K. M., Maeshima K., Frangakis A. S., Dubochet J., Proc. Nat. Acad. Sci. USA, 2008, 105(50), 19732 |
| 41 | Marko M., Hsieh C., Schalek R., Frank J., Mannella C., Nat. Methods, 2007, 4(3), 215―217 |
| 42 | Hayles M. F., Matthijs de Winter D. A., Schneijdenberg C. T. W. M., Meeldijk J. D., Luecken U., Persoon H., de Water J., de Jong F., Humbel B. M., Verkleij A. J., J. Struct. Biol., 2010, 172(2), 180―190 |
| 43 | Rigort A., Bäuerlein F. J. B., Leis A., Gruska M., Hoffmann C., Laugks T., Böhm U., Eibauer M., Gnaegi H., Baumeister W., Plitzko J. M., J. Struct. Biol., 2010, 172(2), 169―179 |
| 44 | Schaffer M., Engel B. D., Laugks T., Mahamid J., Plitzko J. M., Baumeister W., Bio⁃protocol, 2015, 5(17), e1575 |
| 45 | Hsieh C., Schmelzer T., Kishchenko G., Wagenknecht T., Marko M., J. Struct. Biol., 2014, 185(1), 32―41 |
| 46 | Harapin J., Börmel M., Sapra K. T., Brunner D., Kaech A., Medalia O., Nat. Methods, 2015, 12(7), 634―636 |
| 47 | Schaffer M., Pfeffer S., Mahamid J., Kleindiek S., Laugks T., Albert S., Engel B. D., Rummel A., Smith A. J., Baumeister W., Plitzko J. M., Nat. Methods, 2019, 16(8), 757―762 |
| 48 | Kaech A., Ziegler U., Electron Microscopy, Humana Press, Totowa, NJ, 2014, 151—171 |
| 49 | Villa E., Schaffer M., Plitzko J. M., Baumeister W., Curr. Opin. Struct. Biol., 2013, 23(5), 771―777 |
| 50 | Kühlbrandt W., Science, 2014, 343(6178), 1443 |
| 51 | Faruqi A. R., Henderson R., Curr. Opin. Struct. Biol., 2007, 17(5), 549―555 |
| 52 | Li X., Mooney P., Zheng S., Booth C. R., Braunfeld M. B., Gubbens S., Agard D. A., Cheng Y., Nat. Methods, 2013, 10(6), 584―590 |
| 53 | Fernández J. J., Li S., Crowther R. A., Ultramicroscopy, 2006, 106(7), 587―596 |
| 54 | Danev R., Nagayama K., Ultramicroscopy, 2001, 88(4), 243―252 |
| 55 | Danev R., Buijsse B., Khoshouei M., Plitzko J. M., Baumeister W., Proc. Nat. Acad. Sci. USA, 2014, 111(44), 15635 |
| 56 | Dubrovsky A., Sorrentino S., Harapin J., Sapra K. T., Medalia O., Arch. Biochem. Biophys., 2015, 58178―85 |
| 57 | Nickell S., Förster F., Linaroudis A., Net W. D., Beck F., Hegerl R., Baumeister W., Plitzko J. M., J. Struct. Biol., 2005, 149(3), 227―234 |
| 58 | Mastronarde D. N., Held S. R., J. Struct. Biol., 2017, 197(2), 102―113 |
| 59 | Chen M., Bell J. M., Shi X., Sun S. Y., Wang Z., Ludtke S. J., Nat. Methods, 2019, 16(11), 1161―1168 |
| 60 | Gilbert P., J. Theor. Biol., 1972, 36(1), 105―117 |
| 61 | Andersen A. H., Kak A. C., Ultrason. Imaging, 1984, 6(1), 81―94 |
| 62 | Frangakis A. S., Böhm J., Förster F., Nickell S., Nicastro D., Typke D., Hegerl R., Baumeister W., Proc. Nat. Acad. Sci. USA, 2002, 99(22), 14153 |
| 63 | Hrabe T., Chen Y., Pfeffer S., Kuhn Cuellar L., Mangold A. V., Förster F., J. Struct. Biol., 2012, 178(2), 177―188 |
| 64 | Volkmann N., Curr. Opin. Cell Biol., 2012, 24(1), 141―147 |
| 65 | Chen M., Dai W., Sun S. Y., Jonasch D., He C. Y., Schmid M. F., Chiu W., Ludtke S. J., Nat. Methods, 2017, 14(10), 983―985 |
| 66 | Bell J. M., Chen M., Durmaz T., Fluty A. C., Ludtke S. J., J. Struct. Biol., 2018, 204(2), 283―290 |
| 67 | LeCun Y., Bengio Y., Hinton G., Nature, 2015, 521(7553), 436―444 |
| 68 | Martinez⁃Sanchez A., Kochovski Z., Laugks U., Meyer zum Alten Borgloh J., Chakraborty S., Pfeffer S., Baumeister W., Lučić V., Nat. Methods, 2020, 17(2), 209―216 |
| 69 | Castaño⁃Díez D., Kudryashev M., Arheit M., Stahlberg H., J. Struct. Biol., 2012, 178(2), 139―151 |
| 70 | Scheres S. H. W., J. Struct. Biol., 2012, 180(3), 519―530 |
| 71 | Kovacs J. A., Wriggers W., Acta Crystallographica Section D, 2002, 58(8), 1282―1286 |
| 72 | Galaz⁃Montoya J. G., Flanagan J., Schmid M. F., Ludtke S. J., J. Struct. Biol., 2015, 190(3), 279―290 |
| 73 | Bharat T. A. M., Scheres S. H. W., Nat. Protoc., 2016, 11(11), 2054―2065 |
| 74 | Henderson R., Sali A., Baker M. L., Carragher B., Devkota B., Downing K. H., Egelman E. H., Feng Z., Frank J., Grigorieff N., Jiang W., Ludtke S. J., Medalia O., Penczek P. A., Rosenthal P. B., Rossmann M. G., Schmid M. F., Schröder G. F., Steven A. C., Stokes D. L., Westbrook J. D., Wriggers W., Yang H., Young J., Berman H. M., Chiu W., Kleywegt G. J., Lawson C. L., Structure, 2012, 20(2), 205―214 |
| 75 | Himes B. A., Zhang P., Nat. Methods, 2018, 15(11), 955―961 |
| 76 | Pfeffer S., Dudek J., Schaffer M., Ng B. G., Albert S., Plitzko J. M., Baumeister W., Zimmermann R., Freeze H. H., Engel B. D., Förster F., Nature Commun., 2017, 8(1), 14516 |
| 77 | Lin J., Yin W., Smith M. C., Song K., Leigh M. W., Zariwala M. A., Knowles M. R., Ostrowski L. E., Nicastro D., Nature Commun., 2014, 5, 5727―5727 |
| 78 | Brandt F., Etchells S. A., Ortiz J. O., Elcock A. H., Hartl F. U., Baumeister W., Cell, 2009, 136(2), 261―271 |
| 79 | Wan W., Kolesnikova L., Clarke M., Koehler A., Noda T., Becker S., Briggs J. A. G., Nature, 2017, 551(7680), 394―397 |
| 80 | Briggs J. A. G., Riches J. D., Glass B., Bartonova V., Zanetti G., Kräusslich H. G., Proc. Nat. Acad. Sci. USA, 2009, 106(27), 11090 |
| 81 | Calder L. J., Rosenthal P. B., Nat. Struct. Mol. Biol., 2016, 23(9), 853―858 |
| 82 | von Appen A., Kosinski J., Sparks L., Ori A., DiGuilio A. L., Vollmer B., Mackmull M. T., Banterle N., Parca L., Kastritis P., Buczak K., Mosalaganti S., Hagen W., Andres⁃Pons A., Lemke E. A., Bork P., Antonin W., Glavy J. S., Bui K. H., Beck M., Nature, 2015, 526(7571), 140―143 |
| 83 | Fernández⁃Busnadiego R., Zuber B., Maurer U. E., Cyrklaff M., Baumeister W., Lučić V., J. Cell Biol., 2010, 188(1), 145―156 |
| 84 | Tao C. L., Liu Y. T., Sun R., Zhang B., Qi L., Shivakoti S., Tian C. L., Zhang P., Lau P. M., Zhou Z. H., Bi G. Q., the Journal of Neuroscience: the Official Journal of the Society for Neuroscience, 2018, 38(6), 1493―1510 |
| 85 | Medalia O., Beck M., Ecke M., Weber I., Neujahr R., Baumeister W., Gerisch G., Curr. Biol., 2007, 17(1), 79―84 |
| 86 | Urban E., Jacob S., Nemethova M., Resch G. P., Small J. V., Nat. Cell Biol., 2010, 12(5), 429―435 |
| 87 | Bharat T. A. M., Kureisaite⁃Ciziene D., Hardy G. G., Yu E. W., Devant J. M., Hagen W. J. H., Brun Y. V., Briggs J. A. G., Löwe J., Nature Microbiology, 2017, 2(7), 17059 |
| 88 | Beck M., Malmström J. A., Lange V., Schmidt A., Deutsch E. W., Aebersold R., Nat. Methods, 2009, 6(11), 817―823 |
| 89 | Hagen W. J. H., Wan W., Briggs J. A. G., J. Struct. Biol., 2017, 197(2), 191―198 |
| [1] | 汪思聪, 庞贝贝, 刘潇康, 丁韬, 姚涛. XAFS技术在单原子电催化中的应用[J]. 高等学校化学学报, 2022, 43(9): 20220487. |
| [2] | 陈嘉敏, 曲孝章, 齐国华, 徐蔚青, 金永东, 徐抒平. SERS纳米探针对电刺激过程中细胞内产生活性氧的检测[J]. 高等学校化学学报, 2022, 43(6): 20220033. |
| [3] | 李伟辉, 李浩博, 曾诚, 梁昊樾, 陈佳俊, 李俊勇, 李会巧. 热压法构筑锂负极聚偏氟乙烯基双功能保护层的研究[J]. 高等学校化学学报, 2022, 43(2): 20210629. |
| [4] | 王玮, 邹秉辰, 侯捷, 周万里, 罗建平, 王康丽, 蒋凯. Li-Ga液态金属电池界面反应过程的原位检测[J]. 高等学校化学学报, 2022, 43(10): 20220261. |
| [5] | 张太文, 郭军, 张丹, 袁常梅, 邱双艳. trz-Cl-Cu-PMo12的合成、 表征及催化氧化碘离子性能[J]. 高等学校化学学报, 2022, 43(10): 20220215. |
| [6] | 王婕, 霍海燕, 王洋, 张仲, 刘术侠. 铜箔上原位合成NENU-n系列多酸基MOFs的通用策略[J]. 高等学校化学学报, 2022, 43(1): 20210557. |
| [7] | 李奕川, 朱国富, 王宇, 柴永明, 刘晨光, 何盛宝. 基底表面性质与前驱液化学环境对原位定向构筑钛硅分子筛膜的影响[J]. 高等学校化学学报, 2021, 42(9): 2934. |
| [8] | 耿传楠, 化五星, 凌国维, 陶莹, 张辰, 杨全红. 锂硫电池中的催化作用: 材料与表征[J]. 高等学校化学学报, 2021, 42(5): 1331. |
| [9] | 陈炳刚, 刘三荣, 于喜飞, 蒋子江. 聚硅氧烷皮肤组织黏接材料的设计合成与性能表征[J]. 高等学校化学学报, 2021, 42(12): 3746. |
| [10] | 马祥英, 廖艳娟, 覃方红, 尹源浩, 黄在银, 陈其锋. 碳自掺杂g-C3N4光催化性能的原位光微量热-荧光光谱研究[J]. 高等学校化学学报, 2020, 41(11): 2526. |
| [11] | 穆鑫, 姜双双, 张舒皓, 任浩, 孙福兴. SIFSIX-3-Ni膜的制备及氢气分离性质[J]. 高等学校化学学报, 2019, 40(9): 1818. |
| [12] | 王文峰, 秦山, 张荣荣, 周盼盼, 杨庆华, 陈天云. 纳米八面体形FeP@PC的制备及催化析氢性能[J]. 高等学校化学学报, 2019, 40(9): 1979. |
| [13] | 王振宇, 李健华, 郭慧君, 杨春才. 原位合成纳米SiO2增强阴离子聚酯型上浆剂及其对碳纤维复合材料层间剪切强度的影响[J]. 高等学校化学学报, 2019, 40(9): 2005. |
| [14] | 曹培明, 尤静林, 周灿栋, LU Liming, 王建, 王敏, 丁雅妮, 徐琰东. CaF2-Al2O3-MgO电渣微结构的原位拉曼光谱及27Al魔角旋转核磁共振研究[J]. 高等学校化学学报, 2019, 40(6): 1242. |
| [15] | 马东玮, 田润赛, 刘振江, 冯源源, 丁泓宇, 冯季军. Na掺杂Li2-xNaxMnSiO4/C正极材料的微波辅助合成与电化学性能[J]. 高等学校化学学报, 2019, 40(6): 1280. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||