

Chem. J. Chinese Universities ›› 2019, Vol. 40 ›› Issue (4): 705.doi: 10.7503/cjcu20180680
• Organic Chemistry • Previous Articles Next Articles
CHANG Junpeng, ZHAO Jiarui, CHEN Sijia, MENG Kai, SHI Weini, LI Ruifang*( )
)
Received:2018-10-09
Online:2019-04-03
Published:2019-01-09
Contact:
LI Ruifang
E-mail:lrf@haut.edu.cn
Supported by:CLC Number:
TrendMD:
CHANG Junpeng,ZHAO Jiarui,CHEN Sijia,MENG Kai,SHI Weini,LI Ruifang. Structure-activity Relationship of Antimicrobial Peptide SAMP1 and Its Analog Peptides†[J]. Chem. J. Chinese Universities, 2019, 40(4): 705.
| Peptide | Sequence* | Molecular mass | tR/min | |
|---|---|---|---|---|
| Calculated | Observed | |||
| SAMP1 | VRLLKKKI | 1218.57 | 1218.2 | 10.5 |
| SAMP1-A1 | VLLIRKKK | 1218.57 | 1218.4 | 10.5 |
| SAMP1-A2 | RVKLKLKI | 1218.57 | 1218.4 | 10.5 |
| SAMP1-A3 | RKKKVLLI | 1218.57 | 1218.4 | 10.6 |
| SAMP1-A4 | VRLLRRRI | 1302.64 | 1302.2 | 10.3 |
| SAMP1-A5 | IRIIKKKI | 1233.60 | 1232.2 | 12.7 |
| SAMP1-A6 | LRLLKKKL | 1233.60 | 1232.2 | 9.8 |
| SAMP1-A7 | VRVVKKKV | 1176.49 | 1175.2 | 11.4 |
| SAMP1-A8 | WRWWKKKW | 1524.83 | 1524.2 | 8.4 |
Table 1 Amino acid sequences, molecular mass and HPLC retention time of SAMP1 and its analog peptides
| Peptide | Sequence* | Molecular mass | tR/min | |
|---|---|---|---|---|
| Calculated | Observed | |||
| SAMP1 | VRLLKKKI | 1218.57 | 1218.2 | 10.5 |
| SAMP1-A1 | VLLIRKKK | 1218.57 | 1218.4 | 10.5 |
| SAMP1-A2 | RVKLKLKI | 1218.57 | 1218.4 | 10.5 |
| SAMP1-A3 | RKKKVLLI | 1218.57 | 1218.4 | 10.6 |
| SAMP1-A4 | VRLLRRRI | 1302.64 | 1302.2 | 10.3 |
| SAMP1-A5 | IRIIKKKI | 1233.60 | 1232.2 | 12.7 |
| SAMP1-A6 | LRLLKKKL | 1233.60 | 1232.2 | 9.8 |
| SAMP1-A7 | VRVVKKKV | 1176.49 | 1175.2 | 11.4 |
| SAMP1-A8 | WRWWKKKW | 1524.83 | 1524.2 | 8.4 |
| Peptide | pI | GRAVY | Instability index | Fat coefficient | Half-life/h | ||
|---|---|---|---|---|---|---|---|
| Lactating cell | Yeast cell | E. coli | |||||
| SAMP1 | 11.26 | 0.013 | -12.48 | 182.5 | 100 | >20 | >10 |
| SAMP1-A1 | 11.26 | 0.013 | 8.75 | 182.5 | 100 | >20 | >10 |
| SAMP1-A2 | 11.26 | 0.013 | -47.91 | 182.5 | 1 | 0.03 | 0.03 |
| SAMP1-A3 | 11.26 | 0.013 | -1.86 | 182.5 | 1 | 0.03 | 0.03 |
| SAMP1-A4 | 12.48 | -0.212 | 176.02 | 182.5 | 100 | >20 | >10 |
| SAMP1-A5 | 11.26 | 0.225 | -12.48 | 195 | 20 | 0.5 | >10 |
| SAMP1-A6 | 11.26 | -0.125 | 11.60 | 195 | 5.5 | 0.05 | 0.05 |
| SAMP1-A7 | 11.26 | 0.075 | -5.46 | 145 | 100 | >20 | >10 |
| SAMP1-A8 | 11.26 | -2.475 | 80.35 | 0 | 2.8 | 0.05 | 0.05 |
Table 2 Physicochemical properties of SAMP1 and its analog peptides
| Peptide | pI | GRAVY | Instability index | Fat coefficient | Half-life/h | ||
|---|---|---|---|---|---|---|---|
| Lactating cell | Yeast cell | E. coli | |||||
| SAMP1 | 11.26 | 0.013 | -12.48 | 182.5 | 100 | >20 | >10 |
| SAMP1-A1 | 11.26 | 0.013 | 8.75 | 182.5 | 100 | >20 | >10 |
| SAMP1-A2 | 11.26 | 0.013 | -47.91 | 182.5 | 1 | 0.03 | 0.03 |
| SAMP1-A3 | 11.26 | 0.013 | -1.86 | 182.5 | 1 | 0.03 | 0.03 |
| SAMP1-A4 | 12.48 | -0.212 | 176.02 | 182.5 | 100 | >20 | >10 |
| SAMP1-A5 | 11.26 | 0.225 | -12.48 | 195 | 20 | 0.5 | >10 |
| SAMP1-A6 | 11.26 | -0.125 | 11.60 | 195 | 5.5 | 0.05 | 0.05 |
| SAMP1-A7 | 11.26 | 0.075 | -5.46 | 145 | 100 | >20 | >10 |
| SAMP1-A8 | 11.26 | -2.475 | 80.35 | 0 | 2.8 | 0.05 | 0.05 |
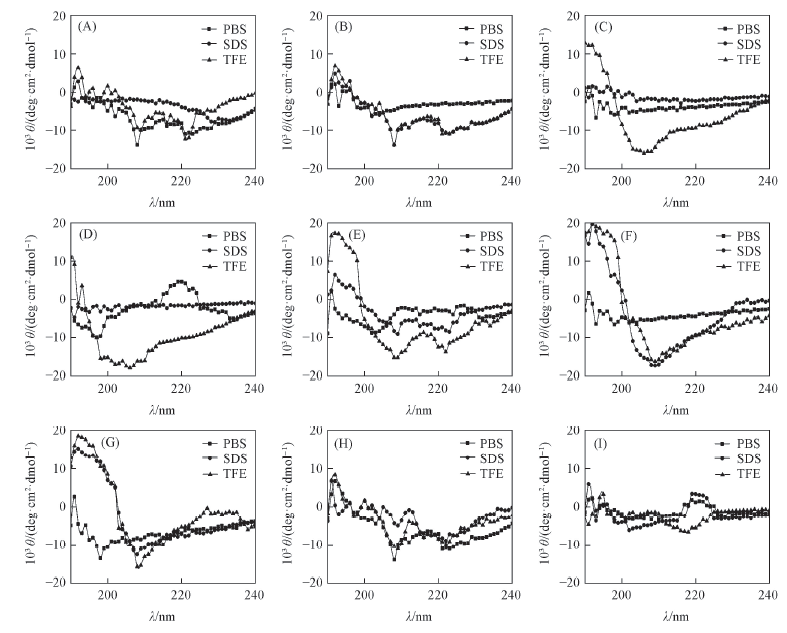
Fig.1 CD spectra of SAMP1 and its analog peptides in PBS, SDS and TFE solutions (A) SAMP1; (B) SAMP1-A1; (C) SAMP1-A2; (D)SAMP1-A3; (E) SAMP1-A4; (F) SAMP1-A5; (G) SAMP1-A6; (H) SAMP1-A7; (I) SAMP1-A8.
| Peptide | Solution | α-Helix(%) | β-Sheet(%) | β-Turn(%) | Random(%) |
|---|---|---|---|---|---|
| SAMP1 | PBS | 54.8 | 6.1 | 16.9 | 22.1 |
| SDS | 39.0 | 6.9 | 20.5 | 33.7 | |
| TFE | 56.3 | 5.1 | 16.6 | 22.0 | |
| SAMP1-A1 | PBS | 20.6 | 24.7 | 19.0 | 35.7 |
| SDS | 54.7 | 5.8 | 17.0 | 22.3 | |
| TFE | 55.0 | 5.7 | 16.9 | 22.3 | |
| SAMP1-A2 | PBS | 34.2 | 13.5 | 22.2 | 30.0 |
| SDS | 18.6 | 29.4 | 22.0 | 30.0 | |
| TFE | 55.0 | 5.7 | 16.9 | 22.3 | |
| SAMP1-A3 | PBS | 34.2 | 13.5 | 22.2 | 30.0 |
| SDS | 18.6 | 29.4 | 22.0 | 30.0 | |
| TFE | 66.0 | 2.1 | 10.6 | 21.1 | |
| SAMP1-A4 | PBS | 19.7 | 23.9 | 20.0 | 36.3 |
| SDS | 42.4 | 6.7 | 29.2 | 21.6 | |
| TFE | 80.8 | 0.5 | 5.0 | 14.3 | |
| SAMP1-A5 | PBS | 31.9 | 15.5 | 21.7 | 31.0 |
| SDS | 74.0 | 3.4 | 6.0 | 16.5 | |
| TFE | 79.8 | 0.4 | 5.4 | 14.5 | |
| SAMP1-A6 | PBS | 33.6 | 7.4 | 21.9 | 37.1 |
| SDS | 78.8 | 0.7 | 5.2 | 15.2 | |
| TFE | 80.7 | 0.9 | 5.0 | 14.4 | |
| SAMP1-A7 | PBS | 55.1 | 5.5 | 17.1 | 22.3 |
| SDS | 56.2 | 5.1 | 16.6 | 22.0 | |
| TFE | 56.4 | 5.0 | 16.5 | 22.0 | |
| SAMP1-A8 | PBS | 20.9 | 26.1 | 21.7 | 31.3 |
| SDS | 25.9 | 16.9 | 17.6 | 39.7 | |
| TFE | 9.3 | 45.9 | 10.8 | 34.1 |
Table 3 Secondary structure content of SAMP1 and its analog peptides
| Peptide | Solution | α-Helix(%) | β-Sheet(%) | β-Turn(%) | Random(%) |
|---|---|---|---|---|---|
| SAMP1 | PBS | 54.8 | 6.1 | 16.9 | 22.1 |
| SDS | 39.0 | 6.9 | 20.5 | 33.7 | |
| TFE | 56.3 | 5.1 | 16.6 | 22.0 | |
| SAMP1-A1 | PBS | 20.6 | 24.7 | 19.0 | 35.7 |
| SDS | 54.7 | 5.8 | 17.0 | 22.3 | |
| TFE | 55.0 | 5.7 | 16.9 | 22.3 | |
| SAMP1-A2 | PBS | 34.2 | 13.5 | 22.2 | 30.0 |
| SDS | 18.6 | 29.4 | 22.0 | 30.0 | |
| TFE | 55.0 | 5.7 | 16.9 | 22.3 | |
| SAMP1-A3 | PBS | 34.2 | 13.5 | 22.2 | 30.0 |
| SDS | 18.6 | 29.4 | 22.0 | 30.0 | |
| TFE | 66.0 | 2.1 | 10.6 | 21.1 | |
| SAMP1-A4 | PBS | 19.7 | 23.9 | 20.0 | 36.3 |
| SDS | 42.4 | 6.7 | 29.2 | 21.6 | |
| TFE | 80.8 | 0.5 | 5.0 | 14.3 | |
| SAMP1-A5 | PBS | 31.9 | 15.5 | 21.7 | 31.0 |
| SDS | 74.0 | 3.4 | 6.0 | 16.5 | |
| TFE | 79.8 | 0.4 | 5.4 | 14.5 | |
| SAMP1-A6 | PBS | 33.6 | 7.4 | 21.9 | 37.1 |
| SDS | 78.8 | 0.7 | 5.2 | 15.2 | |
| TFE | 80.7 | 0.9 | 5.0 | 14.4 | |
| SAMP1-A7 | PBS | 55.1 | 5.5 | 17.1 | 22.3 |
| SDS | 56.2 | 5.1 | 16.6 | 22.0 | |
| TFE | 56.4 | 5.0 | 16.5 | 22.0 | |
| SAMP1-A8 | PBS | 20.9 | 26.1 | 21.7 | 31.3 |
| SDS | 25.9 | 16.9 | 17.6 | 39.7 | |
| TFE | 9.3 | 45.9 | 10.8 | 34.1 |
| Peptide | MIC/(μg·mL-1) | |||||
|---|---|---|---|---|---|---|
| C. tropicalis | E. coli | P. aeruginosa | B. subtilis | L. monocytogenes | S. saureus | |
| SAMP1 | 1.9 | 7.8 | 7.8 | 62.5 | 1.9 | 125 |
| SAMP1-A1 | 1.9 | 31.25 | 250 | 3.9 | 1.9 | >250 |
| SAMP1-A2 | 3.9 | 7.8 | 125 | 7.8 | 3.9 | >250 |
| SAMP1-A3 | 3.9 | 7.8 | 62.5 | 3.9 | 3.9 | 62.5 |
| SAMP1-A4 | 3.9 | 3.9 | 7.8 | 0.95 | 0.95 | 3.9 |
| SAMP1-A5 | 3.9 | >250 | >250 | >250 | >250 | >250 |
| SAMP1-A6 | 1.9 | 1.9 | 3.9 | 3.9 | 3.9 | 3.9 |
| SAMP1-A7 | 3.9 | >250 | >250 | >250 | >250 | >250 |
| SAMP1-A8 | 7.8 | 7.8 | 62.5 | 3.9 | 1.9 | 31.25 |
Table 4 Antimicrobial activity of SAMP1 and its analog peptides
| Peptide | MIC/(μg·mL-1) | |||||
|---|---|---|---|---|---|---|
| C. tropicalis | E. coli | P. aeruginosa | B. subtilis | L. monocytogenes | S. saureus | |
| SAMP1 | 1.9 | 7.8 | 7.8 | 62.5 | 1.9 | 125 |
| SAMP1-A1 | 1.9 | 31.25 | 250 | 3.9 | 1.9 | >250 |
| SAMP1-A2 | 3.9 | 7.8 | 125 | 7.8 | 3.9 | >250 |
| SAMP1-A3 | 3.9 | 7.8 | 62.5 | 3.9 | 3.9 | 62.5 |
| SAMP1-A4 | 3.9 | 3.9 | 7.8 | 0.95 | 0.95 | 3.9 |
| SAMP1-A5 | 3.9 | >250 | >250 | >250 | >250 | >250 |
| SAMP1-A6 | 1.9 | 1.9 | 3.9 | 3.9 | 3.9 | 3.9 |
| SAMP1-A7 | 3.9 | >250 | >250 | >250 | >250 | >250 |
| SAMP1-A8 | 7.8 | 7.8 | 62.5 | 3.9 | 1.9 | 31.25 |
| Peptide | Hemolysis ratio(%) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 250 μg/mL | 125 μg/mL | 62. 5 μg/mL | 31.25 μg/mL | 15.63 μg/mL | 7.8 μg/mL | 3.9 μg/mL | 1.9 μg/mL | 0.95 μg/mL | 0.475 μg/mL | |
| SAMP1 | 40.0 | 28.4 | 13.0 | 4.5 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A1 | 75.7 | 41.0 | 26.5 | 15.4 | 5.6 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A2 | 100 | 93.2 | 92.5 | 33.4 | 27.6 | 16.6 | 5.4 | 0 | 0 | 0 |
| SAMP1-A3 | 16.5 | 4.1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A4 | 30.3 | 7.3 | 2.7 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A5 | 36.0 | 16.4 | 6.5 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A6 | 100 | 52.0 | 22 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A7 | 16.7 | 1.3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A8 | 100 | 100 | 100 | 74.8 | 31.2 | 7.5 | 0 | 0 | 0 | 0 |
| Melittin | 100 | 100 | 100 | 100 | 86 | 58 | 49 | 21 | 0 | 0 |
Table 5 Hemolysis ratio of SAMP1 and its analog peptides
| Peptide | Hemolysis ratio(%) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 250 μg/mL | 125 μg/mL | 62. 5 μg/mL | 31.25 μg/mL | 15.63 μg/mL | 7.8 μg/mL | 3.9 μg/mL | 1.9 μg/mL | 0.95 μg/mL | 0.475 μg/mL | |
| SAMP1 | 40.0 | 28.4 | 13.0 | 4.5 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A1 | 75.7 | 41.0 | 26.5 | 15.4 | 5.6 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A2 | 100 | 93.2 | 92.5 | 33.4 | 27.6 | 16.6 | 5.4 | 0 | 0 | 0 |
| SAMP1-A3 | 16.5 | 4.1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A4 | 30.3 | 7.3 | 2.7 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A5 | 36.0 | 16.4 | 6.5 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A6 | 100 | 52.0 | 22 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A7 | 16.7 | 1.3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| SAMP1-A8 | 100 | 100 | 100 | 74.8 | 31.2 | 7.5 | 0 | 0 | 0 | 0 |
| Melittin | 100 | 100 | 100 | 100 | 86 | 58 | 49 | 21 | 0 | 0 |
| [1] | Bahar A. A., Ren D., Pharmaceuticals,2013, 6(12), 1543—1575 |
| [2] | Feng H. Y., Gao L., Ye X. H., Wang L., Xue Z. C., Kong J. M., Li L. Z., Chem. Res. Chinese Universities,2017, 33(1), 155—159 |
| [3] | Zhang Y. H., Li L. C., Yuan W. C., Zhang X. M., Chem. Res. Chinese Universities,2015, 31(3), 381—387 |
| [4] | Gaspar D., Veiga A. S., Castanho M. A. R. B., Front.Microbiol., 2013, 4(4), 294-1—6 |
| [5] | Wu D. D., Gao Y. F., Qi Y. M., Chen L. X., Ma Y. F., Li Y. Z., Cancer Lett., 2014, 351(1), 13—22 |
| [6] | Zhang Q. H., Wang Q., Shang T. T., Zhao Z. Y., Xu Y. Z., Hu J. H., China Anim. Hus. Veteri. Medic., 2014, 41(5), 163—167 |
| (张庆华, 王青, 尚田田, 赵志雨, 徐彦召, 胡建和. 中国畜牧兽医, 2014, 41(5), 163—167) | |
| [7] | Huang Y. B., Huang J. F., Chen Y. X., Protein Cell,2010, 1(2), 143—152 |
| [8] | Ghosh C., Manjunath G. B., Akkapeddi P., Yarlagadda V., Hoque J., Uppu D. S., J. Med.Chem., 2014, 57(4), 1428—1436 |
| [9] | Schmidtchen A., Pasupuleti M., Malmsten M., Adv. Colloid Interface., 2014, 205(12), 265—274 |
| [10] | Thaker H., Cankaya A., Scott R. W., Tew G. N., ACS Med. Chem.Lett., 2013, 4(5), 481—485 |
| [11] | Yin L. M., Edwards M. A., Li J., Yip C. M., Deber C. M., J. Biol.Chem., 2012, 287(10), 7738—7745 |
| [12] | De L. E., Rajabi M., Zou G., Pazgier M., Lu W., Febs Lett., 2009, 583(15), 2507—2512 |
| [13] | Hirose H., Takeuchi T., Osakada H., Pujals S., Katayama S., Nakase I., Kobayashi S., Haraguchi T., Futaki S., Mol.Ther., 2012, 20(5), 984—993 |
| [14] | Wimley W. C., ACS Chem.Biol., 2010, 5(10), 905—917 |
| [15] | Sun Y. N., Li R. F., Cheng Y. B., Lu Z. F., J. Jiangsu Univ. Sci. Technol., 2016, 30(2), 177—182 |
| (孙亚楠, 李瑞芳, 程艳波, 陆志方. 江苏科技大学学报(自然科学版), 2016, 30(2), 177—182) | |
| [16] | Lu Y. L., Zhang H. R., Li R. F., Lu Y. B., Zhai J. Y., Yan X. H., J. Pathog.Biol., 2014, 9(1), 37—47 |
| (卢亚丽, 张慧茹, 李瑞芳, 卢研博, 翟晋豫, 闫晓慧. 中国病原生物学杂志,2014, 9(1), 37—47) | |
| [17] | Li R. F., Lu Z. F., Sun Y. N., Chen S. H., Yi Y. J., Zhang H. R., Yang S. Y., Yu G. H., Huang L., Li C. N.,Interdiscip.Sci., 2016, 8(3), 319—326 |
| [18] | Tyagi A., Kapoor P., Kumar R., Chaudhary K., Gautam A., Raghava G. P. S., Sci.Rep., 2013, 3(10), 2984-1—8 |
| [19] | Mojsoska B., Zuckermann R. N., Jenssen H., Antimicrob. Agents Chemother., 2015, 59(7), 4112—4120 |
| [20] | Saviello M. R., Malfi S., Campiglia P., Cavalli A., Grieco P., Novellino E., Carotenuto A., Biochem.,2010, 49(7), 1477—1485 |
| [21] | Mangoni M. L., Carotenuto A., Auriemma L., Saviello M. R., Campiglia P., Gomez-Monterrey I., Malfi S., Marcellini L., Barra D., Novellino E., Grieco P., J. Med.Chem., 2015, 54(5), 1298—1307 |
| [22] | Dong W., Dong Z., Mao X., Sun Y., Li F., Shang D., Acta Biomater., 2016, 37, 59—68 |
| [23] | Gopal R., Seo C. H., Song P. I., Park Y., Amino Acids,2013, 44(2), 645—660 |
| [1] | YANG Dan, LIU Xu, DAI Yihu, ZHU Yan, YANG Yanhui. Research Progress in Electrocatalytic CO2 Reduction Reaction over Gold Clusters [J]. Chem. J. Chinese Universities, 2022, 43(7): 20220198. |
| [2] | ZHAO Yingzhe, ZHANG Jianling. Applications of Metal-organic Framework-based Material in Carbon Dioxide Photocatalytic Conversion [J]. Chem. J. Chinese Universities, 2022, 43(7): 20220223. |
| [3] | HOU Hua, WANG Baoshan. Group Additivity Theoretical Model for the Prediction of Dielectric Strengths of the Alternative Gases to SF6 [J]. Chem. J. Chinese Universities, 2021, 42(12): 3709. |
| [4] | YE Xiaodong, QI Guodong, XU Jun, DENG Feng. Glucose Oxidation on Au-supported SBA-15 Molecular Sieve † [J]. Chem. J. Chinese Universities, 2020, 41(5): 960. |
| [5] | LIU Li, MA Yangyang, WANG Kuan, JIA Yunjing, LI Wan, ZHU Huajie. Anti-tumor and Antimicrobial Activities of β-Carbolines† [J]. Chem. J. Chinese Universities, 2018, 39(4): 674. |
| [6] | ZHAO Wencai, HAN Lili, PENG Yingjun, WANG Xiaojing, LIU Shengyu, LI Pengfei, HUANG Yibing, CHEN Yuxin. Effect of Basic Amino Acids on the Biological Activity of Helical Antimicrobial Peptide† [J]. Chem. J. Chinese Universities, 2018, 39(4): 681. |
| [7] | YU Min, HUANG Jingjing, MA Min, FU Ruiyan, YAN Yan, ZHANG Fusheng, YIN Junfeng, XIE Ningning. Zinc Chelating Activity and Quantitative Structure-activity Relationship of Tripeptides† [J]. Chem. J. Chinese Universities, 2018, 39(2): 234. |
| [8] | HOU Hua, YU Xiaojuan, ZHOU Wenjun, LUO Yunbai, WANG Baoshan. Theoretical Investigations on the Structure-activity Relationship to the Dielectric Strength of the Insulation Gases† [J]. Chem. J. Chinese Universities, 2018, 39(11): 2477. |
| [9] | WANG Lei, ZHENG Guojun, JI Qi, CHEN Bo, GONG Longlong, GAO Congmin, DU Zhenjian, ZHANG Xingmin. Synthesis and Biological Activity of Novel PI3K/mTOR Inhibitors† [J]. Chem. J. Chinese Universities, 2017, 38(9): 1590. |
| [10] | LIU Yuming, TIAN Lijun, HU Dong, NIE Jianbing. yntheses and Anti-cholinesterase Activity of 4-N-Phenylaminoquinoline Derivatives † [J]. Chem. J. Chinese Universities, 2017, 38(3): 392. |
| [11] | LIU Benguo, LIU Jiangwei, LI Jiaqi, GENG Sheng, MO Haizhen, LIANG Guizhao. 3D-QSAR and Interaction Mechanism of Flavonoids s P-glycoprotein Inhibitors† [J]. Chem. J. Chinese Universities, 2017, 38(1): 41. |
| [12] | GUO Liang, CAO Rihui, FAN Wenxi, GAN Ziyun, MA Qin. Design, Synthesis and in vitro Antitumor Activities of Novel Bivalent β-Carbolines† [J]. Chem. J. Chinese Universities, 2016, 37(6): 1093. |
| [13] | ZHANG Futing, ZHOU Jinlong, HUANG Hairui, JI Zhenyan, YANG Liangjiong, FU Min, FANG Bijun, ZHANG Hongwen, JIANG Yan, YU Qiang, ZHOU Wanlin. Preparation of Polyimide Nanofiber with Antimicrobial Activity by Electrospinning† [J]. Chem. J. Chinese Universities, 2016, 37(4): 781. |
| [14] | ZHANG Jie, ZHOU Changjian, XIE Jianwei, DAI Bin. Synthesis and Antitumor Activities of Rhein-Valine Adducts† [J]. Chem. J. Chinese Universities, 2016, 37(12): 2159. |
| [15] | ZHANG Jing, MU Boshuai, WU Meng, BIAN Yanqing, LI Yuan. Synthesis, Antifungal Activity and Structure-activity Relationship of -Fluorophenyl-2,3-dihydro-1,5-benzothiazepines Derivatives† [J]. Chem. J. Chinese Universities, 2015, 36(4): 687. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||