

高等学校化学学报 ›› 2016, Vol. 37 ›› Issue (5): 912.doi: 10.7503/cjcu20150817
收稿日期:2015-10-23
出版日期:2016-05-10
发布日期:2016-04-18
作者简介:联系人简介: 郑积敏, 男, 博士, 教授, 博士生导师, 主要从事磷酸化蛋白质及钙调蛋白的结构与功能研究. E-mail:基金资助:
WANG Nan1, CHENG Xiaoheng2, ZHENG Jimin1,*( ), CHEN Guangju1, JIA Zongchao3
), CHEN Guangju1, JIA Zongchao3
Received:2015-10-23
Online:2016-05-10
Published:2016-04-18
Contact:
ZHENG Jimin
E-mail:jimin_z@bnu.edu.cn
Supported by:摘要:
测试并优化了哺乳动物雷帕霉素靶蛋白(mTOR)的FAT结构域及其亚结构域TRD2和DEPTOR蛋白在大肠杆菌中的表达条件, 获得了TRD2和DEPTOR蛋白成功表达并大量纯化的条件. 结果表明, 麦芽糖结合蛋白(MBP)融合mTOR的TRD2亚结构域可以在菌株BL21(DE3)中大量表达. DEPTOR-G270P突变比野生型DEPTOR在BL21(DE3)中的表达量提高约5倍, 且该突变并不影响DEPTOR自身的二级结构. 通过凝胶过滤实验发现, mTOR的TRD2亚结构域和DEPTOR之间可能存在相互作用.
中图分类号:
TrendMD:
王男, 程小桁, 郑积敏, 陈光巨, 贾宗超. DEPTOR蛋白的表达纯化条件及与mTOR-TRD2亚结构域的相互作用. 高等学校化学学报, 2016, 37(5): 912.
WANG Nan, CHENG Xiaoheng, ZHENG Jimin, CHEN Guangju, JIA Zongchao. Expression and Purification of DEPTOR and Its Interaction with mTOR-TRD2†. Chem. J. Chinese Universities, 2016, 37(5): 912.
| Primer name | Sequence(5' to 3') |
|---|---|
| FAT foward primer | CTGCTGGGTGAGAGAGCT |
| FAT reverse primer | AGACTTAGAAGCCACTGTCAGT |
| TRD2 forward primer | ATCCAGGCTACCTGGTATGA |
| TRD2 reverse primer | CTTGTCCCCGAGCGAC |
Table 1 FAT domain and TRD2 subdomain PCR primer sequence
| Primer name | Sequence(5' to 3') |
|---|---|
| FAT foward primer | CTGCTGGGTGAGAGAGCT |
| FAT reverse primer | AGACTTAGAAGCCACTGTCAGT |
| TRD2 forward primer | ATCCAGGCTACCTGGTATGA |
| TRD2 reverse primer | CTTGTCCCCGAGCGAC |
| Primer name | Sequence(5' to 3') |
|---|---|
| DEPTOR foward primer | GCTTGGTACCGAGCTCGG |
| DEPTOR reverse primer | ATGGTGATGGTGATGATG |
| DEPTOR-G270P-forward | GGCAGCAGCCCCTACTTCAGCAGCAGCCCCACCC |
| DEPTOR-G270P-reverse | TGAAGTAGGGGCTGCTGCCACAGCTGCTCATGCTGCTTCT |
Table 2 DEPTOR PCR Primer and G270P Mutation Primer Sequence
| Primer name | Sequence(5' to 3') |
|---|---|
| DEPTOR foward primer | GCTTGGTACCGAGCTCGG |
| DEPTOR reverse primer | ATGGTGATGGTGATGATG |
| DEPTOR-G270P-forward | GGCAGCAGCCCCTACTTCAGCAGCAGCCCCACCC |
| DEPTOR-G270P-reverse | TGAAGTAGGGGCTGCTGCCACAGCTGCTCATGCTGCTTCT |
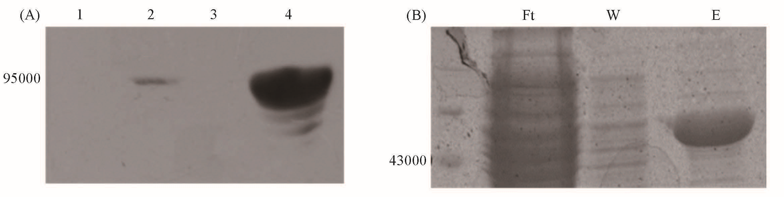

Fig.2 FAT domain and subdomain expression assay in E.coli(A) FAT-GST fusion protein in JM109 expression assay by Western-Blot, lane 1: negative control (incubate without adding IPTG), lane 2: whole-cell-extract after induced by 1 mmol/L IPTG, lanes 3, 4: supernatant/pellet of whole-cell-extract lysate. (B) MBP fusion TRD2 protein in BL21(DE3) expression assay by SDS-PAGE(Ft: flow-through; W: wash; E: eluted protein).
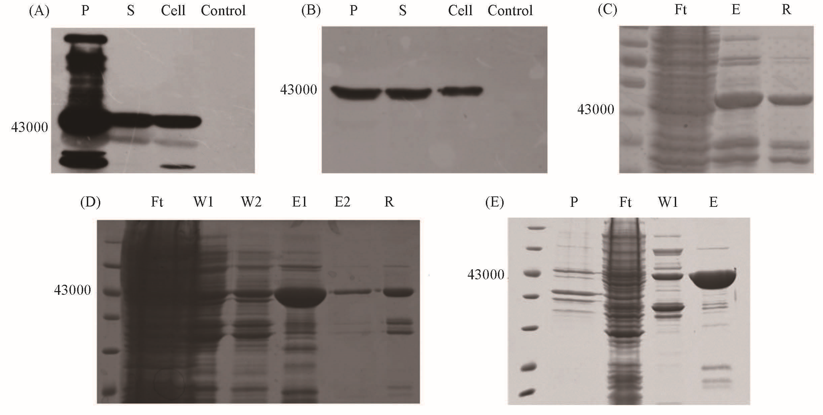
Fig.3 DEPTOR expression assay in E.coliAll the lanes are samples from each purification step, respectively. (A) DEPTOR-His wild-type expression assay by Western-Blot in BL21(DE3); (B) DEPTOR-His wild-type expression assay by Western-Blot in arctic express; (C) DEPTOR-His wild-type expression assay by SDS-PAGE in BL21(DE3); (D) DEPTOR-His G270P expression assay by SDS-PAGE; (E) DEPTOR-His G270P expression assay by SDS-PAGE with glycerol(10%, volume fraction) in purification buffer. P: pellet; S: supernatant; Cell: whole-cell-extract; Control: cell without induced by IPTG; Ft: flow-through; W1, W2: wash; E, E1 and E2: eluted protein; R: resin after elution.
| Secondary structure | Percentage of secondary structure(%) | |||||
|---|---|---|---|---|---|---|
| 180—260 nm | 185—260 nm | 190—260 nm | 195—260 nm | 200—260 nm | 205—260 nm | |
| α-Helix | 29.1 | 28.8 | 28.8 | 27.8 | 26.7 | 27.6 |
| Antiparallel β-strand | 14.2 | 14.6 | 15.0 | 13.4 | 11.0 | 9.4 |
| Parallel β-strand | 9.1 | 9.0 | 8.9 | 9.8 | 10.5 | 10.9 |
| β-Turns | 18.3 | 18.3 | 18.2 | 18.1 | 18.4 | 17.8 |
| Coiled-coil | 31.8 | 32.2 | 31.8 | 33.6 | 35.9 | 37.7 |
| Total | 102.4 | 102.9 | 102.8 | 102.7 | 102.5 | 103.3 |
Table 3 Percentage of secondary structure in DEPTOR-wt*
| Secondary structure | Percentage of secondary structure(%) | |||||
|---|---|---|---|---|---|---|
| 180—260 nm | 185—260 nm | 190—260 nm | 195—260 nm | 200—260 nm | 205—260 nm | |
| α-Helix | 29.1 | 28.8 | 28.8 | 27.8 | 26.7 | 27.6 |
| Antiparallel β-strand | 14.2 | 14.6 | 15.0 | 13.4 | 11.0 | 9.4 |
| Parallel β-strand | 9.1 | 9.0 | 8.9 | 9.8 | 10.5 | 10.9 |
| β-Turns | 18.3 | 18.3 | 18.2 | 18.1 | 18.4 | 17.8 |
| Coiled-coil | 31.8 | 32.2 | 31.8 | 33.6 | 35.9 | 37.7 |
| Total | 102.4 | 102.9 | 102.8 | 102.7 | 102.5 | 103.3 |
| Secondary structure | Percentage of secondary structure(%) | |||||
|---|---|---|---|---|---|---|
| 180—260 nm | 185—260 nm | 190—260 nm | 195—260 nm | 200—260 nm | 205—260 nm | |
| α-Helix | 28.8 | 29.4 | 29.3 | 28.6 | 28.0 | 29.4 |
| Antiparallel β-strand | 14.9 | 14.2 | 14.7 | 12.9 | 10.5 | 8.8 |
| Parallel β-strand | 9.1 | 8.8 | 8.7 | 9.5 | 10.0 | 10.3 |
| β-Turns | 18.4 | 18.2 | 18.1 | 17.9 | 18.1 | 17.4 |
| Coiled-coil | 31.0 | 31.2 | 31.0 | 32.6 | 34.7 | 36.2 |
| Total | 102.2 | 101.8 | 101.9 | 101.6 | 101.3 | 102.1 |
Table 4 Percentage of secondary structure in DEPTOR-G270P*
| Secondary structure | Percentage of secondary structure(%) | |||||
|---|---|---|---|---|---|---|
| 180—260 nm | 185—260 nm | 190—260 nm | 195—260 nm | 200—260 nm | 205—260 nm | |
| α-Helix | 28.8 | 29.4 | 29.3 | 28.6 | 28.0 | 29.4 |
| Antiparallel β-strand | 14.9 | 14.2 | 14.7 | 12.9 | 10.5 | 8.8 |
| Parallel β-strand | 9.1 | 8.8 | 8.7 | 9.5 | 10.0 | 10.3 |
| β-Turns | 18.4 | 18.2 | 18.1 | 17.9 | 18.1 | 17.4 |
| Coiled-coil | 31.0 | 31.2 | 31.0 | 32.6 | 34.7 | 36.2 |
| Total | 102.2 | 101.8 | 101.9 | 101.6 | 101.3 | 102.1 |
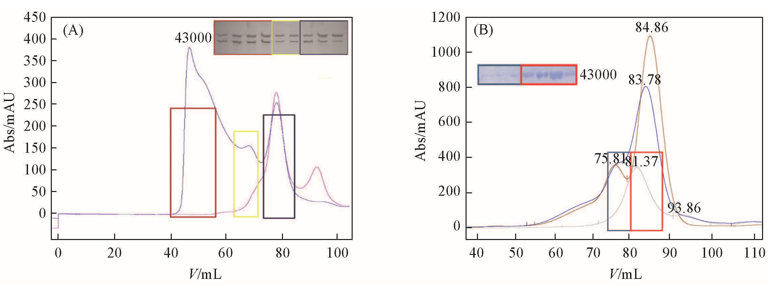
Fig.5 Interaction assay of MBP-TRD2 and DEPTOR(A) MBP-TRD2 and DEPTOR co-purification by gel-filtration (Hi-Load S200) in 20 mmol/L Tris, pH=8.0 and 150 mmol/L NaCl; (B) MBP and DEPTOR co-purification by gel-filtration in the same condition as a negative control. Samples of the putative complexes are picked and analyzed by SDS-PAGE. In each figure, x-axis indicates the elution volume which reflects molecular mass of the protein and y-axis indicates the absorption value which is in direct proportion to protein concentration.
| [1] | Wang J. B., Jiang Y., Liang H., Li P., Xiao H. J., Ji J., Xiang W., Shi J. F., Fan Y. G., Li L., Wang D., Deng S. S., Chen W. Q., Wei W. Q., Qiao Y. L., Boffetta P., Ann.Oncol., 2001, 23(11), 2983—2989 |
| [2] | Qiao Y. L., Chin. J.Oncol., 2009, 34(7), 483—485 |
| [3] | Wei K. R., Liang Z. H., Liu J., Wang Y. N., Chinese Journal of Medical History, 2012, 42(1), 21—25 |
| (魏矿荣, 梁智恒, 刘静, 王亚娜. 中华医史杂志, 2012, 42(1), 21—25) | |
| [4] | Fan W. H., Zhang L. H., Zhang Z. L., Guo J. F., Ren A. M., Ge P. F., Chem. J. Chinese Universities,2013, 34(7), 1731—1738 |
| (范文海, 张丽红, 张子龙, 郭景富, 任爱民, 葛鹏飞. 高等学校化学学报, 2013, 34(7), 1731—1738) | |
| [5] | Peterson T. R., Laplante M., Thoreen C. C., Sancak Y., Kang S. A., Kuehl W. M., Gray N. S., Sabatini D. M., Cell, 2009, 137(5), 873—886 |
| [6] | Fingar D. C., Salama S., Tsou C., Harlow E., Belnis J., GenesDev., 2002, 16(12), 1472—1487 |
| [7] | Gera J., Lichtenstein A., LeukLymphoma, 2011, 52(10), 1857—1866 |
| [8] | Lu M., Wang J., Elves H., Pearce D., J. Biol.Chem., 2011, 286(35), 30647—30654 |
| [9] | Chen J., Zheng X. F., Brown E. J., Scheriber S. L., Proc. Natl. Acad.Sci., 1995, 92(11), 4947—4951 |
| [10] | Yang H., Rudge D. G., Koos J. D., Vaidialingam B., Yang H. J., Pavletich N. P., Nature,2013, 497(7448), 217—223 |
| [11] | Dames S. A., J. Biol.Chem., 2010, 285(10), 7766—7775 |
| [12] | Hardt M., Chantaravisoot N., Tamanoi F., Genes toCells, 2011, 16(2), 141—151 |
| [13] | Sturgill T. W., Hall M. N., ACS Chem.Biol., 2009, 4(12), 999—1015 |
| [14] | Bakkenist C. J., Kastan M. B., Nature,2003, 421(6922), 499—506 |
| [15] | Mehtab P., Mohammed M. A., Akhtar A., Mahboob A., Chem. Res. ChineseUniversities, 2014, 30(1), 55—62 |
| [16] | Imamura S., Kishi S., Int. J. Biochem. Cell. Biol., 2005, 37(5), 1105—1116 |
| [17] | Tanaka A., Weinel S., Nagy N., O’Driscoll M., Lai-Cheong J. E., Kulp-Shorten C. L., Knable A., Carpenter G., Fisher S. A., Hiragun M., Yanase Y., Hide M., Callen J., McGrath J. A., Am. J. Hum.Genet., 2012, 90(3), 511—517 |
| [18] | Jiang X., Sun Y., Chen S., Roy K., Price B. D., J. Biol.Chem., 2006, 281(23), 15741—15746 |
| [19] | Rivera-Calzada A., Maman J. D., Spagnolo L., Pearl L. H., Llorca O., Structure, 2005, 13(2), 243—255 |
| [20] | de la Rubia J., Such E., LeukLymphoma, 2010, 51(11), 1960—1961 |
| [21] | Tsai W. B., Chung Y. M., Takahashi Y., Xu Z., Hu M. C., Nat. Cell.Biol., 2008, 10(4), 460—467 |
| [22] | Kazi A. A., Hong-Brown L., Lang S. M., Lang C. H., Mol.Med., 2011, 17(9), 925—936 |
| [23] | Zhang X. Y., Deng D. J., Tan J. J., He Y., Li C. H., Wang C. X., Chem. Res. ChineseUniversities, 2014, 30(2), 297—305 |
| [1] | 唐玉静 胡敏 王霞 王启刚. 载酶纳米催化体系用于疾病诊疗研究进展[J]. 高等学校化学学报, 0, (): 20220640. |
| [2] | 常丽颖 凌鑫宇 陈和祺 王雪 刘涛. 基因编辑在线粒体疾病中的应用[J]. 高等学校化学学报, 2022, 43(Album-4): 20220363. |
| [3] | 曹舒杰, 李泓君, 管文丽, 任梦田, 周传政. 硫代磷酸酯寡聚核苷酸的立体控制合成研究进展[J]. 高等学校化学学报, 2022, 43(Album-4): 20220304. |
| [4] | 徐永斌 冯帅霞 陈洁 龚欢 施松善 王辉俊 王顺春. 红花均一多糖的结构表征及其抑制HepG2增殖活性[J]. 高等学校化学学报, 0, (): 20220600. |
| [5] | 仵宇帅, 尚颖旭, 蒋乔, 丁宝全. 可控自组装DNA折纸结构作为药物载体的研究进展[J]. 高等学校化学学报, 2022, 43(8): 20220179. |
| [6] | 刘文婷 刘柳宜 朱博琛 毛宗万. 核酸G-四链体的识别、复合物结构与细胞内探测的研究进展[J]. 高等学校化学学报, 0, (): 20220419. |
| [7] | 胡玉灿 曹朝辉 郑灵刚 沈俊涛 赵维 戴磊. CRISPR-Cas基因编辑技术在微生物组工程的应用[J]. 高等学校化学学报, 0, (): 20220362. |
| [8] | 方鑫 赵瑞奇 莫婧 王雅芬 翁小成. 检测核酸表观遗传修饰的测序方法[J]. 高等学校化学学报, 0, (): 20220342. |
| [9] | 张凯嵩 王少儒 张雨桐 田沺. 基于超分子化学的核酸表观遗传修饰研究[J]. 高等学校化学学报, 0, (): 20220335. |
| [10] | 朱凯, 利婕, 武潇逸, 胡薇薇, 吴冬梅, 虞诚潇, 葛志伟, 叶兴乾, 陈士国. 基于多孔石墨化碳柱-四极杆-飞行时间质谱解析甜菜果胶精细结构[J]. 高等学校化学学报, 2022, 43(6): 20220023. |
| [11] | 付俊, 吴美婵, 王书珍, 邵秀丽, 何峰. 基于标记法的福白菊精油抗真菌机理研究[J]. 高等学校化学学报, 2021, 42(12): 3657. |
| [12] | 赵卓, 王雪强. 核酸适体偶联药物的生物偶联构建技术与应用[J]. 高等学校化学学报, 2021, 42(11): 3367. |
| [13] | 陈旺, 胡代花, 刘格歌. 由醋酸去氢表雄酮合成熊去氧胆酸[J]. 高等学校化学学报, 2021, 42(9): 2782. |
| [14] | 胡皓程, 李文利, 张嘉宁, 刘宇博. 黑木耳寡糖的提取、 结构表征及生物活性[J]. 高等学校化学学报, 2021, 42(8): 2465. |
| [15] | 杨依然, 姚华, 闫江红, 孙志恒, 张余, 房雪晴, 李绪文, 金永日. 薤中新的甾体皂苷类化学成分[J]. 高等学校化学学报, 2021, 42(6): 1742. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||