

Chem. J. Chinese Universities ›› 2019, Vol. 40 ›› Issue (6): 1229.doi: 10.7503/cjcu20180837
• Physical Chemistry • Previous Articles Next Articles
SUN Haofan1,2, ZHANG Lingyi1, PATRICK Norman2, ZHANG Weibing1( )
)
Received:2018-12-13
Online:2019-06-10
Published:2019-04-04
Supported by:CLC Number:
TrendMD:
SUN Haofan,ZHANG Lingyi,PATRICK Norman,ZHANG Weibing. Molecular Dynamic of Various DNA Sequences Binding of Dithienylethenes†[J]. Chem. J. Chinese Universities, 2019, 40(6): 1229.
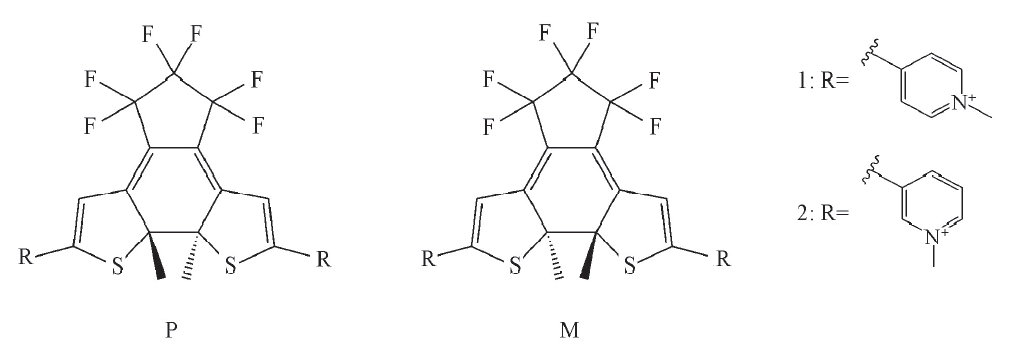
Fig.1 Illustration of molecular structures for the dithienylethene(DTE) derivatives under study in their P and M enantiomeric forms as for closed forms DTE1 and DTE2 differ in the para and meta position of the nitrogen in the pyridinium ligand.
| Conformation | Binding position | |||
|---|---|---|---|---|
| MaGB | MaGI | MiGB | MiGI | |
| P-DTE1 | MaGB | MaGI | MiGB | MiGI |
| M-DTE1 | MiGB | MaGI | MiGB | MiGI |
| P-DTE2 | Bottom binding | MaGI | MiGB | MiGB |
| M-DTE2 | MiGB | MaGI | MiGB | MiGB |
Table 1 Final binding positions from different initial positions between DTE and DNA(A-T pairs)
| Conformation | Binding position | |||
|---|---|---|---|---|
| MaGB | MaGI | MiGB | MiGI | |
| P-DTE1 | MaGB | MaGI | MiGB | MiGI |
| M-DTE1 | MiGB | MaGI | MiGB | MiGI |
| P-DTE2 | Bottom binding | MaGI | MiGB | MiGB |
| M-DTE2 | MiGB | MaGI | MiGB | MiGB |
| Conformation | Binding position | |||
|---|---|---|---|---|
| MaGB | MaGI | MiGB | MiGI | |
| P-DTE1 | MiGB | MaGI | MiGB | MiGI |
| M-DTE1 | MiGB | MaGI | MiGB | MiGI |
| P-DTE2 | Bottom binding | MaGI | MiGB | MiGI |
| M-DTE2 | Bottom binding | MaGI | MiGB | Bottom binding |
Table 2 Final binding positions from different initial positions between DTE and DNA(C-G pairs)
| Conformation | Binding position | |||
|---|---|---|---|---|
| MaGB | MaGI | MiGB | MiGI | |
| P-DTE1 | MiGB | MaGI | MiGB | MiGI |
| M-DTE1 | MiGB | MaGI | MiGB | MiGI |
| P-DTE2 | Bottom binding | MaGI | MiGB | MiGI |
| M-DTE2 | Bottom binding | MaGI | MiGB | Bottom binding |
| DNA | Conformation | Final position | Ecoul/(kJ·mol-1) | EvdW/(kJ·mol-1) |
|---|---|---|---|---|
| (6A-T +6C-G)×2 | P-DTE1 | No favorable binding | -259±75 | -84±21 |
| M-DTE1 | MiGB(A-T) | -828±71 | -217±17 | |
| P-DTE2 | MiGB(between A-T and C-G) | -660±100 | -113±21 | |
| M-DTE2 | MiGB(between A-T and C-G) | -598±88 | -121±25 |
Table 3 Final position and average interaction energies between DTE and DNA
| DNA | Conformation | Final position | Ecoul/(kJ·mol-1) | EvdW/(kJ·mol-1) |
|---|---|---|---|---|
| (6A-T +6C-G)×2 | P-DTE1 | No favorable binding | -259±75 | -84±21 |
| M-DTE1 | MiGB(A-T) | -828±71 | -217±17 | |
| P-DTE2 | MiGB(between A-T and C-G) | -660±100 | -113±21 | |
| M-DTE2 | MiGB(between A-T and C-G) | -598±88 | -121±25 |
| DNA | Conformation | Final position | Ecoul/(kJ·mol-1) | EvdW/(kJ·mol-1) |
|---|---|---|---|---|
| (6A-T +6C-G) ×2 | P-DTE1 | MiGB(A-T) | -493±42 | -138±21 |
| M-DTE1 | MiGB(C-G) | -623±92 | -171±29 | |
| P-DTE2 | MiGB(A-T) | -702±71 | -142±21 | |
| M-DTE2 | No favorable binding | -38±67 | -13±29 | |
| 12A-T +12C-G | P-DTE1 | MiGB(A-T) | -727±105 | -176±25 |
| M-DTE1 | MiGB(A-T) | -869±63 | -226±13 | |
| P-DTE2 | MiGB(C-G) | -359±67 | -109±42 | |
| M-DTE2 | Bottom binding | -150±109 | -75±13 |
Table 4 Final position and average interaction energies among DTE and DNA
| DNA | Conformation | Final position | Ecoul/(kJ·mol-1) | EvdW/(kJ·mol-1) |
|---|---|---|---|---|
| (6A-T +6C-G) ×2 | P-DTE1 | MiGB(A-T) | -493±42 | -138±21 |
| M-DTE1 | MiGB(C-G) | -623±92 | -171±29 | |
| P-DTE2 | MiGB(A-T) | -702±71 | -142±21 | |
| M-DTE2 | No favorable binding | -38±67 | -13±29 | |
| 12A-T +12C-G | P-DTE1 | MiGB(A-T) | -727±105 | -176±25 |
| M-DTE1 | MiGB(A-T) | -869±63 | -226±13 | |
| P-DTE2 | MiGB(C-G) | -359±67 | -109±42 | |
| M-DTE2 | Bottom binding | -150±109 | -75±13 |
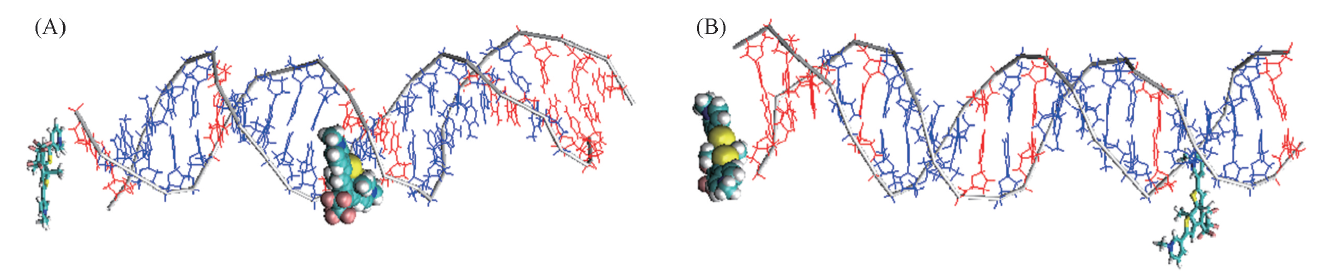
Fig.4 Two parallel sets of unbiased docking for most favorable binding modes to random sequence DNA of P-DTE1(A) and P-DTE2(B) VDW drawing method: DTE1; licorice drawing method: DTE2; blue: A-T pairs, red: C-G pairs.
| DNA | Conformation | Final position | Ecoul/(kJ·mol-1) | EvdW/(kJ·mol-1) |
|---|---|---|---|---|
| A-T/C-G mixture pairs | P-DTE1 | MiGB(A-T) | -594±54 | -150±13 |
| P-DTE2 | Bottom binding | -96±33 | -71±13 | |
| M-DTE1 | Bottom binding | -171±67 | -54±17 | |
| M-DTE2 | No favorable binding | -314±222 | -25±21 | |
| Random pairs | P-DTE1 | MiGB/bottom binding | (-598±63/-146±230) | (-155±17/-12±25) |
| P-DTE2 | Bottom binding/MiGB | (-113±42/-556±46) | (-71±8/-117±13) | |
| M-DTE1 | MiGB | -602±63 | -130±25 | |
| M-DTE2 | MiGB | -564±33 | -121±13 |
Table 5 Final position and average interaction energies between P-DTE or M-DTE and DNA
| DNA | Conformation | Final position | Ecoul/(kJ·mol-1) | EvdW/(kJ·mol-1) |
|---|---|---|---|---|
| A-T/C-G mixture pairs | P-DTE1 | MiGB(A-T) | -594±54 | -150±13 |
| P-DTE2 | Bottom binding | -96±33 | -71±13 | |
| M-DTE1 | Bottom binding | -171±67 | -54±17 | |
| M-DTE2 | No favorable binding | -314±222 | -25±21 | |
| Random pairs | P-DTE1 | MiGB/bottom binding | (-598±63/-146±230) | (-155±17/-12±25) |
| P-DTE2 | Bottom binding/MiGB | (-113±42/-556±46) | (-71±8/-117±13) | |
| M-DTE1 | MiGB | -602±63 | -130±25 | |
| M-DTE2 | MiGB | -564±33 | -121±13 |
| [1] | Wishart D. S., Nat. Rev. Drug Discov., 2016, 15, 473-484 |
| [2] | Nau H., Hauck R. S., Ehlers K., Pharmacol.Toxico.,1991, 69, 310-321 |
| [3] | Zheng Y., Qiang X., Xu R., Song Q., Tian C., Liu H., Li W., Tan Z., Deng Y., Bioorg. Chem.,2018, 78, 298-306 |
| [4] | Kasatani K., Kambe S., Irie M., J. Photoch. Photobio. A,1999, 122, 11-15 |
| [5] | Irie M., Fukaminato T., Matsuda K., Kobatake S., Chem.Rev.,2014, 114, 12174-12277 |
| [6] | Zhang J., Tian H., Adv. Opt. Mater.,2018, 6, 1701278 |
| [7] | Delbaere S., Vermeersch G., J. Photoch. Photobio. C,2008, 9, 61-80 |
| [8] | Irie M., Kobatake S., Horichi M., Science,2001, 291, 1769-1772 |
| [9] | Kay E. R., Leigh D. A., Zerbetto F., Angew. Chem. Int. Ed.,2007, 46, 72-191 |
| [10] | Liddell P. A., Kodis G., Moore A. L., Moore T. A., Gust D., J. Am. Chem. Soc.,2002, 124, 7668-7669 |
| [11] | Irie M., Chem. Rev., 2000, 100, 1685-1716 |
| [12] | Pace T. C., Müller V., Li S., Lincoln P., Andréasson J., Angew. Chem. Int. Ed.,2013, 125, 4489-4492 |
| [13] | Linares M., Sun H., Biler M., Andréasson J., Norman P., Phys. Chem. Chem. Phys.,2019, 21, 3637-3643 |
| [14] | Gilat S. L., Kawai S. H., Lehn J. M., Chem. Eur. J.,1995, 1, 275-284 |
| [15] | Vosko S. H., Wilk L., Nusair M., Can. J. Phys.,1980, 58, 1200-1211 |
| [16] | Dunning T. H. Jr., J. Chem. Phys., 1989, 90, 1007-1023 |
| [17] | Dupradeau F. Y., Pigache A., Zaffran T., Savineau C., Lelong R., Grivel N., Lelong D., Rosanski W., Cieplak P., Phys. Chem. Chem. Phys.,2010, 12, 7821-7839 |
| [18] | Becke A. D., J. Chem. Phys., 1993, 98, 5648-5652 |
| [19] | Lee C., Yang W., Parr R. G., Phys. Rev. B,1988, 37, 785-789 |
| [20] | Pearlman D. A., Case D. A., Caldwell J. W., Ross W. S., Cheatham III T. E., DeBolt S., Ferguson D., Seibel G., Kollman P., Comput. Phys. Commun.,1995, 91, 1-41 |
| [21] | Wang J., Wolf R. M., Caldwell J. W., Kollman P. A., Case D. A., J. Comput. Chem.,2004, 25, 1157-1174 |
| [22] | Ivani I., Dans P. D., Noy A., Pérez A., Faustino I., Walther J., Andrio P., Goñi R., Balaceanu A., Portella G., Nat. Methods,2016, 13, 55-58 |
| [23] | Jorgensen W. L., Chandrasekhar J., Madura J. D., Impey R. W., Klein M. L., J. Chem. Phys.,1983, 79, 926-935 |
| [24] | Berendsen H. J., Postma J. V., van Gunsteren W. F., DiNola A., Haak J., J. Chem. Phys.,1984, 81, 3684-3690 |
| [25] | Grimme S., Antony J., Ehrlich S., Krieg H., J. Chem. Phys.,2010, 132, 154104 |
| [26] | Nagy Á., Phys. Rep., 1998, 298, 1-79 |
| [27] | Miertuš S., Scrocco E., Tomasi J., Chem. Phys.,1981, 55, 117-129 |
| [1] | LIU Suyu, DING Fei, LI Qian, FAN Chunhai, FENG Jing. Azobenzene-integrated DNA Nanomachine [J]. Chem. J. Chinese Universities, 2022, 43(8): 20220122. |
| [2] | WU Yushuai, SHANG Yingxu, JIANG Qiao, DING Baoquan. Research Progress of Controllable Self-assembled DNA Origami Structure as Drug Carrier [J]. Chem. J. Chinese Universities, 2022, 43(8): 20220179. |
| [3] | WANG Junyang, LIU Zheng, ZHANG Qian, SUN Chunyan, LI Hongxia. Application of DNA Silver Nanoclusters in the Fluorescence Biosensors based on Functional Nucleic Acids [J]. Chem. J. Chinese Universities, 2022, 43(6): 20220010. |
| [4] | GAO Zhiwei, LI Junwei, SHI Sai, FU Qiang, JIA Junru, AN Hailong. Analysis of Gating Characteristics of TRPM8 Channel Based on Molecular Dynamics [J]. Chem. J. Chinese Universities, 2022, 43(6): 20220080. |
| [5] | ZENG Xianyang, ZHAO Xi, HUANG Xuri. Mechanism of Inhibition of Glucose and Proton Cotransport Protein GlcPSe by Cytochalasin B [J]. Chem. J. Chinese Universities, 2022, 43(4): 20210822. |
| [6] | CHEN Hanxiang, BIAN Shaoju, HU Bin, LI Wu. Molecular Simulation of the Osmotic Pressures for LiCl-NaCl-KCl-H2O Solution System [J]. Chem. J. Chinese Universities, 2022, 43(3): 20210727. |
| [7] | HU Bo, ZHU Haochen. Dielectric Constant of Confined Water in a Bilayer Graphene Oxide Nanosystem [J]. Chem. J. Chinese Universities, 2022, 43(2): 20210614. |
| [8] | ZHANG Mi, TIAN Yafeng, GAO Keli, HOU Hua, WANG Baoshan. Molecular Dynamics Simulation of the Physicochemical Properties of Trifluoromethanesulfonyl Fluoride Dielectrics [J]. Chem. J. Chinese Universities, 2022, 43(11): 20220424. |
| [9] | ZHANG Lingyu, ZHANG Jilong, QU Zexing. Dynamics Study of Intramolecular Vibrational Energy Redistribution in RDX Molecule [J]. Chem. J. Chinese Universities, 2022, 43(10): 20220393. |
| [10] | LEI Xiaotong, JIN Yiqing, MENG Xuanyu. Prediction of the Binding Site of PIP2 in the TREK-1 Channel Based on Molecular Modeling [J]. Chem. J. Chinese Universities, 2021, 42(8): 2550. |
| [11] | LI Congcong, LIU Minghao, HAN Jiarui, ZHU Jingxuan, HAN Weiwei, LI Wannan. Theoretical Study of the Catalytic Activity of VmoLac Non-specific Substrates Based on Molecular Dynamics Simulations [J]. Chem. J. Chinese Universities, 2021, 42(8): 2518. |
| [12] | LIU Shasha, ZHANG Heng, YUAN Shiling, LIU Chengbu. Molecular Dynamics Simulation of Pulsed Electric Field O/W Emulsion Demulsification [J]. Chem. J. Chinese Universities, 2021, 42(7): 2170. |
| [13] | ZENG Yonghui, YAN Tianying. Vibrational Density of States Analysis of Proton Hydration Structure [J]. Chem. J. Chinese Universities, 2021, 42(6): 1855. |
| [14] | WU Yangyi, CHEN Jianping, Ai Yijing, WANG Qingxiang, GAO Fei, GAO Feng. Synthesis of 2-(2-Hydroxy-3-methoxyphenyl)-C60 and Its Application for Sensing of Cauliflower Mosaic Virus 35S Promotor [J]. Chem. J. Chinese Universities, 2021, 42(6): 1754. |
| [15] | QI Renrui, LI Minghao, CHANG Hao, FU Xueqi, GAO Bo, HAN Weiwei, HAN Lu, LI Wannan. Theoretical Study on the Unbinding Pathway of Xanthine Oxidase Inhibitors Based on Steered Molecular Dynamics Simulation [J]. Chem. J. Chinese Universities, 2021, 42(3): 758. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||