

Chem. J. Chinese Universities ›› 2014, Vol. 35 ›› Issue (12): 2605.doi: 10.7503/cjcu20140713
• Physical Chemistry • Previous Articles Next Articles
WU Yunjian, CUI Yinglu, ZHENG Qingchuan*( ), ZHANG Hongxing
), ZHANG Hongxing
Received:2014-08-01
Online:2014-12-10
Published:2014-11-10
Contact:
ZHENG Qingchuan
E-mail:zhengqc@jlu.edu.cn
Supported by:CLC Number:
TrendMD:
WU Yunjian, CUI Yinglu, ZHENG Qingchuan, ZHANG Hongxing. Theoretical Studies on the Substrate Binding Mode and Regioselectivity of Human CYP2C9 with S- and R-Warfarin†[J]. Chem. J. Chinese Universities, 2014, 35(12): 2605.
| System | ΔEele/ (kJ·mol-1) | ΔEvdw/ (kJ·mol-1) | ΔGGB/ (kJ·mol-1) | ΔGSA/ (kJ·mol-1) | ΔGMM-GB/SA/ (kJ·mol-1) | -TΔS/ (kJ·mol-1) | ΔGbind/ (kJ·mol-1) | KD(expt.) | ΔGbind-expt./ (kJ·mol-1) |
|---|---|---|---|---|---|---|---|---|---|
| cs-2C9-S-Warfarin | -112.14 | -157.18 | 191.67 | -12.39 | -90.04 | 83.09 | -6.95 | ||
| rd-2C9-S-Warfarin | -100.25 | -171.79 | 170.53 | -11.93 | -113.44 | 82.63 | -30.81 | 6 | -31.02 |
Table 1 Binding free energies for the ligands in the complexes*
| System | ΔEele/ (kJ·mol-1) | ΔEvdw/ (kJ·mol-1) | ΔGGB/ (kJ·mol-1) | ΔGSA/ (kJ·mol-1) | ΔGMM-GB/SA/ (kJ·mol-1) | -TΔS/ (kJ·mol-1) | ΔGbind/ (kJ·mol-1) | KD(expt.) | ΔGbind-expt./ (kJ·mol-1) |
|---|---|---|---|---|---|---|---|---|---|
| cs-2C9-S-Warfarin | -112.14 | -157.18 | 191.67 | -12.39 | -90.04 | 83.09 | -6.95 | ||
| rd-2C9-S-Warfarin | -100.25 | -171.79 | 170.53 | -11.93 | -113.44 | 82.63 | -30.81 | 6 | -31.02 |
| Residue | ΔEvdw/(kJ·mol-1) | ΔEele/(kJ·mol-1) | ΔGGB/(kJ·mol-1) | ΔGSASA/(kJ·mol-1) | Δ |
|---|---|---|---|---|---|
| Leu366 | -7.70 | -2.26 | 0.84 | -1.09 | -10.21 |
| Ala103 | -7.12 | -6.70 | 6.03 | -1.09 | -8.87 |
| Pro367 | -7.79 | -3.18 | 3.01 | -0.67 | -8.62 |
| Phe100 | -7.41 | -1.30 | 1.21 | -0.50 | -7.99 |
| Ile99 | -6.07 | -5.32 | 3.77 | -0.29 | -7.91 |
| Leu102 | -3.81 | -5.02 | 4.86 | -0.21 | -4.19 |
| Phe476 | -3.39 | 0.25 | 0.63 | -0.84 | -3.35 |
| Leu208 | -3.39 | 0.08 | 1.59 | -0.88 | -2.60 |
| Phe114 | -2.55 | 0.46 | -0.04 | -0.25 | -2.39 |
Table 2 Decomposition of binding free energy on per-residue basis for cs-2C9-S-Warfarin system
| Residue | ΔEvdw/(kJ·mol-1) | ΔEele/(kJ·mol-1) | ΔGGB/(kJ·mol-1) | ΔGSASA/(kJ·mol-1) | Δ |
|---|---|---|---|---|---|
| Leu366 | -7.70 | -2.26 | 0.84 | -1.09 | -10.21 |
| Ala103 | -7.12 | -6.70 | 6.03 | -1.09 | -8.87 |
| Pro367 | -7.79 | -3.18 | 3.01 | -0.67 | -8.62 |
| Phe100 | -7.41 | -1.30 | 1.21 | -0.50 | -7.99 |
| Ile99 | -6.07 | -5.32 | 3.77 | -0.29 | -7.91 |
| Leu102 | -3.81 | -5.02 | 4.86 | -0.21 | -4.19 |
| Phe476 | -3.39 | 0.25 | 0.63 | -0.84 | -3.35 |
| Leu208 | -3.39 | 0.08 | 1.59 | -0.88 | -2.60 |
| Phe114 | -2.55 | 0.46 | -0.04 | -0.25 | -2.39 |
| Residue | ΔEvdw/(kJ·mol-1) | ΔEele/(kJ·mol-1) | ΔGGB/(kJ·mol-1) | ΔGSASA/(kJ·mol-1) | ΔGbind/(kJ·mol-1) |
|---|---|---|---|---|---|
| Glu300 | -0.50 | -47.63 | 38.43 | -0.96 | -10.67 |
| Leu208 | -4.56 | 0.46 | -5.19 | -1.17 | -10.46 |
| Asn204 | -0.96 | -2.76 | -4.10 | -0.21 | -8.04 |
| Ala297 | -6.20 | -2.64 | 2.68 | -0.46 | -6.61 |
| Leu366 | -4.60 | -0.75 | 0.25 | -0.71 | -5.82 |
| Val113 | -3.93 | -2.97 | 1.67 | -0.33 | -5.57 |
| Phe114 | -5.02 | -2.26 | 3.47 | -1.09 | -4.90 |
| Phe476 | -0.38 | -0.25 | -2.64 | -0.04 | -3.31 |
| Thr301 | -1.88 | 1.21 | -2.60 | 0.04 | -3.22 |
| Leu362 | -2.34 | 0.21 | -0.50 | -0.46 | -3.10 |
Table 3 Decomposition of binding free energy on per-residue basis for rd-2C9-S-Warfarin system
| Residue | ΔEvdw/(kJ·mol-1) | ΔEele/(kJ·mol-1) | ΔGGB/(kJ·mol-1) | ΔGSASA/(kJ·mol-1) | ΔGbind/(kJ·mol-1) |
|---|---|---|---|---|---|
| Glu300 | -0.50 | -47.63 | 38.43 | -0.96 | -10.67 |
| Leu208 | -4.56 | 0.46 | -5.19 | -1.17 | -10.46 |
| Asn204 | -0.96 | -2.76 | -4.10 | -0.21 | -8.04 |
| Ala297 | -6.20 | -2.64 | 2.68 | -0.46 | -6.61 |
| Leu366 | -4.60 | -0.75 | 0.25 | -0.71 | -5.82 |
| Val113 | -3.93 | -2.97 | 1.67 | -0.33 | -5.57 |
| Phe114 | -5.02 | -2.26 | 3.47 | -1.09 | -4.90 |
| Phe476 | -0.38 | -0.25 | -2.64 | -0.04 | -3.31 |
| Thr301 | -1.88 | 1.21 | -2.60 | 0.04 | -3.22 |
| Leu362 | -2.34 | 0.21 | -0.50 | -0.46 | -3.10 |
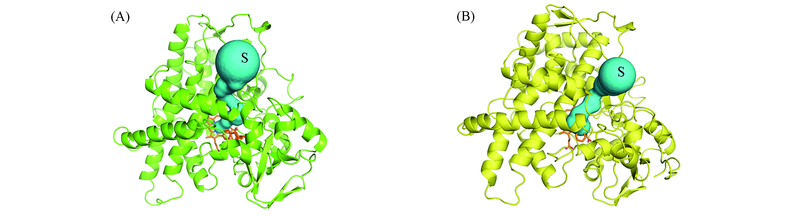
Fig.5 Access channels identified from the representative structures of CYP2C9-substrate complexes for cs-2C9-S-Warfarin system(A) and rd-2C9-S-Warfarin system(B)
| ΔEele/ (kJ·mol-1) | ΔEvdw/ (kJ·mol-1) | ΔGGB/ (kJ·mol-1) | ΔGSA/ (kJ·mol-1) | ΔGMM-GB/SA/ (kJ·mol-1) | -TΔS/ (kJ·mol-1) | ΔGbind/ (kJ·mol-1) |
|---|---|---|---|---|---|---|
| -85.27 | -133.74 | 145.67 | -9.63 | -82.96 | 87.94 | 4.98 |
Table 4 Binding free energies for 2C9-R-Warfarin complex
| ΔEele/ (kJ·mol-1) | ΔEvdw/ (kJ·mol-1) | ΔGGB/ (kJ·mol-1) | ΔGSA/ (kJ·mol-1) | ΔGMM-GB/SA/ (kJ·mol-1) | -TΔS/ (kJ·mol-1) | ΔGbind/ (kJ·mol-1) |
|---|---|---|---|---|---|---|
| -85.27 | -133.74 | 145.67 | -9.63 | -82.96 | 87.94 | 4.98 |
| Residue | ΔEvdw/(kJ·mol-1) | ΔEele/(kJ·mol-1) | ΔGGB/(kJ·mol-1) | ΔGSASA/(kJ·mol-1) | ΔGbind/(kJ·mol-1) |
|---|---|---|---|---|---|
| Phe476 | -5.48 | -0.75 | -0.17 | -0.54 | -6.95 |
| Leu208 | -6.95 | -0.13 | 1.55 | -1.34 | -6.86 |
| Leu366 | -6.53 | -1.05 | 1.88 | -0.92 | -6.61 |
| Phe114 | -3.73 | -1.26 | 1.88 | -0.88 | -3.98 |
| Val113 | -3.18 | -0.88 | 0.63 | -0.50 | -3.93 |
| Phe100 | -2.72 | -0.21 | -0.38 | -0.38 | -3.68 |
| Leu362 | -2.97 | 0.33 | 0.08 | -0.50 | -3.06 |
| Ala103 | -2.43 | -1.72 | 2.09 | -0.63 | -2.68 |
| Ala297 | -2.18 | -0.08 | 0.33 | -0.42 | -2.34 |
| Ile213 | -2.05 | 0.59 | -0.33 | -0.38 | -2.18 |
Table 5 Decomposition of binding free energy on per-residue basis for 2C9-R-Warfarin system
| Residue | ΔEvdw/(kJ·mol-1) | ΔEele/(kJ·mol-1) | ΔGGB/(kJ·mol-1) | ΔGSASA/(kJ·mol-1) | ΔGbind/(kJ·mol-1) |
|---|---|---|---|---|---|
| Phe476 | -5.48 | -0.75 | -0.17 | -0.54 | -6.95 |
| Leu208 | -6.95 | -0.13 | 1.55 | -1.34 | -6.86 |
| Leu366 | -6.53 | -1.05 | 1.88 | -0.92 | -6.61 |
| Phe114 | -3.73 | -1.26 | 1.88 | -0.88 | -3.98 |
| Val113 | -3.18 | -0.88 | 0.63 | -0.50 | -3.93 |
| Phe100 | -2.72 | -0.21 | -0.38 | -0.38 | -3.68 |
| Leu362 | -2.97 | 0.33 | 0.08 | -0.50 | -3.06 |
| Ala103 | -2.43 | -1.72 | 2.09 | -0.63 | -2.68 |
| Ala297 | -2.18 | -0.08 | 0.33 | -0.42 | -2.34 |
| Ile213 | -2.05 | 0.59 | -0.33 | -0.38 | -2.18 |
| [1] | de Graaf C., Vermeulen N. P. E., Feenstra K. A., J. Med. Chem., 2005, 48, 2725—2755 |
| [2] | Denisov I. G., Makris T. M., Sligar S. G., Schlichting I., Chem. Rev., 2005, 105, 2253—2278 |
| [3] | Hasemann C. A., Kurumbail R. G., Boddupalli S. S., Peterson J. A., Deisenhofer J., Structure,1995, 3(1), 41—62 |
| [4] | Presnell S. R., Cohen F. E., Proc. Natl. Acad. Sci., 1989, 86(17), 6592—6596 |
| [5] | Gotoh O., J. Biol. Chem., 1992, 267(1), 83—90 |
| [6] | Otyepka M., Berka K., Anzenbacher P., Curr. Drug Metab., 2012, 13, 130—142 |
| [7] | Cui Y. L., Zhang J. L., Zheng Q. C., Niu R. J., Xu Y., Zhang H. X., Sun C. C., Chem. Euro. J., 2013, 19, 549—557 |
| [8] | Cui Y. L., Zheng Q. C., Zhang J. L., Xue Q., Wang Y., Zhang H. X., J. Chem. Inf. Model., 2013, 53, 3308—3317 |
| [9] | Meng X. Y., Zheng Q. C., Zhang H. X., BBA-Proteins Proteom., 2009, 1794, 1066—1072 |
| [10] | Rendic S., Carlo F. J. D., Drug Metab. Rev., 1997, 29, 413—580 |
| [11] | Aithal G. P., Day C. P., Kesteven P. J., Daly A. K., The Lancet, 1999, 353, 717—719 |
| [12] | Rettie A. E., Korzekwa K. R., Kunze K. L., Lawrence R. F., Eddy A. C., Aoyama T., Gelboin H. V., Gonzalez F. J., Trager W. F., Chem. Res. Toxicol., 1992, 5, 54—59 |
| [13] | Williams P. A., Cosme J., Ward A., Angove H. C., Vinković D. M., Jhoti H., Nature, 2003, 424, 464—468 |
| [14] | Wester M. R., Yano J. K., Schoch G. A., Yang C., Griffin K. J., Stout C. D., Johnson E. F., J. Bio. Chem., 2004, 279, 35630—35637 |
| [15] | Jones G., Willett P., Glen R. C., Leach A. R., Discovery Studio 2.5, Accelrys Inc., San Diego, 2009 |
| [16] | Wu G., Robertson D. H., Brooks C. L., Vieth M., J. Comput. Chem., 2003, 24, 1549—1562 |
| [17] | Xu Y., Cui Y. L., Zheng Q. C., Zhang H. X., Sun C. C., Chem. J. Chinese Universities, 2013, 34(5), 1226—1232 |
| (徐钰, 崔颖璐, 郑清川, 张红星, 孙家锺. 高等学校化学学报, 2013, 34(5), 1226—1232) | |
| [18] | Case D., Darden T., Cheatham III T., Simmerling C., Wang J., Duke R., Luo R., Walker R., Zhang W., Merz K., AMBER 11, University of California, San Francisco, 2010 |
| [19] | Lindorff-Larsen K., Piana S., Palmo K., Maragakis P., Klepeis J. L., Dror R.O., Shaw D.E., Proteins, 2010, 78, 1950—1958 |
| [20] | Frisch M. J., Trucks G. W., Schlegel H. B., Scuseria G. E., Robb M. A., Cheeseman J. R., Scalmani G., Barone V., Mennucci B., Petersson G. A., Nakatsuji H., Caricato M., Li X., Hratchian H. P., Izmaylov A. F., Bloino J., Zheng G., Sonnenberg J. L., Hada M., Ehara M., Toyota K., Fukuda R., Hasegawa J., Ishida M., Nakajima T., Honda Y., Kitao O., Nakai H., Vreven T., Montgomery J. A. Jr., Peralta J. E., Ogliaro F., Bearpark M., Heyd J. J., Brothers E., Kudin K. N., Staroverov V. N., Keith T., Kobayashi R., Normand J., Raghavachari K., Rendell A., Burant J. C., Iyengar S. S., Tomasi J., Cossi M., Rega N., Millam J. M., Klene M., Knox J. E., Cross J. B., Bakken V., Adamo C., Jaramillo J., Gomperts R., Stratmann R. E., Yazyev O., Austin A. J., Cammi R., Pomelli C., Ochterski J. W., Martin R. L., Morokuma K., Zakrzewski V. G., Voth G. A., Salvador P., Dannenberg J. J., Dapprich S. Daniels A. D., Farkas O., Foresman J. B., Ortiz J. V., Cioslowski J., Fox D. J., Gaussian 09, Gaussian Inc., Wallingford CT, 2010 |
| [21] | Wang J., Wang W., Kollman P.A., Case D.A., J. Am. Chem. Soc., 2001, 222, U403 |
| [22] | Xue Q., Zhang J. L., Zheng Q. C., Cui Y. L., Chen L., Chu W. T., Zhang H. X., Langmuir,2013, 29, 11135—11144 |
| [23] | Xia D. H, Ren X. D, Jiao L., Li H., Chem. Res. Chinese Universities, 2012, 28(2), 282—286 |
| [24] | Chovancova E., Pavelka A., Benes P., Strnad O., Brezovsky J., Kozlikova B., Gora A., Sustr V., Klvana M., Medek P., PLOS Comput. Bio., 2012, 8, e1002708 |
| [25] | DeLano W. L., The PyMOL Molecular Graphics System, Schrödinger, LLC, Cambridge, 2002 |
| [1] | GAO Zhiwei, LI Junwei, SHI Sai, FU Qiang, JIA Junru, AN Hailong. Analysis of Gating Characteristics of TRPM8 Channel Based on Molecular Dynamics [J]. Chem. J. Chinese Universities, 2022, 43(6): 20220080. |
| [2] | HU Bo, ZHU Haochen. Dielectric Constant of Confined Water in a Bilayer Graphene Oxide Nanosystem [J]. Chem. J. Chinese Universities, 2022, 43(2): 20210614. |
| [3] | ZHANG Mi, TIAN Yafeng, GAO Keli, HOU Hua, WANG Baoshan. Molecular Dynamics Simulation of the Physicochemical Properties of Trifluoromethanesulfonyl Fluoride Dielectrics [J]. Chem. J. Chinese Universities, 2022, 43(11): 20220424. |
| [4] | LEI Xiaotong, JIN Yiqing, MENG Xuanyu. Prediction of the Binding Site of PIP2 in the TREK-1 Channel Based on Molecular Modeling [J]. Chem. J. Chinese Universities, 2021, 42(8): 2550. |
| [5] | LI Congcong, LIU Minghao, HAN Jiarui, ZHU Jingxuan, HAN Weiwei, LI Wannan. Theoretical Study of the Catalytic Activity of VmoLac Non-specific Substrates Based on Molecular Dynamics Simulations [J]. Chem. J. Chinese Universities, 2021, 42(8): 2518. |
| [6] | ZENG Yonghui, YAN Tianying. Vibrational Density of States Analysis of Proton Hydration Structure [J]. Chem. J. Chinese Universities, 2021, 42(6): 1855. |
| [7] | QI Renrui, LI Minghao, CHANG Hao, FU Xueqi, GAO Bo, HAN Weiwei, HAN Lu, LI Wannan. Theoretical Study on the Unbinding Pathway of Xanthine Oxidase Inhibitors Based on Steered Molecular Dynamics Simulation [J]. Chem. J. Chinese Universities, 2021, 42(3): 758. |
| [8] | LIU Aiqing, XU Wensheng, XU Xiaolei, CHEN Jizhong, AN Lijia. Molecular Dynamics Simulation of Polymer/rod Nanocomposite [J]. Chem. J. Chinese Universities, 2021, 42(3): 875. |
| [9] | SHUAI Die, ZHAO Meijuan, CHEN Bingnian, WANG Li. Inhibitory Effect of Four Kinds of Keegin-type Phosphomolybdate on Tyrosinase and Melanin Formation and Its Antioxidant Activities [J]. Chem. J. Chinese Universities, 2021, 42(12): 3579. |
| [10] | YANG Ju, SU Lijiao, LI Canhua, LU Jiajia, YANG Junli, GU Jie, YANG Li, YANG Lijuan. Host-guest Complexation Behavior of Nardosinone and Water-soluble Phosphate Salt Pillar[6]arene [J]. Chem. J. Chinese Universities, 2021, 42(10): 3099. |
| [11] | ZHANG Aiqin, WANG Man, SHEN Gangyi, JIN Jun. Interactions Between Polybrominated Diphenyl Ethers and Human Serum Albumin Using SPR and Molecular Docking [J]. Chem. J. Chinese Universities, 2020, 41(9): 2054. |
| [12] | WANG Lianping,LI Qingjie,LIU Xiaoyan,REN Yueying,YANG Xiuwei. Screening of Cholinesterase Inhibitors in Fructus Evodiae Alkaloids Based on UFLC-MS/molecular Simulation † [J]. Chem. J. Chinese Universities, 2020, 41(1): 111. |
| [13] | WANG Xiaoxia, MA Litong, NIE Zhihua, WANG Zhengde, CUI Jinlong, ZHAO Wenyuan, SAI Huazheng. Interaction Between Fulvic Acid and Pepsin Investigated by Multispectral and Molecular Docking Simulation † [J]. Chem. J. Chinese Universities, 2019, 40(9): 1840. |
| [14] | QU Siying, XU Qin. Different Roles of Some Key Residues in the S4 Pocket of Coagulation Factor Xa for Rivaroxaban Binding † [J]. Chem. J. Chinese Universities, 2019, 40(9): 1918. |
| [15] | MA Yucong, FAN Baomin, WANG Manman, YANG Biao, HAO Hua, SUN Hui, ZHANG Huijuan. Two-step Preparation of Trazodone and Its Corrosion Inhibition Mechanism for Carbon Steel [J]. Chem. J. Chinese Universities, 2019, 40(8): 1706. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||