

Chem. J. Chinese Universities ›› 2019, Vol. 40 ›› Issue (9): 1918.doi: 10.7503/cjcu20190261
• Physical Chemistry • Previous Articles Next Articles
Received:2019-05-08
Online:2019-09-10
Published:2019-07-16
Contact:
XU Qin
E-mail:xuqin523@sjtu.edu.cn
Supported by:CLC Number:
TrendMD:
QU Siying, XU Qin. Different Roles of Some Key Residues in the S4 Pocket of Coagulation Factor Xa for Rivaroxaban Binding †[J]. Chem. J. Chinese Universities, 2019, 40(9): 1918.
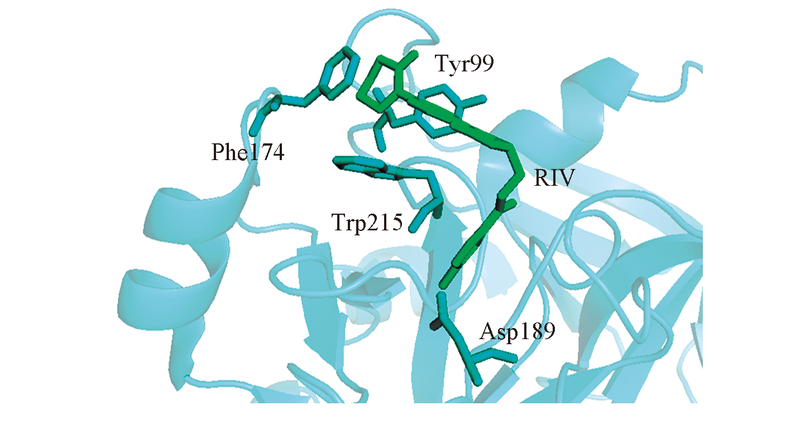
Fig.1 Model of RIV binding to FXa Based on PDB structure 2W26, RIV is shown by sticks in green, the key residues Tyr99, Phe174, Trp215 and Asp189 are shown by sticks in cyan, while the backbone of FXa is shown by cartoon.
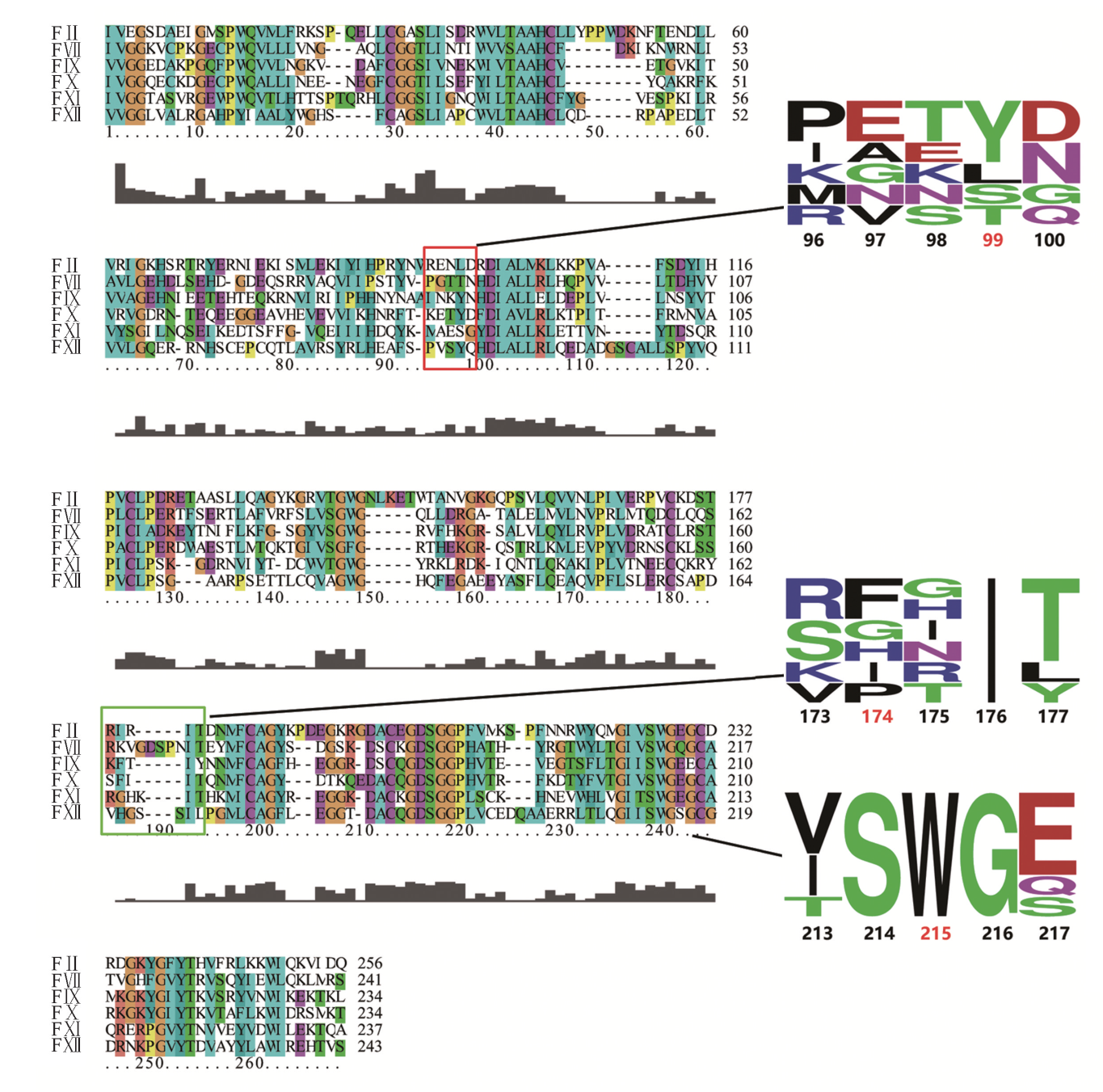
Fig.2 Sequence alignment of the serine protease domain and conservation analysis of residues around the S4 pockets among homologous coagulation factors Surrounding the S4 pocket, the critical residues of 99 loop, 170 loop and 215 strand are framed in red, green and blue, respectively. The corresponding conservation analyses are shown on the right with the residues labeled by Chymotrypsin numbering.
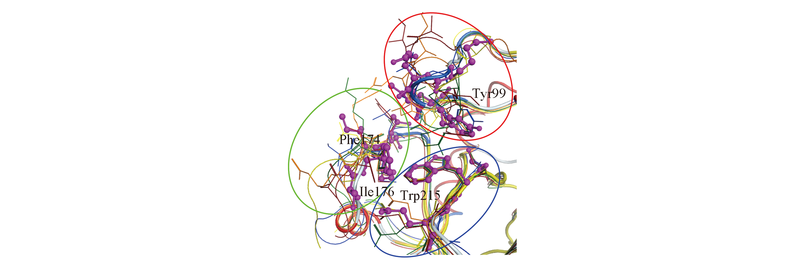
Fig.3 Superposition of the S4 pockets of homologous coagulation factors The backbone of FX is shown as tube and colored by secondary structure, with the residues shown by balls and sticks and colored in magenta. In comparison, the residues and backbones of other coagulation factors are shown by lines and colored in orange for FⅡ, yellow for FⅦ, brown for FⅨ, green for FⅪ and blue for FⅫ.
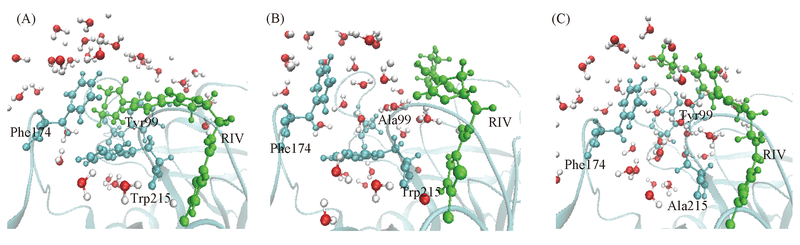
Fig.4 Water molecules around S4 pockets with WT(A), Y99A(B) and W215A(C) RIV is shown by CPK in green; the key residues Tyr99, Phe174, Trp215 and Asp189 are shown by CPK in cyan; the backbone of FXa is shown by cartoon.
| Species | E/(kJ·mol-1) | ||
|---|---|---|---|
| Wild type | Y99A | W215A | |
| van der Waals energy | -188.141±13.438 | -145.748±21.088 | -108.356±19.709 |
| Electrostatic energy | -52.079±15.698 | -39.173±31.740 | -22.613±25.232 |
| Polar solvation energy | 175.618±22.205 | 127.588±42.215 | 89.180±38.211 |
| SASA energy | -20.516±1.214 | -14.328±2.153 | -12.718±1.969 |
| Binding energy | -85.119±15.624 | -71.660±21.972 | -54.507±20.198 |
| Species | E/(kJ·mol-1) | ||
|---|---|---|---|
| Wild type | Y99A | W215A | |
| van der Waals energy | -188.141±13.438 | -145.748±21.088 | -108.356±19.709 |
| Electrostatic energy | -52.079±15.698 | -39.173±31.740 | -22.613±25.232 |
| Polar solvation energy | 175.618±22.205 | 127.588±42.215 | 89.180±38.211 |
| SASA energy | -20.516±1.214 | -14.328±2.153 | -12.718±1.969 |
| Binding energy | -85.119±15.624 | -71.660±21.972 | -54.507±20.198 |
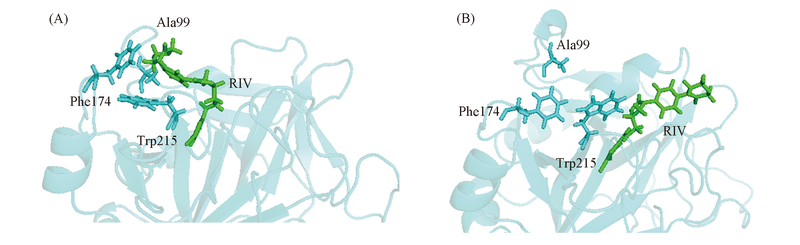
Fig.5 Swing of Trp215 side chain in Y99A mutant The side chain of Trp215 swings from the bottom of S4 pocket(A) to the entry of S1 pocket(B). RIV is shown by sticks in green, the key residues Tyr99, Phe174, Trp215 and Asp189 are shown by sticks in cyan, while the backbone of FXa is shown by cartoon in cyan.
| Residue | E/(kJ·mol-1) | ||
|---|---|---|---|
| Wild type | Y99A | W215A | |
| Y/A99 | -5.088±0.292 | 0.016±0.015 | -0.609±0.189 |
| W/A215 | -9.990±0.259 | -6.101±0.204 | 0.216±0.072 |
| Residue | E/(kJ·mol-1) | ||
|---|---|---|---|
| Wild type | Y99A | W215A | |
| Y/A99 | -5.088±0.292 | 0.016±0.015 | -0.609±0.189 |
| W/A215 | -9.990±0.259 | -6.101±0.204 | 0.216±0.072 |
| [1] | Macfarlane R. G ., Nature 1964, 202, 498— 499 |
| [2] | Davie E. W. Ratnoff O. D. ., Science, 1964, 145, 1310— 1312 |
| [3] | Davie E. W., Fujikawa K., Kisiel W ., Biochemistry 1991, 30, 10363— 10370 |
| [4] | Mann K., Nesheim M., Church W., Haley P., Krishnaswamy S ., Blood 1990, 76, 1— 16 |
| [5] | Samama M., Amiral J., Guinet C., Flem L. L., Seghatchian J., ,. Expert Review of Hematology 2013, 6, 155— 164 |
| [6] | Padmanabhan K., Padmanabhan K. P., Tulinsky A., Park C. H., Bode W., Huber R., Blankenship D. T., Cardin A. D., Kisiel W., ,. J. Mole. Biology 1993, 232, 947— 966 |
| [7] | Lee Y., Player M. R., ,. Medicinal Research Reviews 2011, 31, 202— 283 |
| [8] | Wang W., Yuan J., Fu X., Meng F., Zhang S., Xu W., Xu Y., Huang C ., Molecules 2016, 21( 4), 491 |
| [9] | Perzborn E., Roehrig S., Straub A., Kubitza D., Misselwitz F., , Nature Reviews Drug Discovery 2011, 10, 61— 75 |
| [10] | Wong P. C Pinto D. J. P Zhang D. , J. Thrombosis Thrombolysis 2011, 31, 478— 492 |
| [11] | Maignan S ., Guilloteau J ., Choisledeski Y.M Becker M. R Ewing W. R Pauls H. W Spada A. P Mikol V., ., J. Med.Chem 2003, 46, 685— 690 |
| [12] | Verhoef D ., Visscher K. M Vosmeer C. R Cheung K. L Reitsma P. H Geerke D. P Bos M. H. A ' , Nature Communications 2017, 8, 528 |
| [13] | Roehrig S., Straub A., Pohlmann J., Lampe T., Pernerstorfer J., Schlemmer K., Reinemer P., Perzborn E., ., J. Med.Chem 2005, 48, 5900— 5908 |
| [14] | Nazare M ., Will D. W Matter H ., Schreuder H ., Ritter K ., Urmann M ., Essrich M ., Bauer A ., Wagner M ., Czech J ., J. Med.Chem 2005, 48, 4511— 4525 |
| [15] | Abdelazeim S ., Oliva R ., Chermak E ., De Cristofaro R ., Cavallo L. ,. Biochem., 2014, 53, 6992— 7001 |
| [16] | Rezaie A. R . ,. Thrombosis and Haemostasis , 2003, 89, 112— 121 |
| [17] | Rezaie A. R J. Biolog. Chem ., 1996, 271, 23807— 23814 |
| [18] | Kumar S., Stecher G., Tamura K., Molecular Biology and Evolution, 2016, 33, 1870— 1874 |
| [19] | Larkin M. A Blackshields G ., Brown N. P Chenna R ., Mcgettigan P. A Mcwilliam H. ., Valentin F ., Wallace I. M Wilm A ., Lopez R. ,. Bioinformatics, 2007, 23, 2947— 2948 |
| [20] | Crooks G. E Hon G. C Chandonia J. M . Brenner S. E. , Genome Research , 2004, 14, 1188— 1190 |
| [21] | Schneider T. D Stephens R. M , Nucleic Acids Research 1990, 18, 6097— 6100 |
| [22] | Abraham M.J Murtola T ., Schulz R ., Pall S ., Smith J. C Hess B ., Lindahl E. ,. SoftwareX, 2015, 1, 19— 25 |
| [23] | Klauda J. B Venable R. M Freites J. A Connor J. W. O Tobias D. J Mondragonramirez C ., Vorobyov I., Mackerell A. D Pastor R. W , J. Phys. Chem.B 2010, 114, 7830— 7843 |
| [24] | Jorgensen W. L Chandrasekhar J ., Madura J. D Impey R ., Klein M. L ., J. Chem. Phys 1983, 79, 926— 935 |
| [25] | Vanommeslaeghe K., Hatcher E., Acharya C., Kundu S., Zhong S., Shim J., Darian E., Guvench O., Lopes P.E. M ., Vorobyov I ., J. Computat.Chem 2009, 31, 671— 690 |
| [26] | Vanommeslaeghe K ., Ghosh J ., Polani N. K ., Sheetz M ., Pamidighantam S. V Connolly J. W. D MacKerell A. D. Jr ., Biophys.J 2011, 100, 611— 611 |
| [27] | Vanommeslaeghe K ., Raman E. P Mackerell D. A ., J. Chem. Inform.Model 2012, 52, 3155— 3168 |
| [28] | Hoover W. G.., Phys. Rev. A: Gen. Phys., 1985, 31, 1695— 1697 |
| [29] | Parrinello M., Rahman A., ., J. Appl.Phys 1981, 52, 7182— 7190 |
| [30] | Kumari R Kumar R., Lynn A. M. . J. Chem. Inform. Model., 2014, 54, 1951— 1962 |
| [31] | Sichler K ., Kopetzki E ., Huber R ., Bode W ., Hopfner K ., Brandstetter H ., ., J. Biolog.Chem 2003, 278, 4121— 4126 |
| [32] | Soejima K., Yuguchi M., Mizuguchi J., Tomokiyo K., Nakashima T., Nakagaki T., Iwanaga S., ., J. Biolog.Chem 2002, 277, 49027— 49035 |
| [33] | Neuenschwander P. F Williamson S. R Nalian A. , Bakerdeadmond K. J ., J. Biolog.Chem 2006, 281, 23066— 23074 |
| [34] | Wang J., Hao P., Li Y., Dai J., Li X., , J. Molecular Modeling 2012, 18, 2717— 2725 |
| [1] | GAO Zhiwei, LI Junwei, SHI Sai, FU Qiang, JIA Junru, AN Hailong. Analysis of Gating Characteristics of TRPM8 Channel Based on Molecular Dynamics [J]. Chem. J. Chinese Universities, 2022, 43(6): 20220080. |
| [2] | HU Bo, ZHU Haochen. Dielectric Constant of Confined Water in a Bilayer Graphene Oxide Nanosystem [J]. Chem. J. Chinese Universities, 2022, 43(2): 20210614. |
| [3] | ZHANG Mi, TIAN Yafeng, GAO Keli, HOU Hua, WANG Baoshan. Molecular Dynamics Simulation of the Physicochemical Properties of Trifluoromethanesulfonyl Fluoride Dielectrics [J]. Chem. J. Chinese Universities, 2022, 43(11): 20220424. |
| [4] | LI Congcong, LIU Minghao, HAN Jiarui, ZHU Jingxuan, HAN Weiwei, LI Wannan. Theoretical Study of the Catalytic Activity of VmoLac Non-specific Substrates Based on Molecular Dynamics Simulations [J]. Chem. J. Chinese Universities, 2021, 42(8): 2518. |
| [5] | LEI Xiaotong, JIN Yiqing, MENG Xuanyu. Prediction of the Binding Site of PIP2 in the TREK-1 Channel Based on Molecular Modeling [J]. Chem. J. Chinese Universities, 2021, 42(8): 2550. |
| [6] | ZENG Yonghui, YAN Tianying. Vibrational Density of States Analysis of Proton Hydration Structure [J]. Chem. J. Chinese Universities, 2021, 42(6): 1855. |
| [7] | LIU Aiqing, XU Wensheng, XU Xiaolei, CHEN Jizhong, AN Lijia. Molecular Dynamics Simulation of Polymer/rod Nanocomposite [J]. Chem. J. Chinese Universities, 2021, 42(3): 875. |
| [8] | QI Renrui, LI Minghao, CHANG Hao, FU Xueqi, GAO Bo, HAN Weiwei, HAN Lu, LI Wannan. Theoretical Study on the Unbinding Pathway of Xanthine Oxidase Inhibitors Based on Steered Molecular Dynamics Simulation [J]. Chem. J. Chinese Universities, 2021, 42(3): 758. |
| [9] | MA Yucong, FAN Baomin, WANG Manman, YANG Biao, HAO Hua, SUN Hui, ZHANG Huijuan. Two-step Preparation of Trazodone and Its Corrosion Inhibition Mechanism for Carbon Steel [J]. Chem. J. Chinese Universities, 2019, 40(8): 1706. |
| [10] | ZHANG Zhang,WANG Dong,WANG Xiaolei,XU Yan. Regulation of Ester Synthesis Activity of Rhizopus chinensis Lipase† [J]. Chem. J. Chinese Universities, 2019, 40(4): 747. |
| [11] | MA Lan,RONG Jingjing,ZHU Youliang,HUANG Yineng,SUN Zhaoyan. Simulation on the Dynamic Process of Formation of Particle Cluster by Generalized Exponential Model† [J]. Chem. J. Chinese Universities, 2019, 40(1): 195. |
| [12] | ZHU Jingxuan,YU Zhengfei,LIU Ye,ZHAN Dongling,HAN Jiarui,TIAN Xiaopian,HAN Weiwei. Exploration of Increasing the Non-specificity Substrates Activity for the Phosphotriesterase-like Lactonase Using Molecular Dynamics Simulations† [J]. Chem. J. Chinese Universities, 2019, 40(1): 138. |
| [13] | WU Hongmei,LI Huiting,LI Yongcheng,WANG Hongqing,WANG Meng. Using Group Contribution Method and Molecular Dynamics to Predict the Glass Transition Temperature of Poly(p-phenylene isophthalamide)† [J]. Chem. J. Chinese Universities, 2019, 40(1): 180. |
| [14] | LIU Yanfang, YANG Hua, ZHANG Hui. Molecular Dynamics Simulation on the Orientation of Alkane Mixture on Graphene† [J]. Chem. J. Chinese Universities, 2018, 39(8): 1729. |
| [15] | LIU Haichun, LU Shuai, ZHANG Yanmin, ZHOU Weineng, YIN Lingfeng, ZHU Lu, ZHAO Junnan, LU Tao, CHEN Yadong. Molecular Dynamics Simulation of the Selectivity of Fedratinib Complex with JAK2/JAK3† [J]. Chem. J. Chinese Universities, 2018, 39(7): 1540. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
