

Chem. J. Chinese Universities ›› 2018, Vol. 39 ›› Issue (9): 1893.doi: 10.7503/cjcu20180262
• Analytical Chemistry • Previous Articles Next Articles
QIAO Junqin, LIANG Chao, CAO Zhaoming, LIAN Hongzhen*( )
)
Received:2018-04-04
Online:2018-09-07
Published:2018-06-26
Contact:
LIAN Hongzhen
E-mail:hzlian@nju.edu.cn
Supported by:CLC Number:
TrendMD:
QIAO Junqin,LIANG Chao,CAO Zhaoming,LIAN Hongzhen. Retention Behavior of Oligonucleotides under System Containing Mixed Ion-pair Reagents by IP-RPLC†[J]. Chem. J. Chinese Universities, 2018, 39(9): 1893.
| Oligonucleotide | Sequence(5'-3') | Percentage(%) | ΔG*/(J·mol-1) | |||
|---|---|---|---|---|---|---|
| G | C | A | T | |||
| Oligo20-1 | CTTAGTGAAGAGCTCAGTTA | 25 | 15 | 30 | 30 | -3.97×104 |
| Oligo20-2 | CTTAGTGAAGAGTCTCTAAG | 25 | 15 | 30 | 30 | -2.55×104 |
| Oligo20-3 | GACAGGAAAGACATTCTGGC | 30 | 20 | 35 | 15 | -1.48×104 |
| Oligo20-4 | GACAGGAAAGACATTCCGGT | 30 | 20 | 35 | 15 | -4.08×104 |
| Oligo32-1 | GTCGTTATCATCAGAGTAGCCCAGGAAGCTTC | 25 | 25 | 25 | 25 | -5.60×104 |
| Oligo32-2 | GCGTACAGTATAGCCCAGTCTTGAGTGCCATA | 25 | 25 | 25 | 25 | -1.53×104 |
| Oligo32-3 | GGTATGGTTTACGAGTATTGCCTGAAGCGAGG | 37.5 | 12.5 | 21.9 | 28.1 | -1.95×104 |
| Oligo32-4 | CCTCGCTTCAGGCAATACTCGTAAACCATACC | 12.5 | 37.5 | 28.1 | 21.9 | -1.95×104 |
Table 1 Hetero-oligonucleotides used in the experiment
| Oligonucleotide | Sequence(5'-3') | Percentage(%) | ΔG*/(J·mol-1) | |||
|---|---|---|---|---|---|---|
| G | C | A | T | |||
| Oligo20-1 | CTTAGTGAAGAGCTCAGTTA | 25 | 15 | 30 | 30 | -3.97×104 |
| Oligo20-2 | CTTAGTGAAGAGTCTCTAAG | 25 | 15 | 30 | 30 | -2.55×104 |
| Oligo20-3 | GACAGGAAAGACATTCTGGC | 30 | 20 | 35 | 15 | -1.48×104 |
| Oligo20-4 | GACAGGAAAGACATTCCGGT | 30 | 20 | 35 | 15 | -4.08×104 |
| Oligo32-1 | GTCGTTATCATCAGAGTAGCCCAGGAAGCTTC | 25 | 25 | 25 | 25 | -5.60×104 |
| Oligo32-2 | GCGTACAGTATAGCCCAGTCTTGAGTGCCATA | 25 | 25 | 25 | 25 | -1.53×104 |
| Oligo32-3 | GGTATGGTTTACGAGTATTGCCTGAAGCGAGG | 37.5 | 12.5 | 21.9 | 28.1 | -1.95×104 |
| Oligo32-4 | CCTCGCTTCAGGCAATACTCGTAAACCATACC | 12.5 | 37.5 | 28.1 | 21.9 | -1.95×104 |
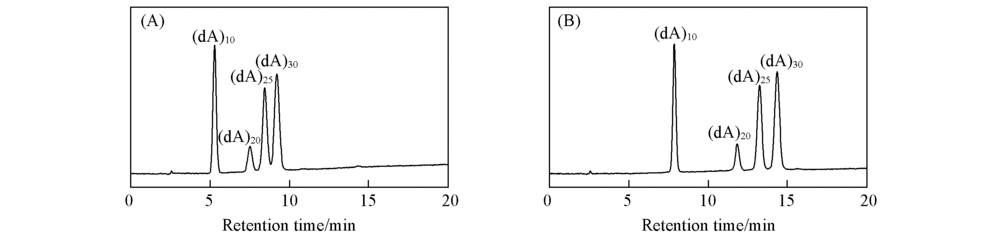
Fig.1 Chromatograms of (dA)n under mobile phases containing either 10 mmol/L TEA/10 mmol/L PA(A)or 20 mmol/L TEA(B)Purospher© STAR RP-18 endcapped column(50 mm×4.6 mm i.d., 5 μm); column temperature: 30 ℃; mobile phase A(10 mmol/L TEA/10 mmol/L PA-20 mmol/LAA-5%CH3CN, pH=7.0) or (20 mmol/L TEAA-5%CH3CN, pH=7.0), mobile phase B(10 mmol/L TEA/10 mmol/L PA-20 mmol/L AA-25%CH3CN, pH=7.0) or (20 mmol/L TEAA-25%CH3CN, pH=7.0); gradient elution: 0—40 min, 15%B—45%B; 40—43 min, 45%B—15%B; 43—58 min, 15%B—15%B; flow rate: 1.0 mL/min; detection wavelength: 260 nm.
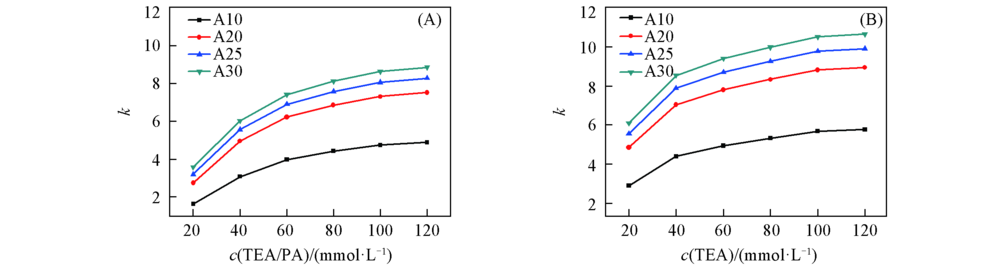
Fig.2 Retention factor k of (dA)n under mobile phases containing different concentrations of ion-pair reagent(A) TEA/PA; (B) TEA. The concentrations of ion-pair reagent in mobile phase were 20, 40, 60, 80, 100 and 120 mmol/L, respectively. Other chromatographic conditions were the same as in Fig.1.
| TEA/PA | TEA | |||||
|---|---|---|---|---|---|---|
| A10/A20 | A20/A25 | A25/A30 | A10/A20 | A20/A25 | A25/A30 | |
| 20 | 5.02 | 1.76 | 1.34 | 9.06 | 2.73 | 1.97 |
| 40 | 9.20 | 2.66 | 1.89 | 12.47 | 3.47 | 2.55 |
| 60 | 10.90 | 2.96 | 2.12 | 13.36 | 3.74 | 2.77 |
| 80 | 11.58 | 3.17 | 2.29 | 13.98 | 3.92 | 2.89 |
| 100 | 12.37 | 3.33 | 2.37 | 14.76 | 4.03 | 2.96 |
| 120 | 12.55 | 3.34 | 2.39 | 14.42 | 3.99 | 2.94 |
Table 2 Resolution(Rs) of adjacent peaks in (dA)n under different concentrations of ion-pair reagent
| TEA/PA | TEA | |||||
|---|---|---|---|---|---|---|
| A10/A20 | A20/A25 | A25/A30 | A10/A20 | A20/A25 | A25/A30 | |
| 20 | 5.02 | 1.76 | 1.34 | 9.06 | 2.73 | 1.97 |
| 40 | 9.20 | 2.66 | 1.89 | 12.47 | 3.47 | 2.55 |
| 60 | 10.90 | 2.96 | 2.12 | 13.36 | 3.74 | 2.77 |
| 80 | 11.58 | 3.17 | 2.29 | 13.98 | 3.92 | 2.89 |
| 100 | 12.37 | 3.33 | 2.37 | 14.76 | 4.03 | 2.96 |
| 120 | 12.55 | 3.34 | 2.39 | 14.42 | 3.99 | 2.94 |
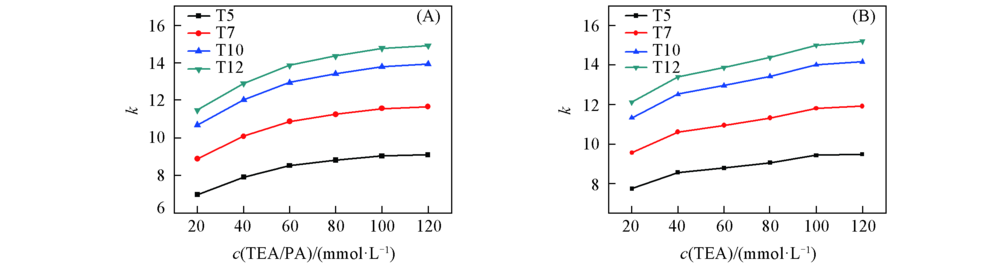
Fig.3 Retention factor k of (dT)n under mobile phases containing different concentrations of ion-pair reagent (A) TEA/PA; (B) TEA. The chromatographic conditions are the same as those in Fig.2.
| cp/(mmol·L-1) | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| T5/T7 | T7/T10 | T10/T12 | T5/T7 | T7/T10 | T10/T12 | |
| 20 | 10.28 | 9.24 | 3.78 | 9.79 | 9.26 | 4.08 |
| 40 | 11.79 | 10.42 | 4.44 | 10.96 | 10.20 | 4.53 |
| 60 | 12.50 | 10.96 | 4.76 | 11.58 | 10.48 | 4.85 |
| 80 | 12.92 | 11.32 | 4.85 | 11.83 | 10.98 | 5.06 |
| 100 | 13.32 | 11.63 | 5.06 | 12.24 | 11.40 | 5.07 |
| 120 | 13.49 | 11.78 | 5.31 | 12.54 | 11.60 | 5.26 |
Table 3 Resolution(Rs) of adjacent peaks in (dT)n under different concentrations of ion-pair reagent
| cp/(mmol·L-1) | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| T5/T7 | T7/T10 | T10/T12 | T5/T7 | T7/T10 | T10/T12 | |
| 20 | 10.28 | 9.24 | 3.78 | 9.79 | 9.26 | 4.08 |
| 40 | 11.79 | 10.42 | 4.44 | 10.96 | 10.20 | 4.53 |
| 60 | 12.50 | 10.96 | 4.76 | 11.58 | 10.48 | 4.85 |
| 80 | 12.92 | 11.32 | 4.85 | 11.83 | 10.98 | 5.06 |
| 100 | 13.32 | 11.63 | 5.06 | 12.24 | 11.40 | 5.07 |
| 120 | 13.49 | 11.78 | 5.31 | 12.54 | 11.60 | 5.26 |
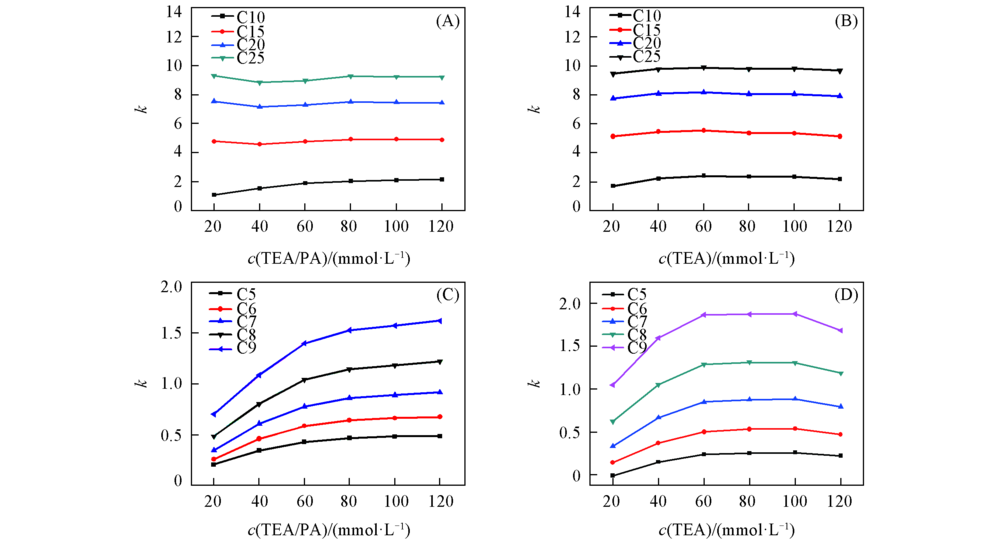
Fig.4 Retention factor(k) of (dC)n under mobile phases containing different concentrations of ion-pair reagent (A), (C) TEA/PA; (B), (D) TEA. The chromatographic conditions are the same as those in Fig.2.
| System | cp/(mmol·L-1) | C5/C6 | C6/C7 | C7/C8 | C8/C9 | C10/C15 | C15/C20 | C20/C25 |
|---|---|---|---|---|---|---|---|---|
| TEA/PA | 20 | 0.43 | 0.76 | 1.15 | 1.53 | 12.90 | 5.24 | 1.88 |
| 40 | 1.21 | 1.33 | 1.38 | 1.68 | 12.03 | 5.44 | 2.09 | |
| 60 | 1.54 | 1.58 | 1.80 | 2.02 | 11.46 | 5.37 | 2.01 | |
| 80 | 1.67 | 1.81 | 1.91 | 2.12 | 11.37 | 5.47 | 1.93 | |
| 100 | 1.72 | 1.86 | 1.94 | 2.10 | 11.43 | 5.09 | 2.06 | |
| 120 | 1.79 | 1.95 | 2.02 | 2.16 | 10.94 | 5.45 | 2.15 | |
| TEA | 20 | 1.16 | 1.25 | 1.67 | 2.00 | 13.01 | 5.57 | 2.00 |
| 40 | 1.69 | 1.86 | 1.98 | 2.35 | 13.46 | 5.53 | 1.94 | |
| 60 | 1.82 | 1.98 | 2.12 | 2.46 | 13.05 | 5.51 | 1.97 | |
| 80 | 1.91 | 1.98 | 2.16 | 2.41 | 12.48 | 5.62 | 1.99 | |
| 100 | 1.86 | 1.93 | 2.07 | 2.29 | 12.36 | 5.31 | 2.01 | |
| 120 | 1.73 | 1.90 | 1.96 | 2.15 | 12.04 | 5.71 | 2.04 |
Table 4 Resolution(Rs) of adjacent peaks in (dC)n under different concentrations of ion-pair reagent
| System | cp/(mmol·L-1) | C5/C6 | C6/C7 | C7/C8 | C8/C9 | C10/C15 | C15/C20 | C20/C25 |
|---|---|---|---|---|---|---|---|---|
| TEA/PA | 20 | 0.43 | 0.76 | 1.15 | 1.53 | 12.90 | 5.24 | 1.88 |
| 40 | 1.21 | 1.33 | 1.38 | 1.68 | 12.03 | 5.44 | 2.09 | |
| 60 | 1.54 | 1.58 | 1.80 | 2.02 | 11.46 | 5.37 | 2.01 | |
| 80 | 1.67 | 1.81 | 1.91 | 2.12 | 11.37 | 5.47 | 1.93 | |
| 100 | 1.72 | 1.86 | 1.94 | 2.10 | 11.43 | 5.09 | 2.06 | |
| 120 | 1.79 | 1.95 | 2.02 | 2.16 | 10.94 | 5.45 | 2.15 | |
| TEA | 20 | 1.16 | 1.25 | 1.67 | 2.00 | 13.01 | 5.57 | 2.00 |
| 40 | 1.69 | 1.86 | 1.98 | 2.35 | 13.46 | 5.53 | 1.94 | |
| 60 | 1.82 | 1.98 | 2.12 | 2.46 | 13.05 | 5.51 | 1.97 | |
| 80 | 1.91 | 1.98 | 2.16 | 2.41 | 12.48 | 5.62 | 1.99 | |
| 100 | 1.86 | 1.93 | 2.07 | 2.29 | 12.36 | 5.31 | 2.01 | |
| 120 | 1.73 | 1.90 | 1.96 | 2.15 | 12.04 | 5.71 | 2.04 |
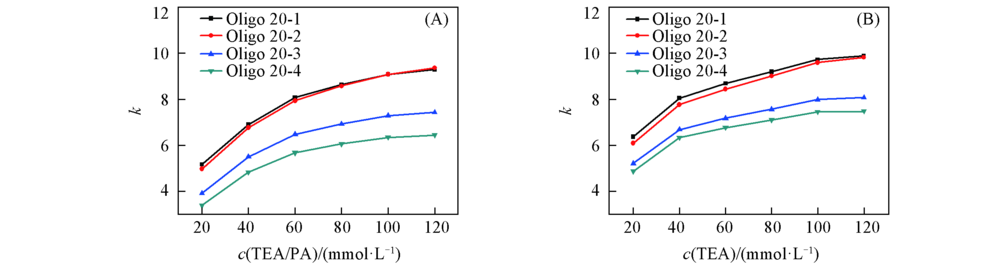
Fig.5 Retention factor(k) of 20mer hetero-oligonucleotides under mobile phases containing different concentrations of ion-pair reagent (A) TEA/PA; (B) TEA. The chromatographic conditions are the same as those in Fig.2.
| cp/(mmol·L-1) | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| 20-4/20-3 | 20-3/20-2 | 20-2/20-1 | 20-4/20-3 | 20-3/20-2 | 20-2/20-1 | |
| 20 | 2.18 | 4.41 | 0.87 | 1.50 | 3.90 | 1.29 |
| 40 | 3.34 | 6.29 | 0.61 | 1.61 | 5.00 | 1.32 |
| 60 | 3.73 | 6.88 | 0.61 | 1.88 | 5.88 | 1.15 |
| 80 | 4.01 | 7.59 | 0.28 | 2.12 | 6.65 | 0.84 |
| 100 | 4.39 | 8.52a | 0.02b | 2.42 | 7.29 | 0.60 |
| 120 | 4.53 | 8.74a | 0.34b | 2.65 | 7.84 | 0.28 |
Table 5 Resolution(Rs) of adjacent peaks in 20mer hetero-oligonucleotides under different concentrations of ion-pair reagent
| cp/(mmol·L-1) | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| 20-4/20-3 | 20-3/20-2 | 20-2/20-1 | 20-4/20-3 | 20-3/20-2 | 20-2/20-1 | |
| 20 | 2.18 | 4.41 | 0.87 | 1.50 | 3.90 | 1.29 |
| 40 | 3.34 | 6.29 | 0.61 | 1.61 | 5.00 | 1.32 |
| 60 | 3.73 | 6.88 | 0.61 | 1.88 | 5.88 | 1.15 |
| 80 | 4.01 | 7.59 | 0.28 | 2.12 | 6.65 | 0.84 |
| 100 | 4.39 | 8.52a | 0.02b | 2.42 | 7.29 | 0.60 |
| 120 | 4.53 | 8.74a | 0.34b | 2.65 | 7.84 | 0.28 |
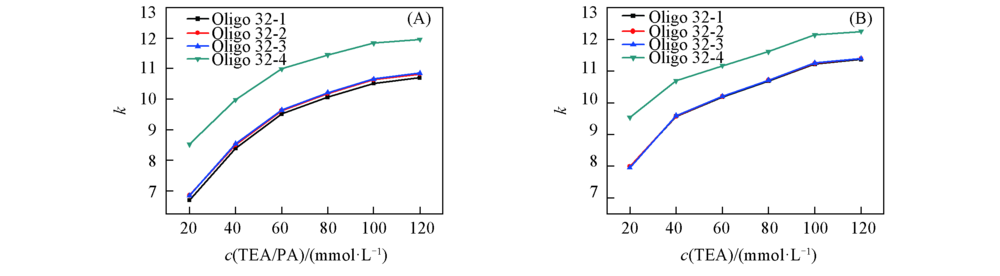
Fig.6 Retention factor(k) of 32mer hetero-oligonucleotides under mobile phases containing different concentrations of ion-pair reagent (A) TEA/PA; (B) TEA. The chromatographic conditions a the same as those in Fig.2.
| cp/(mmol·L-1) | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| 32-1/32-2 | 32-2/32-3 | 32-3/32-4 | 32-1/32-2 | 32-2/32-3 | 32-3/32-4 | |
| 20 | 0.49 | 0.02 | 4.92 | 0.10a | 0.02b | 5.07c |
| 40 | 0.48 | 0.13 | 5.24 | 0.01 | 0.09 | 4.22 |
| 60 | 0.46 | 0.07 | 5.16 | 0.06 | 0.04 | 3.84 |
| 80 | 0.48 | 0.10 | 4.81 | 0.11 | 0.01 | 3.65 |
| 100 | 0.50 | 0.10 | 4.65 | 0.10 | 0.05 | 3.61 |
| 120 | 0.50 | 0.16 | 4.48 | 0.09 | 0.03 | 3.42 |
Table 6 Resolution of adjacent peaks in 32mer hetero-oligonucleotides under different concentrations of ion-pair reagent
| cp/(mmol·L-1) | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| 32-1/32-2 | 32-2/32-3 | 32-3/32-4 | 32-1/32-2 | 32-2/32-3 | 32-3/32-4 | |
| 20 | 0.49 | 0.02 | 4.92 | 0.10a | 0.02b | 5.07c |
| 40 | 0.48 | 0.13 | 5.24 | 0.01 | 0.09 | 4.22 |
| 60 | 0.46 | 0.07 | 5.16 | 0.06 | 0.04 | 3.84 |
| 80 | 0.48 | 0.10 | 4.81 | 0.11 | 0.01 | 3.65 |
| 100 | 0.50 | 0.10 | 4.65 | 0.10 | 0.05 | 3.61 |
| 120 | 0.50 | 0.16 | 4.48 | 0.09 | 0.03 | 3.42 |
| Oligonucleotide | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| Intercept | Slope | R2 | Intercept | Slope | R2 | |
| A10 | 1.6110 | 0.0310 | 0.841 | 2.9782 | 0.0266 | 0.816 |
| A20 | 2.7694 | 0.0452 | 0.830 | 5.0068 | 0.0376 | 0.806 |
| A25 | 3.2356 | 0.0480 | 0.830 | 5.7163 | 0.0399 | 0.804 |
| A30 | 3.6109 | 0.0498 | 0.831 | 6.2791 | 0.0417 | 0.802 |
| T5 | 6.9595 | 0.0204 | 0.834 | 7.6807 | 0.0165 | 0.894 |
| T7 | 8.8489 | 0.0268 | 0.849 | 9.4400 | 0.0225 | 0.911 |
| T10 | 10.6040 | 0.0314 | 0.866 | 11.1440 | 0.0274 | 0.923 |
| T12 | 11.3850 | 0.0333 | 0.868 | 11.9050 | 0.0297 | 0.929 |
| C5 | 0.2155 | 0.0027 | 0.772 | 0.4297 | 0.0017 | 0.496 |
| C6 | 0.2714 | 0.0039 | 0.783 | 0.5674 | 0.0025 | 0.488 |
| C7 | 0.3539 | 0.0054 | 0.808 | 0.7494 | 0.0034 | 0.459 |
| C8 | 0.4859 | 0.0071 | 0.815 | 1.0169 | 0.0041 | 0.397 |
| C9 | 0.6992 | 0.0088 | 0.816 | 1.4032 | 0.0046 | 0.316 |
| Oligo20-1 | 5.0833 | 0.0396 | 0.860 | 6.3444 | 0.0330 | 0.867 |
| Oligo20-2 | 4.8244 | 0.0422 | 0.880 | 5.9811 | 0.0353 | 0.893 |
| Oligo20-3 | 3.9152 | 0.0334 | 0.836 | 5.2573 | 0.0266 | 0.836 |
| Oligo20-4 | 3.4328 | 0.0289 | 0.814 | 4.9940 | 0.0239 | 0.788 |
| Oligo32-1 | 6.6123 | 0.0386 | 0.860 | 7.9326 | 0.0319 | 0.876 |
| Oligo32-2 | 6.7599 | 0.0383 | 0.864 | 7.9301 | 0.0322 | 0.879 |
| Oligo32-3 | 6.7601 | 0.0386 | 0.863 | 7.9182 | 0.0324 | 0.870 |
| Oligo32-4 | 8.4744 | 0.0331 | 0.852 | 9.3990 | 0.02617 | 0.915 |
Table 7 Linear correlation between k and cp
| Oligonucleotide | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| Intercept | Slope | R2 | Intercept | Slope | R2 | |
| A10 | 1.6110 | 0.0310 | 0.841 | 2.9782 | 0.0266 | 0.816 |
| A20 | 2.7694 | 0.0452 | 0.830 | 5.0068 | 0.0376 | 0.806 |
| A25 | 3.2356 | 0.0480 | 0.830 | 5.7163 | 0.0399 | 0.804 |
| A30 | 3.6109 | 0.0498 | 0.831 | 6.2791 | 0.0417 | 0.802 |
| T5 | 6.9595 | 0.0204 | 0.834 | 7.6807 | 0.0165 | 0.894 |
| T7 | 8.8489 | 0.0268 | 0.849 | 9.4400 | 0.0225 | 0.911 |
| T10 | 10.6040 | 0.0314 | 0.866 | 11.1440 | 0.0274 | 0.923 |
| T12 | 11.3850 | 0.0333 | 0.868 | 11.9050 | 0.0297 | 0.929 |
| C5 | 0.2155 | 0.0027 | 0.772 | 0.4297 | 0.0017 | 0.496 |
| C6 | 0.2714 | 0.0039 | 0.783 | 0.5674 | 0.0025 | 0.488 |
| C7 | 0.3539 | 0.0054 | 0.808 | 0.7494 | 0.0034 | 0.459 |
| C8 | 0.4859 | 0.0071 | 0.815 | 1.0169 | 0.0041 | 0.397 |
| C9 | 0.6992 | 0.0088 | 0.816 | 1.4032 | 0.0046 | 0.316 |
| Oligo20-1 | 5.0833 | 0.0396 | 0.860 | 6.3444 | 0.0330 | 0.867 |
| Oligo20-2 | 4.8244 | 0.0422 | 0.880 | 5.9811 | 0.0353 | 0.893 |
| Oligo20-3 | 3.9152 | 0.0334 | 0.836 | 5.2573 | 0.0266 | 0.836 |
| Oligo20-4 | 3.4328 | 0.0289 | 0.814 | 4.9940 | 0.0239 | 0.788 |
| Oligo32-1 | 6.6123 | 0.0386 | 0.860 | 7.9326 | 0.0319 | 0.876 |
| Oligo32-2 | 6.7599 | 0.0383 | 0.864 | 7.9301 | 0.0322 | 0.879 |
| Oligo32-3 | 6.7601 | 0.0386 | 0.863 | 7.9182 | 0.0324 | 0.870 |
| Oligo32-4 | 8.4744 | 0.0331 | 0.852 | 9.3990 | 0.02617 | 0.915 |
| Oligonucleotide | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| A1 | B1 | R2 | A1 | B1 | R2 | |
| A10 | 0.1012 | 10.026 | 0.984 | 0.1339 | 4.1449 | 0.991 |
| A20 | 0.0750 | 5.6674 | 0.985 | 0.0907 | 2.2607 | 0.993 |
| A25 | 0.0737 | 4.6767 | 0.988 | 0.0834 | 1.9013 | 0.994 |
| A30 | 0.0726 | 4.0594 | 0.990 | 0.0784 | 1.6810 | 0.994 |
| T5 | 0.1035 | 0.8199 | 0.987 | 0.1024 | 0.5507 | 0.957 |
| T7 | 0.0791 | 0.7760 | 0.999 | 0.0815 | 0.4808 | 0.955 |
| T10 | 0.0677 | 0.5527 | 0.987 | 0.0686 | 0.4111 | 0.948 |
| T12 | 0.0636 | 0.4860 | 0.980 | 0.0640 | 0.3872 | 0.946 |
| C5 | 1.3103 | 69.715 | 0.984 | 1.4721 | 20.595 | 0.942 |
| C6 | 0.8572 | 58.660 | 0.981 | 1.0707 | 16.734 | 0.937 |
| C7 | 0.6338 | 44.098 | 0.986 | 0.7867 | 13.573 | 0.930 |
| C8 | 0.5083 | 30.667 | 0.990 | 0.6100 | 9.5254 | 0.915 |
| C9 | 0.4201 | 19.968 | 0.993 | 0.4803 | 6.1408 | 0.877 |
| C10 | 0.3614 | 11.452 | 0.994 | 0.3818 | 3.6390 | 0.762 |
| Oligo20-1 | 0.0901 | 2.0868 | 0.998 | 0.0825 | 1.3597 | 0.995 |
| Oligo20-2 | 0.0881 | 2.2844 | 0.998 | 0.0821 | 1.2991 | 0.993 |
| Oligo20-3 | 0.1079 | 2.9454 | 0.999 | 0.0881 | 2.2844 | 0.998 |
| Oligo20-4 | 0.1228 | 3.4268 | 0.997 | 0.1079 | 2.9454 | 0.999 |
| Oligo32-1 | 0.0825 | 1.3597 | 0.995 | 0.0818 | 0.8813 | 0.989 |
| Oligo32-2 | 0.0821 | 1.2991 | 0.993 | 0.0816 | 0.8847 | 0.989 |
| Oligo32-3 | 0.0817 | 1.3088 | 0.994 | 0.0813 | 0.8989 | 0.991 |
| Oligo32-4 | 0.0773 | 0.8182 | 0.988 | 0.0786 | 0.5428 | 0.960 |
Table 8 Correlations of k-1-cp fitted according to Eq.(9)
| Oligonucleotide | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|
| A1 | B1 | R2 | A1 | B1 | R2 | |
| A10 | 0.1012 | 10.026 | 0.984 | 0.1339 | 4.1449 | 0.991 |
| A20 | 0.0750 | 5.6674 | 0.985 | 0.0907 | 2.2607 | 0.993 |
| A25 | 0.0737 | 4.6767 | 0.988 | 0.0834 | 1.9013 | 0.994 |
| A30 | 0.0726 | 4.0594 | 0.990 | 0.0784 | 1.6810 | 0.994 |
| T5 | 0.1035 | 0.8199 | 0.987 | 0.1024 | 0.5507 | 0.957 |
| T7 | 0.0791 | 0.7760 | 0.999 | 0.0815 | 0.4808 | 0.955 |
| T10 | 0.0677 | 0.5527 | 0.987 | 0.0686 | 0.4111 | 0.948 |
| T12 | 0.0636 | 0.4860 | 0.980 | 0.0640 | 0.3872 | 0.946 |
| C5 | 1.3103 | 69.715 | 0.984 | 1.4721 | 20.595 | 0.942 |
| C6 | 0.8572 | 58.660 | 0.981 | 1.0707 | 16.734 | 0.937 |
| C7 | 0.6338 | 44.098 | 0.986 | 0.7867 | 13.573 | 0.930 |
| C8 | 0.5083 | 30.667 | 0.990 | 0.6100 | 9.5254 | 0.915 |
| C9 | 0.4201 | 19.968 | 0.993 | 0.4803 | 6.1408 | 0.877 |
| C10 | 0.3614 | 11.452 | 0.994 | 0.3818 | 3.6390 | 0.762 |
| Oligo20-1 | 0.0901 | 2.0868 | 0.998 | 0.0825 | 1.3597 | 0.995 |
| Oligo20-2 | 0.0881 | 2.2844 | 0.998 | 0.0821 | 1.2991 | 0.993 |
| Oligo20-3 | 0.1079 | 2.9454 | 0.999 | 0.0881 | 2.2844 | 0.998 |
| Oligo20-4 | 0.1228 | 3.4268 | 0.997 | 0.1079 | 2.9454 | 0.999 |
| Oligo32-1 | 0.0825 | 1.3597 | 0.995 | 0.0818 | 0.8813 | 0.989 |
| Oligo32-2 | 0.0821 | 1.2991 | 0.993 | 0.0816 | 0.8847 | 0.989 |
| Oligo32-3 | 0.0817 | 1.3088 | 0.994 | 0.0813 | 0.8989 | 0.991 |
| Oligo32-4 | 0.0773 | 0.8182 | 0.988 | 0.0786 | 0.5428 | 0.960 |
| Oligonucleotide | TEA/PA | TEA | Oligonucleotide | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|---|---|---|
| K4 | K5 | K4 | K5 | K4 | K5 | K4 | K5 | ||
| A10 | 0.010 | 9.881β | 0.0323 | 7.468β | C8 | 0.017 | 1.967β | 0.0640 | 1.639β |
| A20 | 0.013 | 13.333β | 0.0401 | 11.025β | C9 | 0.021 | 2.380β | 0.0782 | 2.082β |
| A25 | 0.016 | 13.568β | 0.0439 | 11.990β | C10 | 0.032 | 2.767β | 0.1049 | 2.619β |
| A30 | 0.018 | 13.774β | 0.0466 | 12.755β | Oligo20-1 | 0.043 | 11.099β | 0.0607 | 12.121β |
| T5 | 0.126 | 9.662β | 0.1859 | 9.766β | Oligo20-2 | 0.039 | 11.351β | 0.0632 | 12.180β |
| T7 | 0.102 | 12.642β | 0.1695 | 12.270β | Oligo20-3 | 0.037 | 9.268β | 0.0386 | 11.351β |
| T10 | 0.122 | 14.771β | 0.1669 | 14.577β | Oligo20-4 | 0.036 | 8.143β | 0.0366 | 9.268β |
| T12 | 0.131 | 15.723β | 0.1653 | 15.625β | Oligo32-1 | 0.061 | 12.121β | 0.0928 | 12.225β |
| C5 | 0.019 | 0.763β | 0.0715 | 0.679β | Oligo32-2 | 0.063 | 12.180β | 0.0922 | 12.255β |
| C6 | 0.015 | 1.167β | 0.0640 | 0.934β | Oligo32-3 | 0.062 | 12.240β | 0.0904 | 12.300β |
| C7 | 0.014 | 1.578β | 0.0580 | 1.271β | Oligo32-4 | 0.094 | 12.937β | 0.1448 | 12.723β |
Table 9 Quilibrium constants of K4 and K5 for oligonucleotides
| Oligonucleotide | TEA/PA | TEA | Oligonucleotide | TEA/PA | TEA | ||||
|---|---|---|---|---|---|---|---|---|---|
| K4 | K5 | K4 | K5 | K4 | K5 | K4 | K5 | ||
| A10 | 0.010 | 9.881β | 0.0323 | 7.468β | C8 | 0.017 | 1.967β | 0.0640 | 1.639β |
| A20 | 0.013 | 13.333β | 0.0401 | 11.025β | C9 | 0.021 | 2.380β | 0.0782 | 2.082β |
| A25 | 0.016 | 13.568β | 0.0439 | 11.990β | C10 | 0.032 | 2.767β | 0.1049 | 2.619β |
| A30 | 0.018 | 13.774β | 0.0466 | 12.755β | Oligo20-1 | 0.043 | 11.099β | 0.0607 | 12.121β |
| T5 | 0.126 | 9.662β | 0.1859 | 9.766β | Oligo20-2 | 0.039 | 11.351β | 0.0632 | 12.180β |
| T7 | 0.102 | 12.642β | 0.1695 | 12.270β | Oligo20-3 | 0.037 | 9.268β | 0.0386 | 11.351β |
| T10 | 0.122 | 14.771β | 0.1669 | 14.577β | Oligo20-4 | 0.036 | 8.143β | 0.0366 | 9.268β |
| T12 | 0.131 | 15.723β | 0.1653 | 15.625β | Oligo32-1 | 0.061 | 12.121β | 0.0928 | 12.225β |
| C5 | 0.019 | 0.763β | 0.0715 | 0.679β | Oligo32-2 | 0.063 | 12.180β | 0.0922 | 12.255β |
| C6 | 0.015 | 1.167β | 0.0640 | 0.934β | Oligo32-3 | 0.062 | 12.240β | 0.0904 | 12.300β |
| C7 | 0.014 | 1.578β | 0.0580 | 1.271β | Oligo32-4 | 0.094 | 12.937β | 0.1448 | 12.723β |
| [1] | Huber C. G., Oefner P. K., Bonn G. K., J. Chromatogr. A, 1992, 599(1/2), 113—118 |
| [2] | Apfell A., Chakel J. A., Fischer S., Lichtenwalter K., Hancock W. S., Anal. Chem., 1997, 69(7), 1320—1325 |
| [3] | Gilar M., Foutain K. J., Budman Y., Neue U. D., Yardley K. R., Rainville P. D., Russell R. J., Gebler J.C., J. Chromatogr. A, 2002, 958(1/2), 167—182 |
| [4] | Huber C. G., Krajete A., Anal. Chem., 1999, 71(17), 3730—3739 |
| [5] | Huber C. G., Krajete A., J. Chromatogr. A, 2000, 870(1/2), 413—424 |
| [6] | Buncek M., Backovska V., Holasova A., Radilova H., Safarova M., Kunc F., Haluza R., Anal. Biochem., 2006, 348(2), 300—306 |
| [7] | Bothner B., Chatman K., Sarkisian M., Siuzdak G., Bioorg. Med. Chem. Lett., 1995, 5(23), 2863—2868 |
| [8] | Apffel A., Chakel J., Fischer S., Lichtenwalter K., Hancock W., J. Chromatogr. A, 1997, 777(1), 3—21 |
| [9] | Kenski D. M., Cooper A. J., Li J. J., Willingham A. T., Haringsma H. J., Young T. A., Kuklin N. A., Jones J. J., Cancilla M. T., McMasters D. R., Nucleic Acids Res., 2010, 38(2), 660—671 |
| [10] | Dai G., Wei X., Liu Z., Liu S., Marcucci G., Chan K. K., J. Chromatogr. B, 2005, 825(2), 201—213 |
| [11] | Deng P., Chen X., Zhang G., Zhong D., J. Pharm. Biomed. Anal., 2010, 52(4), 571—579 |
| [12] | Zhang G., Lin J., Srinivasan K., Kavetskaia O., Duncan J., Anal. Chem., 2007, 79(9), 3416—3424 |
| [13] | Lin Z., Li W., Dai G., J. Pharm. Biomed. Anal., 2007, 44(2), 330—341 |
| [14] | Oberacher H., Parson W., Muhlmann R., Huber C., Anal. Chem., 2001, 73(21), 5109—5115 |
| [15] | Oberacher H., Oefner P., Parson W., Huber C., Angew. Chem. Int. Ed., 2001, 40(20), 3828—3830 |
| [16] | Oberacher H., Niederstatter H., Pitterl F., Parson W., Anal. Chem., 2006, 78(22), 7816—7827 |
| [17] | Erb R., Leithner K., Bernkop-Schnürch A., Oberacher H., AAPS J., 2012, 14(4), 728—736 |
| [18] | McCarthy S. M., Gilar M., Gebler J., Anal. Biochem., 2009, 390(2), 181—188 |
| [19] | Gong L. Z., McCullagh J. S. O., Rapid Commun. Mass Spectrom., 2014, 28(4), 339—350 |
| [20] | Miyaguchi H., JoVE-J. Vis. Exp., 2016, (115), e54402 |
| [21] | Beverly M., Hartsough K., Machemer L., Rapid Commun. Mass Spectrom.,2005, 19(12), 1675—1682 |
| [22] | Beverly M., Hartsough K., Machemer L., Pavco P., Lockridge J., J. Chromatogr. B, 2006, 835(1/2), 62—70 |
| [23] | Levin D. S., Shepperd B. T., Gruenloh C. J., J. Chromatogr. B, 2011, 879(19), 1587—1595 |
| [24] | Yuan N., Han S. Y., Yang J., Qiao J. Q., Liu Y., Lian H. Z., Curr. Anal. Chem.,2012, 8(4), 550—556 |
| [25] | Qiao J. Q., Liang C., Wei L. C., Cao Z. M., Lian H. Z., J. Sep. Sci., 2016, 39(23), 4502—4511 |
| [26] | Tomlinson E., Riley C. M., Jefferies T. M., J. Chromatogr. A, 1979, 173(1), 89—100 |
| [27] | Wang Y. P., Shen G. Q., Zhu M. H., Acta Chim. Sinica, 1993, 51, 392—396 |
| (王延平, 沈国钦, 朱明华. 化学学报, 1993, 51, 392—396) | |
| [28] | Han S. Y., Qiao J. Q., Zhang Y. Y., Yang L. L., Lian H. Z., Ge X., Chen H. Y., Chemosphere, 2011, 83(2), 131—136 |
| [29] | Huber C. G., Oefner P. J., Bonn G. K., Anal. Biochem., 1993, 212(2), 351—358 |
| [30] | Lee D. P., Kindsvater J. H., Anal. Chem., 1980, 52(14), 2425—2428 |
| [31] | Huber C. G., Oefner P. J., Bonn G. K., J. Chromatogr. A, 1992, 599(1/2), 113—118 |
| [32] | Knox J. H., Hartwick R. A., J Chromatogr., 1981, 204(1), 3—21 |
| [33] | Horvath C., Melander W., Monlar I., Anal. Chem., 1977, 49(1), 142—154 |
| [1] | CAO Shujie, LI Hongjun, GUAN Wenli, REN Mengtian, ZHOU Chuanzheng. Progress on the Stereocontrolled Synthesis of Phosphorothioate Oligonucleotides [J]. Chem. J. Chinese Universities, 2022, 43(Album-4): 20220304. |
| [2] | YU Qiongwei, ZHENG Feng, FANG Kaimin, FENG Yuqi. Preparation of Zirconia Deposited Silica Stationary Phase and Its Application to Hydrophilic-interaction Liquid Chromatography† [J]. Chem. J. Chinese Universities, 2019, 40(9): 1857. |
| [3] | LIU Zhongcheng, ZHAO Lijian, LIU Lipeng, ZHANG Yanfen, WANG Nannan, CHEN Yao, WANG Xianghuan. Novel Method Based on Colloidal Gold for Detection of Oligonucleotide Aptamer with Protein Interacting† [J]. Chem. J. Chinese Universities, 2015, 36(4): 638. |
| [4] | CHEN Yan, LIU Gui-Feng, LIU Xia, ZHANG Gui-Zhen, WANG Zhen-Xin. Detecting Type 2 Diabetes-related Single Nucleotide Polymorphisms by Microarray [J]. Chem. J. Chinese Universities, 2013, 34(5): 1078. |
| [5] | GAO Yun-Hua, LI Hai-Feng, LI Jian-Xin, LI Peng,YE Jian, ZHAO Xing-Chun. Quantitative Analysis of Fluorescent Dye\|labeled DNA by High Resolution Inductively Coupled Plasma Mass Spectrometry [J]. Chem. J. Chinese Universities, 2010, 31(12): 2360. |
| [6] | LIANG Yuan-Jun, HE Jun-Lin, XU Liang, ZHANG Di, LIU Ke-Liang. Chemical Synthesis and Stability of Oligonucleotides Modified by Alkyl- or Alkoxy- on the Phosphonates [J]. Chem. J. Chinese Universities, 2007, 28(3): 467. |
| [7] | PAN Qin, XU Li-Jian, WANG Zhi-Fei, LU Zu-Hong*, HE Nong-Yue*. Electrochemical Detection of Sequence Specific DNA Binding Protein with Gold Nanoparticle-catalized Ag Enhancement [J]. Chem. J. Chinese Universities, 2007, 28(12): 2290. |
| [8] | SHEN Jia-Yao, HOU Peng, JI Mei-Ju, LI Song, HE Nong-Yue . A Methylation-specific Oligonuceotide Microarray for Quantitative Analysis of p16 Gene [J]. Chem. J. Chinese Universities, 2005, 26(3): 449. |
| [9] | DONG Li-Qin, ZHOU Jian-Zhang, WU Ling-Ling, DONG Ping, LIN Zhong-Hua . Surface Enhanced Raman Scattering(SERS) Studies on Adsorption Orientation on Self-assembled DNA [J]. Chem. J. Chinese Universities, 2002, 23(12): 2303. |
| [10] | LEI Rong, XING Yu-Jie, YANG Jun-Jiao, ZUO Yu-Min. Evaluation of Polymer Encapsulated-TiO2 Stationary Phase and Comparison to Related Phases Using Linear Solvation Energy Relationships [J]. Chem. J. Chinese Universities, 2001, 22(S1): 45. |
| [11] | YU Hong-Wu, ZHANG Liang-Ren, MA Ling-Tai, ZHANG Li-He. Synthesis and Enzymatic Stability of oligonucleotide Incorporated with Isonucleoside [J]. Chem. J. Chinese Universities, 1997, 18(7): 1103. |
| [12] | LUAN Lian-Jun, ZENG Su, LIU Zhi-Qiang, FU Xu-Chun. Studies on the RP-HPLC Retention Behavior Relationship Between the Structure of Hydroxy Compounds and Their Glucuronides [J]. Chem. J. Chinese Universities, 1997, 18(1): 42. |
| [13] | Zuo Yumin, Xie Mengxia, Yang Zhiqiang, Qi Mei, Yang Xuejin . Studies on the Retention Mechanism for Ester-Containing Porous Polymer Beads as Stationary Phase [J]. Chem. J. Chinese Universities, 1988, 9(5): 493. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||