

Chem. J. Chinese Universities ›› 2019, Vol. 40 ›› Issue (1): 138.doi: 10.7503/cjcu20180445
• Physical Chemistry • Previous Articles Next Articles
ZHU Jingxuan1, YU Zhengfei1, LIU Ye1, ZHAN Dongling2, HAN Jiarui1, TIAN Xiaopian1, HAN Weiwei1,*( )
)
Received:2018-06-19
Online:2019-01-10
Published:2018-12-19
Contact:
HAN Weiwei
E-mail:weiweihan@jlu.edu.cn
Supported by:CLC Number:
TrendMD:
ZHU Jingxuan,YU Zhengfei,LIU Ye,ZHAN Dongling,HAN Jiarui,TIAN Xiaopian,HAN Weiwei. Exploration of Increasing the Non-specificity Substrates Activity for the Phosphotriesterase-like Lactonase Using Molecular Dynamics Simulations†[J]. Chem. J. Chinese Universities, 2019, 40(1): 138.
| Amino acid | Interaction | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Any | Backbone | Sidechain | Polar | Hydrophobcic | H-Bond acceptor | H-Bond donor | Aromatic | Charged | |
| His24 | 0.72 | 0 | 0.72 | 0.28 | 0.44 | 0 | 0 | 0 | 0 |
| His22 | 0.26 | 0 | 0.26 | 0.11 | 0.15 | 0 | 0 | 0 | 0 |
| Tyr97 | 0.85 | 0 | 0.85 | 0.45 | 0.40 | 0 | 0.13 | 0.72 | 0 |
| His170 | 0.81 | 0 | 0.81 | 0.34 | 0.47 | 0 | 0 | 0 | 0 |
| Arg223 | 0.88 | 0 | 0.88 | 0.71 | 0.16 | 0 | 0.24 | 0 | 0.14 |
| Asp256 | 0.85 | 0.68 | 0.85 | 0.67 | 0.18 | 0 | 0 | 0 | 0 |
| Trp278 | 0.84 | 0 | 0.84 | 0.08 | 0.75 | 0 | 0 | 0.79 | 0 |
| Trp263 | 0.89 | 0 | 0.89 | 0 | 0.88 | 0 | 0 | 0.88 | 0 |
| Leu226 | 0.68 | 0 | 0.68 | 0 | 0.68 | 0 | 0 | 0 | 0 |
| Cys258 | 0.47 | 0 | 0.47 | 0.25 | 0.24 | 0 | 0 | 0 | 0 |
| Ile261 | 0.53 | 0.05 | 0.53 | 0.05 | 0.51 | 0 | 0 | 0 | 0 |
| Thr265 | 0.31 | 0.21 | 0.31 | 0.20 | 0.13 | 0 | 0 | 0 | 0 |
| Ala266 | 0.41 | 0.31 | 0.41 | 0 | 0.40 | 0 | 0 | 0 | 0 |
| Leu274 | 0.24 | 0.13 | 0.24 | 0 | 0.21 | 0 | 0 | 0 | 0 |
| Ala275 | 0.22 | 0.12 | 0.22 | 0 | 0.20 | 0 | 0 | 0 | 0 |
| Leu228 | 0.38 | 0 | 0.38 | 0 | 0.38 | 0 | 0 | 0 | 0 |
| Phe229 | 0.48 | 0 | 0.48 | 0 | 0.48 | 0 | 0 | 0.47 | 0 |
| Tyr99 | 0.31 | 0 | 0.31 | 0.06 | 0.25 | 0 | 0 | 0.27 | 0 |
| Val27 | 0.49 | 0 | 0.49 | 0 | 0.49 | 0 | 0 | 0 | 0 |
| Leu72 | 0.48 | 0 | 0.48 | 0 | 0.47 | 0 | 0 | 0 | 0 |
Table 1 Structural interaction fingerprints results(residues number according to PDB ID: 2VC7)
| Amino acid | Interaction | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Any | Backbone | Sidechain | Polar | Hydrophobcic | H-Bond acceptor | H-Bond donor | Aromatic | Charged | |
| His24 | 0.72 | 0 | 0.72 | 0.28 | 0.44 | 0 | 0 | 0 | 0 |
| His22 | 0.26 | 0 | 0.26 | 0.11 | 0.15 | 0 | 0 | 0 | 0 |
| Tyr97 | 0.85 | 0 | 0.85 | 0.45 | 0.40 | 0 | 0.13 | 0.72 | 0 |
| His170 | 0.81 | 0 | 0.81 | 0.34 | 0.47 | 0 | 0 | 0 | 0 |
| Arg223 | 0.88 | 0 | 0.88 | 0.71 | 0.16 | 0 | 0.24 | 0 | 0.14 |
| Asp256 | 0.85 | 0.68 | 0.85 | 0.67 | 0.18 | 0 | 0 | 0 | 0 |
| Trp278 | 0.84 | 0 | 0.84 | 0.08 | 0.75 | 0 | 0 | 0.79 | 0 |
| Trp263 | 0.89 | 0 | 0.89 | 0 | 0.88 | 0 | 0 | 0.88 | 0 |
| Leu226 | 0.68 | 0 | 0.68 | 0 | 0.68 | 0 | 0 | 0 | 0 |
| Cys258 | 0.47 | 0 | 0.47 | 0.25 | 0.24 | 0 | 0 | 0 | 0 |
| Ile261 | 0.53 | 0.05 | 0.53 | 0.05 | 0.51 | 0 | 0 | 0 | 0 |
| Thr265 | 0.31 | 0.21 | 0.31 | 0.20 | 0.13 | 0 | 0 | 0 | 0 |
| Ala266 | 0.41 | 0.31 | 0.41 | 0 | 0.40 | 0 | 0 | 0 | 0 |
| Leu274 | 0.24 | 0.13 | 0.24 | 0 | 0.21 | 0 | 0 | 0 | 0 |
| Ala275 | 0.22 | 0.12 | 0.22 | 0 | 0.20 | 0 | 0 | 0 | 0 |
| Leu228 | 0.38 | 0 | 0.38 | 0 | 0.38 | 0 | 0 | 0 | 0 |
| Phe229 | 0.48 | 0 | 0.48 | 0 | 0.48 | 0 | 0 | 0.47 | 0 |
| Tyr99 | 0.31 | 0 | 0.31 | 0.06 | 0.25 | 0 | 0 | 0.27 | 0 |
| Val27 | 0.49 | 0 | 0.49 | 0 | 0.49 | 0 | 0 | 0 | 0 |
| Leu72 | 0.48 | 0 | 0.48 | 0 | 0.47 | 0 | 0 | 0 | 0 |
Fig.3 Conformational changes in WT, W263F and W263T SsoPox during 100 ns MD(A1, A2) RMSD; (B1, B2) RMSF; (C1, C2) SASA; (A1, B1, C1) paraoxon; (A2, B2, C2) lactone.
| System | WT-lactone | W263T-lactone | WT-paraoxon | W263F-paraoxon |
|---|---|---|---|---|
| ΔEele/(kJ·mol-1) | -47.72 | -90.50 | -15.64 | -93.63 |
| ΔEvdw/(kJ·mol-1) | -126.76 | -129.39 | -72.68 | -101.62 |
| ΔGGB/(kJ·mol-1) | 83.97 | 102.50 | 42.49 | 109.23 |
| ΔGSA/(kJ·mol-1) | -17.73 | -18.69 | -10.79 | -16.18 |
| Δ | 36.26 | 12.00 | 26.85 | 15.64 |
| Δ | -144.49 | -148.08 | -83.47 | -117.81 |
| Δ | -108.23 | -136.08 | -56.62 | -102.17 |
| -TΔS/(kJ·mol-1) | 80.80 | 72.52 | 77.58 | 92.13 |
| Δ | -27.43 | -63.57 | 20.95 | -10.04 |
| (2.36±0.38)×103 | (3.49±1.13)×105 | (5.19±0.95)×102 | (1.21±0.26)×104 |
Table 2 Calculate the binding free energy of a protein-ligand complex using MM-GB/SA methods
| System | WT-lactone | W263T-lactone | WT-paraoxon | W263F-paraoxon |
|---|---|---|---|---|
| ΔEele/(kJ·mol-1) | -47.72 | -90.50 | -15.64 | -93.63 |
| ΔEvdw/(kJ·mol-1) | -126.76 | -129.39 | -72.68 | -101.62 |
| ΔGGB/(kJ·mol-1) | 83.97 | 102.50 | 42.49 | 109.23 |
| ΔGSA/(kJ·mol-1) | -17.73 | -18.69 | -10.79 | -16.18 |
| Δ | 36.26 | 12.00 | 26.85 | 15.64 |
| Δ | -144.49 | -148.08 | -83.47 | -117.81 |
| Δ | -108.23 | -136.08 | -56.62 | -102.17 |
| -TΔS/(kJ·mol-1) | 80.80 | 72.52 | 77.58 | 92.13 |
| Δ | -27.43 | -63.57 | 20.95 | -10.04 |
| (2.36±0.38)×103 | (3.49±1.13)×105 | (5.19±0.95)×102 | (1.21±0.26)×104 |
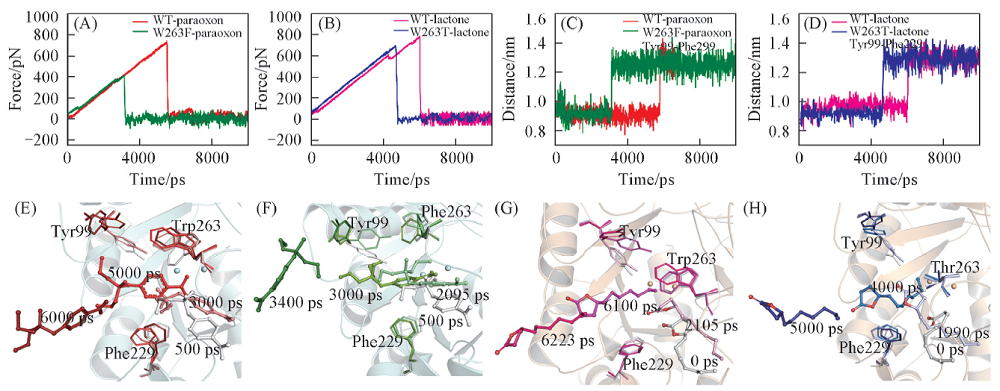
Fig.9 SMD simulation of typical force profiles of paraoxon(A) and lactone(B) pulled out of the binding pocket along the unbinding pathway, the distance between the Cz of Tyr99 and Phe229 in WT-paraoxon and W263F-paraoxon(C), WT-lactone and W263T-lactone(D) and conformational changes of WT-paraoxon(E), W263F-paraoxon(F), WT-lactone(G), W263T-lactone(H) during 10 ns SMD
| [1] | Benning M. M., Shim H., Raushel F. M., Holden H. M., Biochemistry,2001, 40(9), 2712—2722 |
| [2] | Afriat L., Roodveldt C., Manco G., Tawfik D. S., Biochemistry,2006, 45(46), 13677—13686 |
| [3] | Omburo G. A., Kuo J. M., Mullins L. S., Raushel F. M., J. Biol. Chem.,1992, 267(19), 13278—13283 |
| [4] | Gomes D. E. B., Lins R. D., Pascutti P. G., Lei C., Soares T. A., J. Phys. Chem. B,2011, 115(51), 15389—15398 |
| [5] | Galloway W. R. J. D., Hodgkinson J. T., Bowden S. D., Welch M., Spring D. R., Chem. Rev.,2011, 111(1), 28—67 |
| [6] | Fuqua C., Greenberg E. P., Curr. Opin. Microbiol.,1998, 1(2), 183—189 |
| [7] | Elias M., Dupuy J., Merone L., Mandrich L., Porzio E., Moniot S., Rochu D., Lecomte C., Rossi M., Masson P., Manco G., Chabriere E., J. Mol. Biol.,2008, 379(5), 1017—1028 |
| [8] | Brock T. D., Brock K. M., Belly R. T., Weiss R. L., Arch. Mikrobiol.,1972, 84(1), 54—68 |
| [9] | Deng Q. G., Song B., J. Sci. Teachers' College University,2003, 23(2), 26—28 |
| (邓启刚, 宋波. 高师理科学刊, 2003, 23#(2), 26—28) | |
| [10] | Lewis V. E., Donarski W. J., Wild J. R., Raushel F. M., Biochemistry,1988, 27(5), 1591—1597 |
| [11] | Hiblot J., Gotthard G., Elias M., Chabriere E., PloS One,2013, 8(9), e75272 |
| [12] | Frisch M., Trucks G., Schlegel H., Scuseria G., Robb M., Cheeseman J., Scalmani G., Barone V., Mennucci B., Petersson G., Nakatsuji H., Caricato M., Li X., Hratchian H., Izmaylov A., Bloino J., Zheng G., Sonnenberg J., Hada M., Ehara M., Toyota K., Fukuda R., Hasegawa J., Ishida M., Nakajima T., Honda Y., Kitao O., Nakai H., Vreven T., Montgomery J.Jr., Peralta J., Ogliaro F., Bearpark M., Heyd J., Brothers E., Kudin K., Staroverov V., Kobayashi R., Normand J., Raghavachari K., Rendell A., Burant J., Iyengar S., Tomasi J., Cossi M., Rega N., Millam J., Klene M., Knox J., Cross J., Bakken V., Adamo C., Jaramillo J., Gomperts R., Stratmann R., Yazyev O., Austin A., Cammi R., Pomelli C., Ochterski J., Martin R., Morokuma K., Zakrzewski V., Voth G., Salvador P., Dannenberg J., Dapprich S., Daniels A., Farkas Ö., Foresman J., Ortiz J., Cioslowski J., Fox D., Gaussian 09, Revision A.01, Gaussian Inc.,Wallingford CT, 2009 |
| [13] | Lu T., Chen F., J. Comput. Chem.,2012, 33(5), 580—592 |
| [14] | Wu G., Robertson D. H., Brooks C. L., Vieth M., J. Comput. Chem., 2003, 24(13), 1549—1562 |
| [15] | Norgan A. P., Coffman P. K., Kocher J. P. A., Katzmann D. J., Sosa C. P., J. Cheminform.,2011, 3(1), 12 |
| [16] | Hou X., Du J., Zhang J., Du L., Fang H., Li M., J. Chem. Inf. Model,2013, 53(1), 188—200 |
| [17] | Trott O., Olson A. J., J. Comput.Chem.,2010, 31(2), 455—461 |
| [18] | Stigliani J. L., Bernardes-Génisson V., Bernadou J., Pratviel G., Org. Biomol. Chem.,2012, 10(31), 6341—6349 |
| [19] | Wickstrom L., Okur A., Simmerling C., Biophys. J.,2009, 97(3), 853—856 |
| [20] | Wang J., Wolf R. M., Caldwell J. W., Kollman P. A., Case D. A., J. Comput. Chem., 2004, 25(9), 1157—1174 |
| [21] | Van der Spoel D., Lindahl E., Hess B., Groenhof G., Mark A. E., Berendsen H. J. C., J. Comput. Chem., 2005, 26(16), 1701—1718 |
| [22] | Hess B., Kutzner C., Dan Der Spoel D., Lindahl E., J. Chem. Theory Comput., 2008, 4(3), 435—447 |
| [23] | Mark P., Nilsson L., J. Phys. Chem. A,2001, 105(43), 9954—9960 |
| [24] | Sousa da Silva A. W., Vranken W. F., BMC Res. Notes,2012, 5(1), 367 |
| [25] | Darden T., York D., Pedersen L., J. Chem. Phys., 1993, 98(12), 10089—10092 |
| [26] | Hou T., Wang J., Li Y., Wang W., J. Chem. Inf. Model,2011, 51(1), 69—82 |
| [27] | Hou T., Wang J., Li Y., Wang W., J. Comput. Chem.,2015, 32(5), 866—877 |
| [28] | Sun H., Li Y., Shen M., Tian S., Xu L., Pan P., Guan Y., Hou T., Phys. Chem. Chem. Phys., 2014, 16(40), 22035—22045 |
| [29] | Sun H., Li Y., Tian S., Xu L., Hou T., Phys. Chem. Chem. Phys., 2014, 16(31), 16719—16729 |
| [30] | Case D. A., Cheatham T. E., Darden T., Gohlke H., Luo R., Merz K. M., Onufriev A., Simmerling C., Wang B., Woods R. J., J. Comput. Chem., 2005, 26(16), 1668—1688 |
| [31] | Izrailev S., Stepaniants S., Isralewitz B., Kosztin D., Lu H., Molnar F., Wriggers W., Schulten K., Steered Molecular Dynamics, Springer, Berlin,1999, 4(12), 39—65 |
| [32] | Isralewitz B., Gao M., Schulten K., Curr. Opin. Struct. Biol.,2001, 11(2), 224—230 |
| [33] | Hiblot J., Gotthard G., Chabriere E., Elias M., Sci. Rep.,2012, 2(11), 779 |
| [1] | GAO Zhiwei, LI Junwei, SHI Sai, FU Qiang, JIA Junru, AN Hailong. Analysis of Gating Characteristics of TRPM8 Channel Based on Molecular Dynamics [J]. Chem. J. Chinese Universities, 2022, 43(6): 20220080. |
| [2] | HU Bo, ZHU Haochen. Dielectric Constant of Confined Water in a Bilayer Graphene Oxide Nanosystem [J]. Chem. J. Chinese Universities, 2022, 43(2): 20210614. |
| [3] | ZHANG Mi, TIAN Yafeng, GAO Keli, HOU Hua, WANG Baoshan. Molecular Dynamics Simulation of the Physicochemical Properties of Trifluoromethanesulfonyl Fluoride Dielectrics [J]. Chem. J. Chinese Universities, 2022, 43(11): 20220424. |
| [4] | LEI Xiaotong, JIN Yiqing, MENG Xuanyu. Prediction of the Binding Site of PIP2 in the TREK-1 Channel Based on Molecular Modeling [J]. Chem. J. Chinese Universities, 2021, 42(8): 2550. |
| [5] | LI Congcong, LIU Minghao, HAN Jiarui, ZHU Jingxuan, HAN Weiwei, LI Wannan. Theoretical Study of the Catalytic Activity of VmoLac Non-specific Substrates Based on Molecular Dynamics Simulations [J]. Chem. J. Chinese Universities, 2021, 42(8): 2518. |
| [6] | ZENG Yonghui, YAN Tianying. Vibrational Density of States Analysis of Proton Hydration Structure [J]. Chem. J. Chinese Universities, 2021, 42(6): 1855. |
| [7] | LIU Aiqing, XU Wensheng, XU Xiaolei, CHEN Jizhong, AN Lijia. Molecular Dynamics Simulation of Polymer/rod Nanocomposite [J]. Chem. J. Chinese Universities, 2021, 42(3): 875. |
| [8] | QI Renrui, LI Minghao, CHANG Hao, FU Xueqi, GAO Bo, HAN Weiwei, HAN Lu, LI Wannan. Theoretical Study on the Unbinding Pathway of Xanthine Oxidase Inhibitors Based on Steered Molecular Dynamics Simulation [J]. Chem. J. Chinese Universities, 2021, 42(3): 758. |
| [9] | QU Siying, XU Qin. Different Roles of Some Key Residues in the S4 Pocket of Coagulation Factor Xa for Rivaroxaban Binding † [J]. Chem. J. Chinese Universities, 2019, 40(9): 1918. |
| [10] | MA Yucong, FAN Baomin, WANG Manman, YANG Biao, HAO Hua, SUN Hui, ZHANG Huijuan. Two-step Preparation of Trazodone and Its Corrosion Inhibition Mechanism for Carbon Steel [J]. Chem. J. Chinese Universities, 2019, 40(8): 1706. |
| [11] | ZHANG Zhang,WANG Dong,WANG Xiaolei,XU Yan. Regulation of Ester Synthesis Activity of Rhizopus chinensis Lipase† [J]. Chem. J. Chinese Universities, 2019, 40(4): 747. |
| [12] | MA Lan,RONG Jingjing,ZHU Youliang,HUANG Yineng,SUN Zhaoyan. Simulation on the Dynamic Process of Formation of Particle Cluster by Generalized Exponential Model† [J]. Chem. J. Chinese Universities, 2019, 40(1): 195. |
| [13] | WU Hongmei,LI Huiting,LI Yongcheng,WANG Hongqing,WANG Meng. Using Group Contribution Method and Molecular Dynamics to Predict the Glass Transition Temperature of Poly(p-phenylene isophthalamide)† [J]. Chem. J. Chinese Universities, 2019, 40(1): 180. |
| [14] | LIU Yanfang, YANG Hua, ZHANG Hui. Molecular Dynamics Simulation on the Orientation of Alkane Mixture on Graphene† [J]. Chem. J. Chinese Universities, 2018, 39(8): 1729. |
| [15] | SHI Penghui,BIAN Liujiao. Mechanism Study on the Interaction Between Cefoxitin and Metal β-Lactamase BcⅡ Based on Spectroscopic Methods and Computional Simulations† [J]. Chem. J. Chinese Universities, 2018, 39(5): 971. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||