

Chem. J. Chinese Universities ›› 2015, Vol. 36 ›› Issue (6): 1156.doi: 10.7503/cjcu20150050
• Physical Chemistry • Previous Articles Next Articles
ZHANG Fada1,2, LIU Yi2,3,*( ), XU Jingcheng2, LI Shengjuan2, WANG Xiunan1,2, SUN Yue2, ZHAO Xinluo3
), XU Jingcheng2, LI Shengjuan2, WANG Xiunan1,2, SUN Yue2, ZHAO Xinluo3
Received:2015-01-16
Online:2015-06-10
Published:2015-05-22
Contact:
LIU Yi
E-mail:yiliu@shu.edu.cn
Supported by:CLC Number:
TrendMD:
ZHANG Fada, LIU Yi, XU Jingcheng, LI Shengjuan, WANG Xiunan, SUN Yue, ZHAO Xinluo. Molecular Dynamics Study on Binding Strength and Conformation of Dendrimer-based Drug Delivery Systems†[J]. Chem. J. Chinese Universities, 2015, 36(6): 1156.
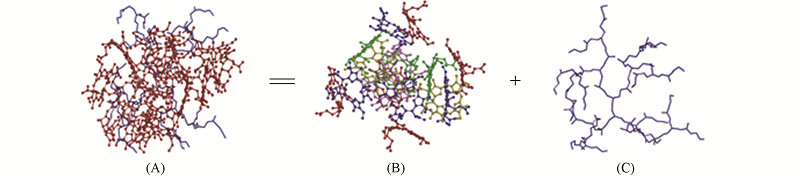
Fig.1 Atomic structures of CE616@G2(A) snapshot and the structures of the CE616(B) and G2(C) separated from the CE616@G2, respectivelyThe various colors in (B) denote the different drug molecules. All the H atoms are omitted for clarity.
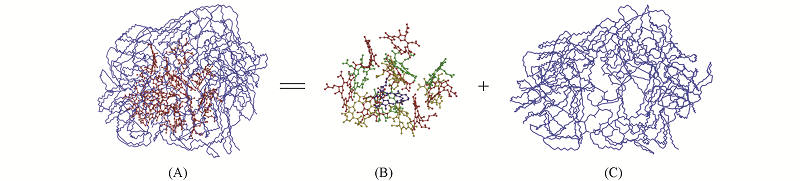
Fig.2 Atomic structures of CE616@G2-PEG(A) snapshot and the structures of the CE616(B) and G2-PEG(C) separated from the CE616@G2-PEG, respectivelyThe various colors in (B) denote the different drug molecules. All the H atoms are omitted for clarity.
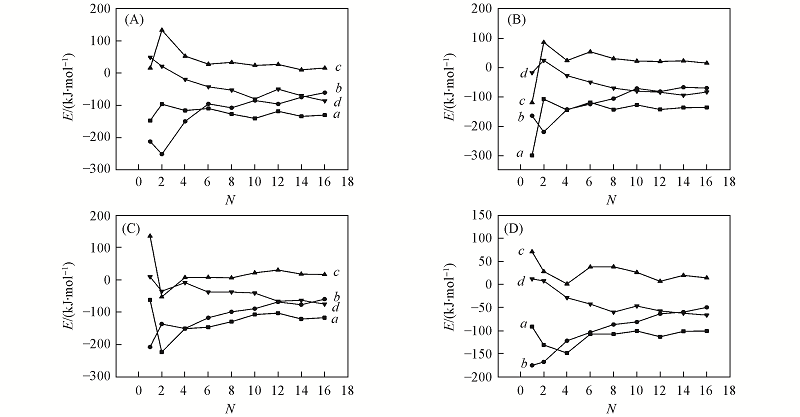
Fig.4 Instantaneous binding energies(Eb-PD, a), interaction energies(Ei-PD, b), deformation energies(Ed-P, c), and deformation-and-interaction energies(Edi-D, d) of the DN@G2 complexes with CE6(A), DOX(B), MTX(C) and SN38(D) as the functions of the number of drug molecules N
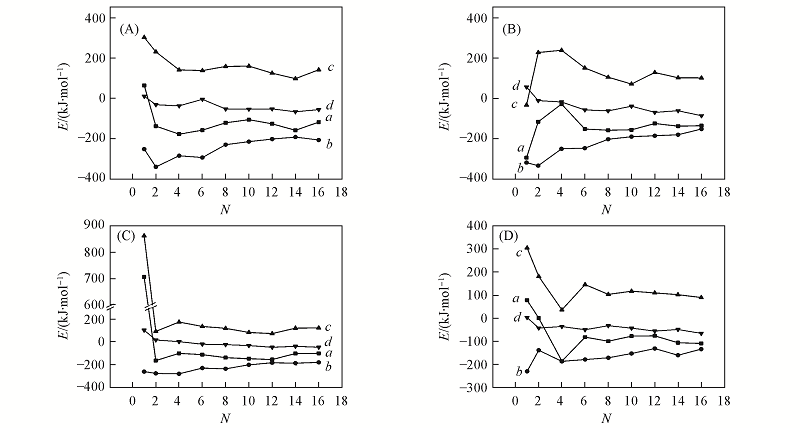
Fig.5 Instantaneous binding energies(Eb-PD, a), interaction energies(Ei-PD, b), deformation energies(Ed-P, c), and deformation-and-interaction(Edi-D, d) energies of the DN@G2-PEG complexes with CE6(A), DOX(B), MTX(C) and SN38(D) as the functions of the number of drug molecules N
| Drug | D1 | D1(D16@G2) | G2 | G2(D16@G2) | D16@G2 | D1(D16@ G2-PEG) | G2-PEG | G2-PEG(D16@ G2-PEG) | D16@G2-PEG |
|---|---|---|---|---|---|---|---|---|---|
| None | 0.896 | 1.965 | |||||||
| CE6 | 0.465 | 0.469±0.05 | 1.294(44.4%) | 1.351(50.8%) | 0.467±0.05 | 2.411(22.7%) | 2.242(14.1%) | ||
| DOX | 0.479 | 0.484±0.07 | 1.314(46.7%) | 1.227(67.0%) | 0.484±0.07 | 2.475(26.0%) | 2.318(18.0%) | ||
| MTX | 0.525 | 0.541±0.42 | 1.228(37.1%) | 1.211(35.2%) | 0.542±0.45 | 2.315(17.8%) | 2.167(10.3%) | ||
| SN38 | 0.434 | 0.434±0.01 | 1.14(27.2%) | 1.189(32.7%) | 0.435±0.01 | 2.36(20.1%) | 2.254(14.7%) |
Table 1 Rg(nm) of drug molecules(D1), dendrimers(G2 or G2-PEG) and drug-dendrimer complexes(DN@G2 or DN@G2-PEG)
| Drug | D1 | D1(D16@G2) | G2 | G2(D16@G2) | D16@G2 | D1(D16@ G2-PEG) | G2-PEG | G2-PEG(D16@ G2-PEG) | D16@G2-PEG |
|---|---|---|---|---|---|---|---|---|---|
| None | 0.896 | 1.965 | |||||||
| CE6 | 0.465 | 0.469±0.05 | 1.294(44.4%) | 1.351(50.8%) | 0.467±0.05 | 2.411(22.7%) | 2.242(14.1%) | ||
| DOX | 0.479 | 0.484±0.07 | 1.314(46.7%) | 1.227(67.0%) | 0.484±0.07 | 2.475(26.0%) | 2.318(18.0%) | ||
| MTX | 0.525 | 0.541±0.42 | 1.228(37.1%) | 1.211(35.2%) | 0.542±0.45 | 2.315(17.8%) | 2.167(10.3%) | ||
| SN38 | 0.434 | 0.434±0.01 | 1.14(27.2%) | 1.189(32.7%) | 0.435±0.01 | 2.36(20.1%) | 2.254(14.7%) |
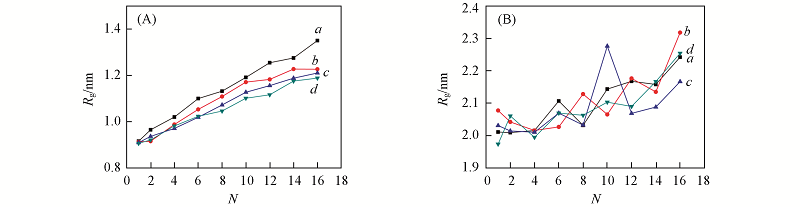
Fig.6 Radius of gyration(Rg) as functions of number of drug molecules in DN@G2(A) and DN@G2-PEG(B) with CE6(a), DOX(b), MTX(c) and SN38(d), respectively
| DN@Gn | Eb(H2O)/ (kJ·mol-1) | Eb(no H2O)/ (kJ·mol-1) | DN@Gn-PEG | Eb(H2O)/ (kJ·mol-1) | Eb(no H2O)/ (kJ·mol-1) |
|---|---|---|---|---|---|
| CE68@G0 | -218.572 | -98.157 | CE68@G0-PEG | 154.808 | -138.951 |
| DOX8@G0 | -201.418 | -111.587 | DOX8@G0-PEG | 118.407 | -146.900 |
| MTX8@G0 | -205.476 | -82.299 | MTX8@G0-PEG | 166.900 | -145.478 |
| SN388@G0 | -137.277 | -84.224 | SN388@G0-PEG | 212.464 | -125.813 |
| CE610@G1 | -254.136 | -109.412 | CE612@G1-PEG | 224.555 | -144.766 |
| DOX10@G1 | -148.699 | -125.478 | DOX12@G1-PEG | 364.719 | -129.160 |
| MTX10@G1 | -210.079 | -101.546 | MTX12@G1-PEG | 301.834 | -112.089 |
| SN3810@G1 | -90.709 | -96.650 | SN3812@G1-PEG | 346.017 | -142.047 |
| CE616@G2 | -213.761 | -122.089 | CE616@G2-PEG | 378.610 | -141.126 |
| DOX16@G2 | -195.351 | -137.612 | DOX16@G2-PEG | 533.251 | -119.077 |
| MTX16@G2 | 114.683 | -115.060 | MTX16@G2-PEG | 486.265 | -136.608 |
| SN3816@G2 | -109.746 | -103.721 | SN3816@G2-PEG | -632.788 | -88.408 |
Table 2 Binding energy(Eb) of DN@PAMAM complexes with(or without) water
| DN@Gn | Eb(H2O)/ (kJ·mol-1) | Eb(no H2O)/ (kJ·mol-1) | DN@Gn-PEG | Eb(H2O)/ (kJ·mol-1) | Eb(no H2O)/ (kJ·mol-1) |
|---|---|---|---|---|---|
| CE68@G0 | -218.572 | -98.157 | CE68@G0-PEG | 154.808 | -138.951 |
| DOX8@G0 | -201.418 | -111.587 | DOX8@G0-PEG | 118.407 | -146.900 |
| MTX8@G0 | -205.476 | -82.299 | MTX8@G0-PEG | 166.900 | -145.478 |
| SN388@G0 | -137.277 | -84.224 | SN388@G0-PEG | 212.464 | -125.813 |
| CE610@G1 | -254.136 | -109.412 | CE612@G1-PEG | 224.555 | -144.766 |
| DOX10@G1 | -148.699 | -125.478 | DOX12@G1-PEG | 364.719 | -129.160 |
| MTX10@G1 | -210.079 | -101.546 | MTX12@G1-PEG | 301.834 | -112.089 |
| SN3810@G1 | -90.709 | -96.650 | SN3812@G1-PEG | 346.017 | -142.047 |
| CE616@G2 | -213.761 | -122.089 | CE616@G2-PEG | 378.610 | -141.126 |
| DOX16@G2 | -195.351 | -137.612 | DOX16@G2-PEG | 533.251 | -119.077 |
| MTX16@G2 | 114.683 | -115.060 | MTX16@G2-PEG | 486.265 | -136.608 |
| SN3816@G2 | -109.746 | -103.721 | SN3816@G2-PEG | -632.788 | -88.408 |
| DN@Gn | Ei-PD(H2O)/ (kJ·mol-1) | Ei-PD(no H2O)/ (kJ·mol-1) | DN@Gn-PEG | Ei-PD(H2O)/ (kJ·mol-1) | Eb(no H2O)/ (kJ·mol-1) |
|---|---|---|---|---|---|
| CE68@G0 | -161.126 | -37.447 | CE68@G0-PEG | -204.137 | -106.734 |
| DOX8@G0 | -174.389 | -20.209 | DOX8@G0-PEG | -253.969 | -133.260 |
| MTX8@G0 | -175.770 | -29.079 | MTX8@G0-PEG | -195.184 | -161.377 |
| SN388@G0 | -122.591 | -21.255 | SN388@G0-PEG | -167.862 | -130.708 |
| CE610@G1 | -164.640 | -54.057 | CE612@G1-PEG | -193.970 | -153.929 |
| DOX10@G1 | -182.841 | -54.099 | DOX12@G1-PEG | -192.087 | -165.184 |
| MTX10@G1 | -157.151 | -42.802 | MTX12@G1-PEG | -177.694 | -153.595 |
| SN3810@G1 | -121.545 | -40.417 | SN3812@G1-PEG | -120.792 | -93.430 |
| CE616@G2 | -153.678 | -60.250 | CE616@G2-PEG | -230.204 | -205.476 |
| DOX16@G2 | -173.887 | -69.036 | DOX16@G2-PEG | -212.589 | -151.586 |
| MTX16@G2 | -194.472 | -59.329 | MTX16@G2-PEG | -192.004 | -178.950 |
| SN3816@G2 | -121.503 | -49.162 | SN3816@G2-PEG | -162.088 | -133.553 |
Table 3 Interaction energies(Ei-PD) of DN@PAMAM complexes with(or without) water
| DN@Gn | Ei-PD(H2O)/ (kJ·mol-1) | Ei-PD(no H2O)/ (kJ·mol-1) | DN@Gn-PEG | Ei-PD(H2O)/ (kJ·mol-1) | Eb(no H2O)/ (kJ·mol-1) |
|---|---|---|---|---|---|
| CE68@G0 | -161.126 | -37.447 | CE68@G0-PEG | -204.137 | -106.734 |
| DOX8@G0 | -174.389 | -20.209 | DOX8@G0-PEG | -253.969 | -133.260 |
| MTX8@G0 | -175.770 | -29.079 | MTX8@G0-PEG | -195.184 | -161.377 |
| SN388@G0 | -122.591 | -21.255 | SN388@G0-PEG | -167.862 | -130.708 |
| CE610@G1 | -164.640 | -54.057 | CE612@G1-PEG | -193.970 | -153.929 |
| DOX10@G1 | -182.841 | -54.099 | DOX12@G1-PEG | -192.087 | -165.184 |
| MTX10@G1 | -157.151 | -42.802 | MTX12@G1-PEG | -177.694 | -153.595 |
| SN3810@G1 | -121.545 | -40.417 | SN3812@G1-PEG | -120.792 | -93.430 |
| CE616@G2 | -153.678 | -60.250 | CE616@G2-PEG | -230.204 | -205.476 |
| DOX16@G2 | -173.887 | -69.036 | DOX16@G2-PEG | -212.589 | -151.586 |
| MTX16@G2 | -194.472 | -59.329 | MTX16@G2-PEG | -192.004 | -178.950 |
| SN3816@G2 | -121.503 | -49.162 | SN3816@G2-PEG | -162.088 | -133.553 |
| DN@Gn | D/(nm2·ps-1) | DN@Gn-PEG | D/(nm2·ps-1) |
|---|---|---|---|
| CE68@G0 | 1.1×10-3 | CE68@G0-PEG | 6.4×10-4 |
| DOX8@G0 | 1.0×10-3 | DOX8@G0-PEG | 8.1×10-4 |
| MTX8@G0 | 1.0×10-3 | MTX8@G0-PEG | 8.8×10-4 |
| SN388@G0 | 9.6×10-4 | SN388@G0-PEG | 7.5×10-4 |
| CE610@G1 | 7.8×10-4 | CE612@G1-PEG | 7.0×10-4 |
| DOX10@G1 | 8.7×10-4 | DOX12@G1-PEG | 7.0×10-4 |
| MTX10@G1 | 1.2×10-3 | MTX12@G1-PEG | 7.9×10-4 |
| SN3810@G1 | 1.2×10-3 | SN3812@G1-PEG | 5.7×10-4 |
| CE616@G2 | 8.8×10-4 | CE616@G2-PEG | 5.0×10-4 |
| DOX16@G2 | 7.1×10-4 | DOX16@G2-PEG | 5.1×10-4 |
| MTX16@G2 | 6.6×10-4 | MTX16@G2-PEG | 5.5×10-4 |
| SN3816@G2 | 7.7×10-4 | SN3816@G2-PEG | 3.2×10-4 |
Table 4 Diffusion coefficients(D) of the DN@Gn and DN@Gn-PEG complexes in water
| DN@Gn | D/(nm2·ps-1) | DN@Gn-PEG | D/(nm2·ps-1) |
|---|---|---|---|
| CE68@G0 | 1.1×10-3 | CE68@G0-PEG | 6.4×10-4 |
| DOX8@G0 | 1.0×10-3 | DOX8@G0-PEG | 8.1×10-4 |
| MTX8@G0 | 1.0×10-3 | MTX8@G0-PEG | 8.8×10-4 |
| SN388@G0 | 9.6×10-4 | SN388@G0-PEG | 7.5×10-4 |
| CE610@G1 | 7.8×10-4 | CE612@G1-PEG | 7.0×10-4 |
| DOX10@G1 | 8.7×10-4 | DOX12@G1-PEG | 7.0×10-4 |
| MTX10@G1 | 1.2×10-3 | MTX12@G1-PEG | 7.9×10-4 |
| SN3810@G1 | 1.2×10-3 | SN3812@G1-PEG | 5.7×10-4 |
| CE616@G2 | 8.8×10-4 | CE616@G2-PEG | 5.0×10-4 |
| DOX16@G2 | 7.1×10-4 | DOX16@G2-PEG | 5.1×10-4 |
| MTX16@G2 | 6.6×10-4 | MTX16@G2-PEG | 5.5×10-4 |
| SN3816@G2 | 7.7×10-4 | SN3816@G2-PEG | 3.2×10-4 |
| [1] | Kavyani S., Amjad-Iranagh S., Modarress H., J. Phys. Chem. B, 2014, 118, 3257—3266 |
| [2] | Tomalia D. A., Baker H., Dewald J., Hall M. E., Kallos G., Martin S., Roeck J., Ryder J., Smith P., Polym. J., 1985, 17, 117—132 |
| [3] | Mignani S., Kazzouli S. E., Bousmina M., Majoral J. P., Adv. Drug. Deliver. Rev., 2013, 65, 1316—1330 |
| [4] | Wu L. P., Zhang Z. P., Guo Y., Wang Y., Lu W., Chem. J. Chinese Universities, 2011, 32(6), 1436—1444 |
| (吴丽平, 张政朴, 郭义, 王勇, 陆伟.高等学校化学学报, 2011,32(6), 1436—1444) | |
| [5] | Zhang Y., Thomas T. P., Lee K. H., Li M., Zong H., Desai A. M., Kotlyar A., Huang B. H., Banaszak Holl M. M., Baker J. R. Jr., Bioorgan. Med. Chem., 2011, 19, 2557—2564 |
| [6] | Cai J., Ai S. Y., Yin H. S., Shi W. J., Acta Chim. Sinica, 2009, 67(19), 2227—2232 |
| (蔡军, 艾仕云, 殷焕顺, 时伟杰.化学学报, 2009,67(19), 2227—2232) | |
| [7] | Esfand R., Tomalia D. A., Drug. Discov. Today, 2001, 6(8), 427—436 |
| [8] | Svenson S., Tomalia D. A., Adv. Drug. Deliver. Rev., 2012, 64, 102—115 |
| [9] | Tomalia D. A., Naylor A. M., Goddard III W. A., Angew. Chem. Int. Ed., 1990, 29, 138—175 |
| [10] | Han Q. R., Wang B. X., He X. M., Ding M. T., Xia H. P., Chem. J. Chinese Universities, 2009, 30(3), 629—631 |
| (韩巧荣, 王炳祥, 何旭敏, 丁马太, 夏海平.高等学校化学学报, 2009,30(3), 629—631) | |
| [11] | Lu W. T., Li G. P., Luo Y. J., Jin Y. J., Acta Chim. Sinica, 2008, 66(20), 2258—2262 |
| (卢文婷, 李国平, 罗运军, 靳玉娟.化学学报, 2008,66(20), 2258—2262) | |
| [12] | Lee C. C., MacKay J. A., Frechet J. M. J., Szoka F. C., Nat. Biotechnol., 2005, 23(12), 1517—1526 |
| [13] | Wiener E. C., Brechbiel M. W., Brothers H., Magin R. L., Gansow O. A., Tomalia D. A., Lauterbur P. C., Magn. Reson. Med., 1994, 31, 1—8 |
| [14] | Radu Daniela R., Lai C. Y., Jeftinija K., Rowe E. W., Jeftinija S., Lin V. S. Y., J. Am. Chem. Soc., 2004, 126(41), 13216—13217 |
| [15] | Domanski D. M., Klajnert B., Bryszewska M., Bioelectrochemistry, 2004, 63, 189—191 |
| [16] | Wang W., Xiong W., Zhu Y., Xu H., Yang X., J. Biomed. Mater. Res. B, 2010, 93B, 59—64 |
| [17] | He H., Li Y., Jia X. R., Du J., Ying X., Lu W. L., Lou J. N., Wei Y., Biomaterials.2011, 32, 478—487 |
| [18] | Sideratou Z., Kontoyianni C., Drossopoulou G. I., Paleos C. M., Bioorg. Med. Chem. Lett.2010, 20, 6513—6517 |
| [19] | Singh P., Gupta U., Asthana A., Jain N. K., Bioconjugate. Chem., 2008, 19, 2239—2252 |
| [20] | Yu H. Y., Tang Z. H., Song W. T., Deng M. X., Chen X. S., Chem. J. Chinese Universities, 2014, 35(5), 903—916 |
| (于海洋, 汤朝晖, 宋万通, 邓明虓, 陈学思.高等学校化学学报, 2014,35(5), 903—916) | |
| [21] | Goldberg D. S., Vijayalakshmi N., Swaan P. W., Ghandehari H., J. Control. Release., 2011, 150, 318—325 |
| [22] | Allen T. M., Cullis P. R., Science, 2004, 303(19), 1818—1822 |
| [23] | Tian W. D., Ma Y. Q., Soft. Matter., 2012, 8, 2627—2632 |
| [24] | Ye L., Jiang X. Z., Yang H., Su J. T., Yun L. H., Chem. J. Chinese Universities, 2005, 26(2), 353—355 |
| (叶玲, 江晓舟, 杨华, 苏健婷, 恽榴红.高等学校化学学报, 2005,26(2), 353—355) | |
| [25] | Liu Y., Bryantsev V. S., Diallo M. S., Goddard III W. A., J. Am. Chem. Soc., 2009, 131, 2798—2799 |
| [26] | Tomalia D. A., Reyna L. A., Svenson S., Biochem. Soc. Trans., 2007, 35, 61—67 |
| [27] | Gurdag S., Khandare J., Stapels S., Matherly L. H., Kannan R. M., Bioconjug. Chem., 2006, 17, 275—283 |
| [28] | Thomas T. P., Majoros I. J., Kotlyar A., Kukowska-Latallo J. F., Bielinska A., Myc A., Baker J. R. Jr., J. Med. Chem., 2005, 48, 3729—3735 |
| [29] | Kwon II. K., Lee S. C., Han B., Park K., J. Control. Release., 2012, 164, 108—114 |
| [30] | Fu Y., Jia G. F., Pang X. Q., Wang R. N., Wang X., Li C. J., Smemo S., Dai Q., Bailey K. A., Nobrega M. A., Han K. L., Cui Q., He C., Nat. Commun., 2013, 4, 1798—1805 |
| [31] | Pang X. Q., Yang M. J., Han K. L., Proteins, 2013, 81, 1399—1410 |
| [32] | Karatasos K., Krystallis M. J., Chem. Phys., 2009, 130, 1—11 |
| [33] | Zhong T. P., Ai P. F., Zhou J., Fluid. Phase. Equilibr., 2011, 302, 43—47 |
| [1] | WU Yushuai, SHANG Yingxu, JIANG Qiao, DING Baoquan. Research Progress of Controllable Self-assembled DNA Origami Structure as Drug Carrier [J]. Chem. J. Chinese Universities, 2022, 43(8): 20220179. |
| [2] | GAO Zhiwei, LI Junwei, SHI Sai, FU Qiang, JIA Junru, AN Hailong. Analysis of Gating Characteristics of TRPM8 Channel Based on Molecular Dynamics [J]. Chem. J. Chinese Universities, 2022, 43(6): 20220080. |
| [3] | HU Bo, ZHU Haochen. Dielectric Constant of Confined Water in a Bilayer Graphene Oxide Nanosystem [J]. Chem. J. Chinese Universities, 2022, 43(2): 20210614. |
| [4] | ZHANG Mi, TIAN Yafeng, GAO Keli, HOU Hua, WANG Baoshan. Molecular Dynamics Simulation of the Physicochemical Properties of Trifluoromethanesulfonyl Fluoride Dielectrics [J]. Chem. J. Chinese Universities, 2022, 43(11): 20220424. |
| [5] | LEI Xiaotong, JIN Yiqing, MENG Xuanyu. Prediction of the Binding Site of PIP2 in the TREK-1 Channel Based on Molecular Modeling [J]. Chem. J. Chinese Universities, 2021, 42(8): 2550. |
| [6] | LI Congcong, LIU Minghao, HAN Jiarui, ZHU Jingxuan, HAN Weiwei, LI Wannan. Theoretical Study of the Catalytic Activity of VmoLac Non-specific Substrates Based on Molecular Dynamics Simulations [J]. Chem. J. Chinese Universities, 2021, 42(8): 2518. |
| [7] | ZENG Yonghui, YAN Tianying. Vibrational Density of States Analysis of Proton Hydration Structure [J]. Chem. J. Chinese Universities, 2021, 42(6): 1855. |
| [8] | QI Renrui, LI Minghao, CHANG Hao, FU Xueqi, GAO Bo, HAN Weiwei, HAN Lu, LI Wannan. Theoretical Study on the Unbinding Pathway of Xanthine Oxidase Inhibitors Based on Steered Molecular Dynamics Simulation [J]. Chem. J. Chinese Universities, 2021, 42(3): 758. |
| [9] | LIU Aiqing, XU Wensheng, XU Xiaolei, CHEN Jizhong, AN Lijia. Molecular Dynamics Simulation of Polymer/rod Nanocomposite [J]. Chem. J. Chinese Universities, 2021, 42(3): 875. |
| [10] | FAN Ye, LI Qian, FANG Yun, XIA Yongmei. Fabrication of Lamellar Liquid Crystals of Conjugated Linoleic Acid as Drug Delivery Systems † [J]. Chem. J. Chinese Universities, 2020, 41(4): 750. |
| [11] | XUE Xuling,CHEN Jun,ZHANG Ziyou,WANG Mengmeng,Lü Mengdi,HAO Yuanyuan,HU Jiongsheng,GE Chao,SU Zhi,QIAN Yong,LIU Hongke. Arene-metal Complexes Based on Naproxen for Antitumor and Anti-inflammatory Applications † [J]. Chem. J. Chinese Universities, 2020, 41(2): 243. |
| [12] | WANG Xinghuo,TANG Jun,YANG Yingwei. Mesoporous Silica Nanoparticles-Based Stimuli-Responsive Drug Delivery Systems Gated by Polymers † [J]. Chem. J. Chinese Universities, 2020, 41(1): 28. |
| [13] | QU Siying, XU Qin. Different Roles of Some Key Residues in the S4 Pocket of Coagulation Factor Xa for Rivaroxaban Binding † [J]. Chem. J. Chinese Universities, 2019, 40(9): 1918. |
| [14] | MA Yucong, FAN Baomin, WANG Manman, YANG Biao, HAO Hua, SUN Hui, ZHANG Huijuan. Two-step Preparation of Trazodone and Its Corrosion Inhibition Mechanism for Carbon Steel [J]. Chem. J. Chinese Universities, 2019, 40(8): 1706. |
| [15] | LI Cuiqin, ZHAI Xue, SUN Peng, GAO Yuxin, ZHANG Zhiqiu, WANG Jun, LI Feng. Synthesis and Antioxidant Capacity of Dendrimer Bridged Antioxidant with Semi-hindered Phenolic Groups† [J]. Chem. J. Chinese Universities, 2019, 40(7): 1535. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||