

高等学校化学学报 ›› 2023, Vol. 44 ›› Issue (3): 20220333.doi: 10.7503/cjcu20220333
收稿日期:2022-05-13
出版日期:2023-03-10
发布日期:2023-03-14
通讯作者:
黄硕
E-mail:shuo.huang@nju.edu.cn
基金资助:
CHEN Jialu1,2, HUANG Shuo1,2( )
)
Received:2022-05-13
Online:2023-03-10
Published:2023-03-14
Contact:
HUANG Shuo
E-mail:shuo.huang@nju.edu.cn
Supported by:摘要:
DNA和RNA上广泛存在着多种化学修饰. 这些核酸修饰参与基因表达的调控, 影响生长发育等生理过程, 并可能会引发癌症等疾病. 对核酸修饰的精准识别与定位有助于理解其功能机制, 帮助相关疾病的诊断与治疗. 纳米孔测序是一种新兴的单分子测序技术, 可以根据修饰碱基与天然碱基之间阻孔信号的差异实现核酸序列中多种修饰的同时检测, 是目前检测核酸修饰最直接的方法. 本文简要介绍了纳米孔测序技术的发展和原理以及识别核酸修饰的算法工具, 总结了纳米孔测序技术在核酸修饰检测中的应用, 并对其发展前景进行了展望.
中图分类号:
TrendMD:
陈佳璐, 黄硕. 纳米孔测序技术在核酸修饰检测中的应用. 高等学校化学学报, 2023, 44(3): 20220333.
CHEN Jialu, HUANG Shuo. Applications of Nanopore Sequencing Technology in the Detection of Nucleic Acid Modifications. Chem. J. Chinese Universities, 2023, 44(3): 20220333.
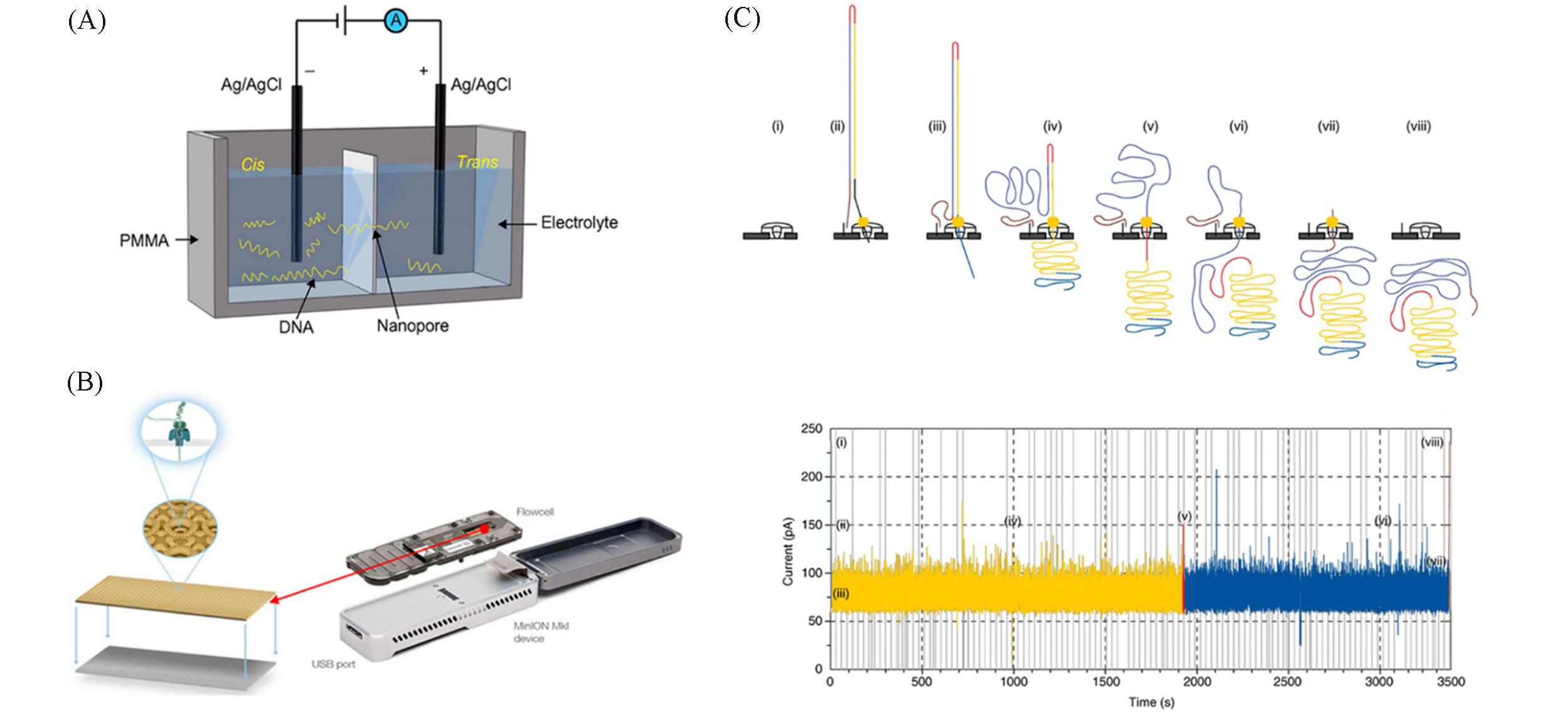
Fig.1 The basic principle of nanopore sequencing and MinION nanopore sequencer(A) Schematic view of nanopore setup comprising a DNA molecule through the nanopore[35]; (B) the composition of the MinION nanopore sequencer[46]; (C) steps in DNA translocation through nanopore and corresponding current trace[34].(A) Copyright 2015, the authors; (B) Copyright 2016, the authors; (C) Copyright 2016, the authors.
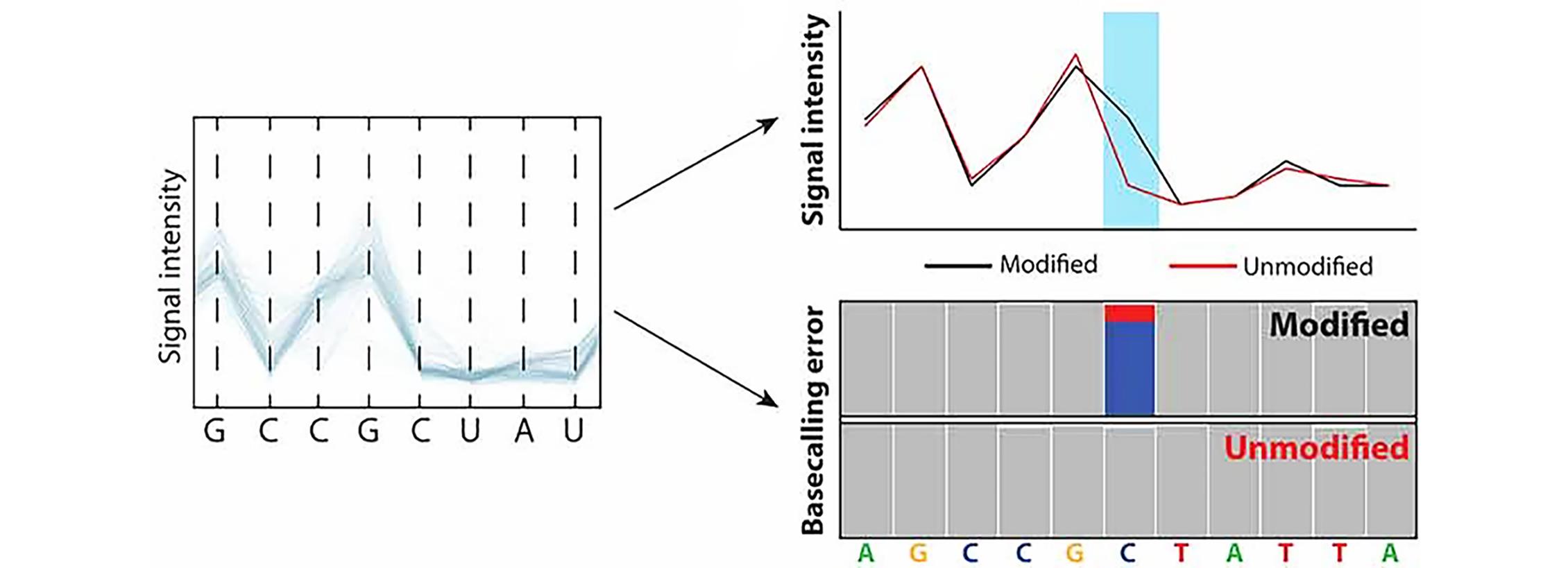
Fig.2 Schematic representation of two major approaches to detect nucleic acid modifications in nanopore sequencing data[51]The detection of nucleic acid modifications in the form of altered ionic current (up), or systematic base-calling errors(bottom). Copyright 2021, the authors.
| Method | Tool | Target | Modification type | Ref. | Method | Tool | Target | Modification type | Ref. |
|---|---|---|---|---|---|---|---|---|---|
| Ionic current based | nanopolish | DNA | 5mC | [ | Ionic current based | nanoraw | DNA | all | [ |
| method | signalAlign | DNA | 5mC, 5hmC, 6mA | [ | method | Nanocompore | RNA | all | [ |
| DeepMod | DNA | 5mC, 6mA | [ | xPore | RNA | all | [ | ||
| mCaller | DNA | 6mA | [ | nanoDoc | RNA | all | [ | ||
| DeepSignal | DNA | 5mC, 6mA | [ | Yanocomp | RNA | all | [ | ||
| CpelNano | DNA | 5mC | [ | Systematic base⁃calling | EpiNano | RNA | m6A | [ | |
| nanodisco | DNA | 4mC, 5mC, 6mA | [ | error based method | ELIGOS | RNA | m6A | [ | |
| Nanom6A | RNA | m6A | [ | DiffErr | RNA | m6A | [ | ||
| Penguin | RNA | Ψ | [ | DRUMMER | RNA | m6A | [ | ||
| MINES | RNA | m6A | [ | nanoRMS | RNA | Ψ, Nm | [ | ||
| NanoMod | DNA | all | [ |
Table1 Tools for nucleic acid modification detection from nanopore sequencing reads
| Method | Tool | Target | Modification type | Ref. | Method | Tool | Target | Modification type | Ref. |
|---|---|---|---|---|---|---|---|---|---|
| Ionic current based | nanopolish | DNA | 5mC | [ | Ionic current based | nanoraw | DNA | all | [ |
| method | signalAlign | DNA | 5mC, 5hmC, 6mA | [ | method | Nanocompore | RNA | all | [ |
| DeepMod | DNA | 5mC, 6mA | [ | xPore | RNA | all | [ | ||
| mCaller | DNA | 6mA | [ | nanoDoc | RNA | all | [ | ||
| DeepSignal | DNA | 5mC, 6mA | [ | Yanocomp | RNA | all | [ | ||
| CpelNano | DNA | 5mC | [ | Systematic base⁃calling | EpiNano | RNA | m6A | [ | |
| nanodisco | DNA | 4mC, 5mC, 6mA | [ | error based method | ELIGOS | RNA | m6A | [ | |
| Nanom6A | RNA | m6A | [ | DiffErr | RNA | m6A | [ | ||
| Penguin | RNA | Ψ | [ | DRUMMER | RNA | m6A | [ | ||
| MINES | RNA | m6A | [ | nanoRMS | RNA | Ψ, Nm | [ | ||
| NanoMod | DNA | all | [ |
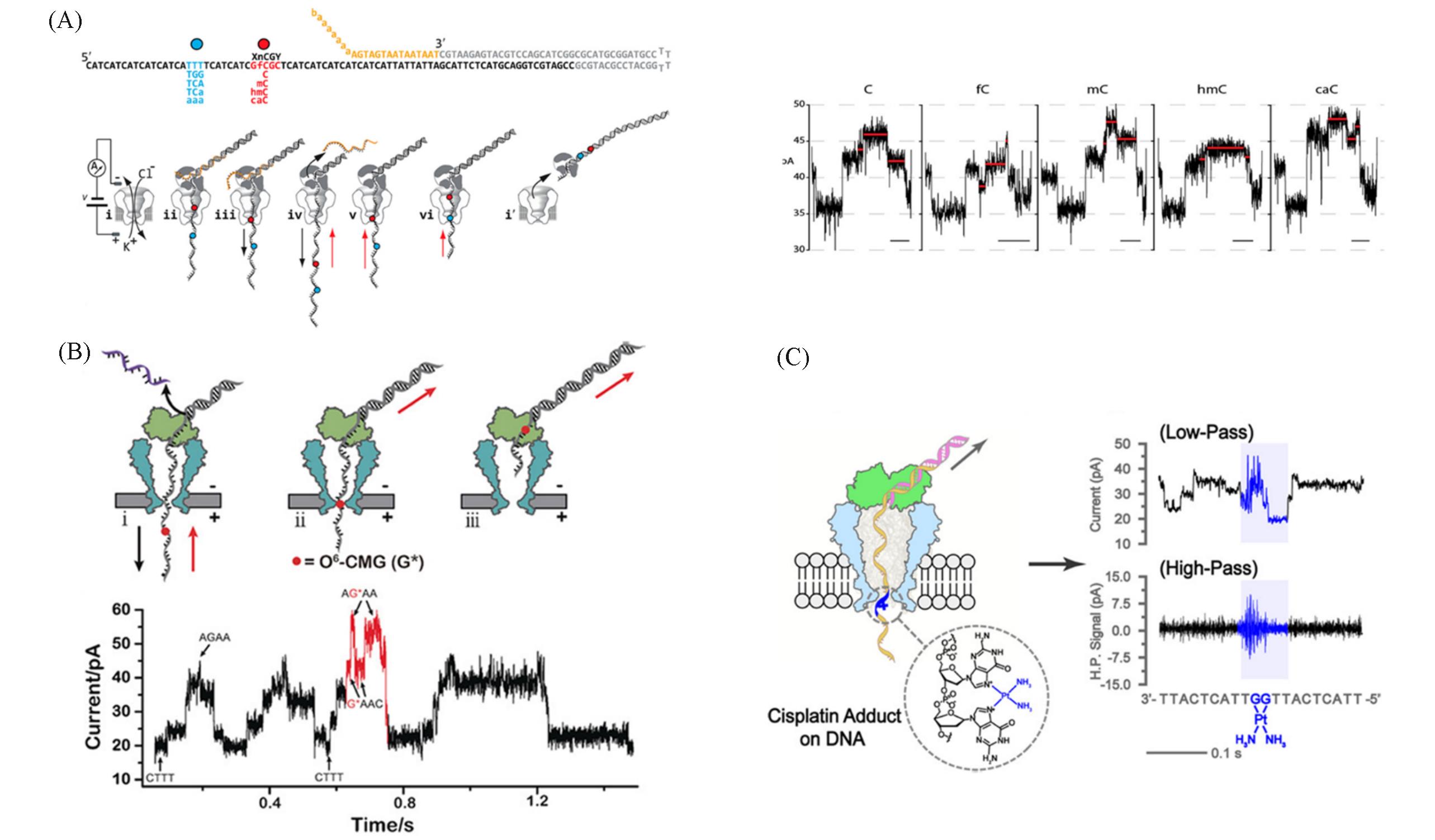
Fig.3 Discrimination of DNA modifications by nanopore sequencing(A) Strategy for reading C5-cytosine modifications along individual DNA templates using a nanopore and representative current traces for C5-cytosine modifications in the GnCGC 4-mer context[75]; (B) schematic of O6-CMG detection during nanopore sequencing and a representative current trace is shown below[76]; (C) schematic of cisplatin adduct detection during nanopore sequencing. Representative traces with low-pass or high-pass digital signal filtration treatment were shown on the right[77].(A) Copyright 2014, American Chemical Society; (B) Copyright 2019, Wiley-VCH; (C) Copyright 2021, American Chemical Society.
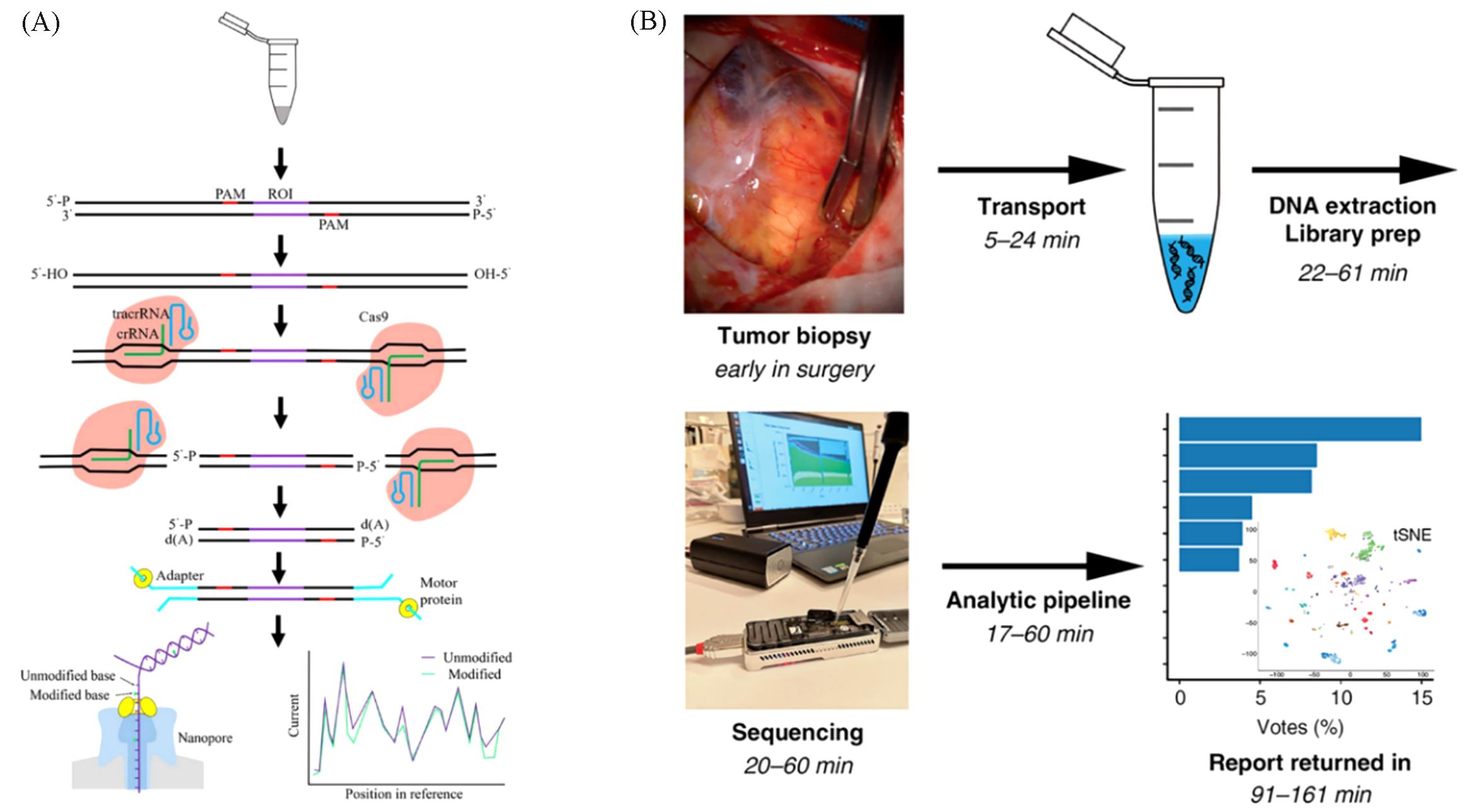
Fig.4 Application of nanopore sequencing to identify DNA modifications(A) Schematic of nanopore Cas9-targeted sequencing (nCATS)[95]; (B) the workflow of nanopore DNA methylation analysis which enables the detailed and rapid classification of brain tumors[100].(A) Copyright 2021, the authors; (B) Copyright 2021, the authors.
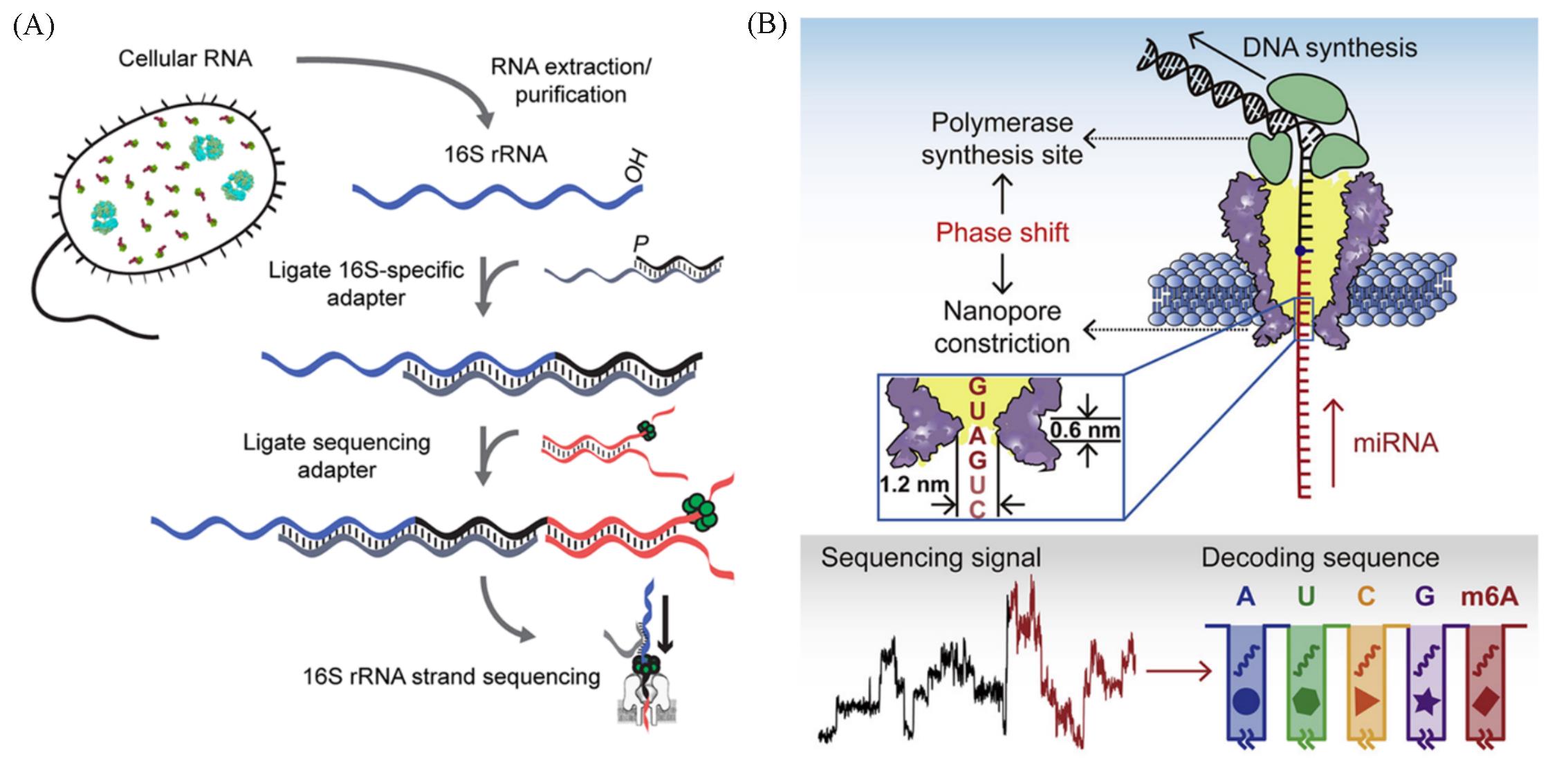
Fig.5 Nanopore sequencing and modification detection of different kinds of RNA(A) Library preparation for MinION sequencing of 16S rRNA strands[102]; (B) the principle and sequencing signal of nanopore-induced phase-shift sequencing (NIPSS) strategy for direct miRNA sequencing[104].(A) Copyright 2019, the authors; (B) Copyright 2021, the authors.
| 1 | Sood A. J., Viner C., Hoffman M. M., J. Cheminform., 2019, 11(1), 30 |
| 2 | Boccaletto P., Stefaniak F., Ray A., Cappannini A., Mukherjee S., Purta E., Kurkowska M., Shirvanizadeh N., Destefanis E., Groza P., Avşar G., Romitelli A., Pir P., Dassi E., Conticello S. G., Aguilo F., Bujnicki J. M., Nucleic Acids Res., 2021, 50(D1), D231—D235 |
| 3 | Chen K., Zhao Boxuan S., He C., Cell Chem. Biol., 2016, 23(1), 74—85 |
| 4 | Fu Y., He C., Curr. Opin. Chem. Biol., 2012, 16(5), 516—524 |
| 5 | Deobagkar D. D., Chandra H. S., J. Genet., 2003, 82(1), 13 |
| 6 | Yamaguchi S., Shen L., Liu Y., Sendler D., Zhang Y., Nature, 2013, 504(7480), 460—464 |
| 7 | Smith Z. D., Chan M. M., Mikkelsen T. S., Gu H., Gnirke A., Regev A., Meissner A., Nature, 2012, 484(7394), 339—344 |
| 8 | La H., Ding B., Mishra G. P., Zhou B., Yang H., Bellizzi M. D. R., Chen S., Meyers B. C., Peng Z., Zhu J. K., Wang G. L., Proc. Natl. Acad. Sci., 2011, 108(37), 15498—15503 |
| 9 | Guo J. U., Su Y., Zhong C., Ming G. l., Song H., Cell Cycle, 2011, 10(16), 2662—2668 |
| 10 | Plongthongkum N., Diep D. H., Zhang K., Nat. Rev. Genet., 2014, 15(10), 647—661 |
| 11 | Wang X., Lu Z., Gomez A., Hon G. C., Yue Y., Han D., Fu Y., Parisien M., Dai Q., Jia G., Ren B., Pan T., He C., Nature, 2014, 505(7481), 117—120 |
| 12 | Zhao X., Yang Y., Sun B. F., Shi Y., Yang X., Xiao W., Hao Y. J., Ping X. L., Chen Y. S., Wang W. J., Jin K. X., Wang X., Huang C. M., Fu Y., Ge X. M., Song S. H., Jeong H. S., Yanagisawa H., Niu Y., Jia G. F., Wu W., Tong W. M., Okamoto A., He C., Danielsen J. M. R., Wang X. J., Yang Y. G., Cell Res., 2014, 24(12), 1403—1419 |
| 13 | Alarcón C. R., Lee H., Goodarzi H., Halberg N., Tavazoie S. F., Nature, 2015, 519(7544), 482—485 |
| 14 | Coots R. A., Liu X. M., Mao Y., Dong L., Zhou J., Wan J., Zhang X., Qian S. B., Mol. Cell, 2017, 68(3), 504—514 |
| 15 | Vu L. P., Pickering B. F., Cheng Y., Zaccara S., Nguyen D., Minuesa G., Chou T., Chow A., Saletore Y., MacKay M., Schulman J., Famulare C., Patel M., Klimek V. M., Garrett⁃Bakelman F. E., Melnick A., Carroll M., Mason C. E., Jaffrey S. R., Kharas M. G., Nat. Med., 2017, 23(11), 1369—1376 |
| 16 | Liu J., Eckert M. A., Harada B. T., Liu S. M., Lu Z., Yu K., Tienda S. M., Chryplewicz A., Zhu A. C., Yang Y., Huang J. T., Chen S. M., Xu Z. G., Leng X. H., Yu X. C., Cao J., Zhang Z., Liu J., Lengyel E., He C., Nat. Cell Biol., 2018, 20(9), 1074—1083 |
| 17 | Yue B., Song C., Yang L., Cui R., Cheng X., Zhang Z., Zhao G., Mol. Cancer, 2019, 18(1), 142 |
| 18 | Wu G., Adachi H., Ge J., Stephenson D., Query C. C., Yu Y. T., EMBO J., 2016, 35(6), 654—667 |
| 19 | Chen C., Zhao X., Kierzek R., Yu Y. T., Mol. Cell Biol., 2010, 30(17), 4108—4119 |
| 20 | Lorenz C., Lünse C. E., Mörl M., Biomolecules, 2017, 7(2), 35 |
| 21 | Li X., Ma S., Yi C., Curr. Opin. Chem. Biol., 2016, 33, 108—116 |
| 22 | Leppik M., Liiv A., Remme J., Nucleic Acids Res., 2017, 45(10), 6098—6108 |
| 23 | Kierzek E., Malgowska M., Lisowiec J., Turner D. H., Gdaniec Z., Kierzek R., Nucleic Acids Res., 2013, 42(5), 3492—3501 |
| 24 | Jora M., Lobue P. A., Ross R. L., Williams B., Addepalli B., BBA⁃Gene Regul. Mech., 2019, 1862(3), 280—290 |
| 25 | Lai W., Mo J., Yin J., Lyu C., Wang H., Trends. Anal. Chem., 2019, 110, 173—182 |
| 26 | Dominissini D., Moshitch⁃Moshkovitz S., Schwartz S., Salmon⁃Divon M., Ungar L., Osenberg S., Cesarkas K., Jacob⁃Hirsch J., Amariglio N., Kupiec M., Sorek R., Rechavi G., Nature, 2012, 485(7397), 201—206 |
| 27 | Meyer Kate D., Saletore Y., Zumbo P., Elemento O., Mason C. E., Affrey S. R., Cell, 2012, 149(7), 1635—1646 |
| 28 | Frommer M., McDonald L. E., Millar D. S., Collis C. M., Watt F., Grigg G. W., Molloy P. L., Paul C. L., Proc. Natl. Acad. Sci., 1992, 89(5), 1827—1831 |
| 29 | Schutsky E. K., DeNizio J. E., Hu P., Liu M. Y., Nabel C. S., Fabyanic E. B., Hwang Y., Bushman F. D., Wu H., Kohli R. M., Nat. Biotechnol., 2018, 36(11), 1083—1090 |
| 30 | Liu Y., Siejka⁃Zielińska P., Velikova G., Bi Y., Yuan F., Tomkova M., Bai C., Chen L., Schuster⁃Böckler B., Song C. X., Nat. Biotechnol., 2019, 37(4), 424—429 |
| 31 | Flusberg B. A., Webster D. R., Lee J. H., Travers K. J., Olivares E. C., Clark T. A., Korlach J., Turner S. W., Nat. Meth., 2010, 7(6), 461—465 |
| 32 | Huang S., Chin. Sci. Bull., 2014, 59(35), 4918—4928 |
| 33 | Xu L., Seki M., J. Hum. Genet., 2020, 65(1), 25—33 |
| 34 | Jain M., Olsen H. E., Paten B., Akeson M., Genome Biol., 2016, 17(1), 239 |
| 35 | Feng Y., Zhang Y., Ying C., Wang D., Du C., Genom. Proteom. Bioinf., 2015, 13(1), 4—16 |
| 36 | Stoddart D., Heron A. J., Mikhailova E., Maglia G., Bayley H., Proc. Natl. Acad. Sci., 2009, 106(19), 7702—7707 |
| 37 | Deamer D., Akeson M., Branton D., Nat. Biotechnol., 2016, 34(5), 518—524 |
| 38 | Wolna A. H., Fleming A. M., An N., He L., White H. S., Burrows C. J., Isr. J. Chem., 2013, 53(6/7), 417—430 |
| 39 | Schibel A. E. P., An N., Jin Q., Fleming A. M., Burrows C. J., White H. S., J. Am. Chem. Soc., 2010, 132(51), 17992—17995 |
| 40 | Wallace E. V. B., Stoddart D., Heron A. J., Mikhailova E., Maglia G., Donohoe T. J., Bayley H., Chem. Commun., 2010, 46(43), 8195—8197 |
| 41 | An N., White H. S., Burrows C. J., Chem. Commun., 2012, 48(93), 11410—11412 |
| 42 | Ayub M., Bayley H., Nano Lett., 2012, 12(11), 5637—5643 |
| 43 | Li W. W., Gong L. Z., Bayley H., Angew. Chem. Int. Ed., 2013, 52(16), 4350—4355 |
| 44 | Cherf G. M., Lieberman K. R., Rashid H., Lam C. E., Karplus K., Akeson M., Nat. Biotechnol., 2012, 30(4), 344—348 |
| 45 | Manrao E. A., Derrington I. M., Laszlo A. H., Langford K. W., Hopper M. K., Gillgren N., Pavlenok M., Niederweis M., Gundlach J. H., Nat. Biotechnol., 2012, 30(4), 349—353 |
| 46 | Lu H., Giordano F., Ning Z., Genom. Proteom. Bioinf., 2016, 14(5), 265—279 |
| 47 | Wang Y., Zhao Y., Bollas A., Wang Y., Au K. F., Nat. Biotechnol., 2021, 39(11), 1348—1365 |
| 48 | Garalde D. R., Snell E. A., Jachimowicz D., Sipos B., Lloyd J. H., Bruce M., Pantic N., Admassu T., James P., Warland A., Jordan M., Ciccone J., Serra S., Keenan J., Martin S., McNeill L., Wallace E. J., Jayasinghe L., Wright C., Blasco J., Young S., Brocklebank D., Juul S., Clarke J., Heron A. J., Turner D. J., Nat. Meth., 2018, 15(3), 201—206 |
| 49 | Kerkhof L. J., FEMS Microbiol. Ecol., 2021, 97(3), fiab001 |
| 50 | Wan Y. K., Hendra C., Pratanwanich P. N., Göke J., Trends Genet., 2021, 38(3), 246—257 |
| 51 | Furlan M., Delgado⁃Tejedor A., Mulroney L., Pelizzola M., Novoa E. M., Leonardi T., RNA Biol., 2021, 18(sup. 1), 31—40 |
| 52 | Simpson J. T., Workman R. E., Zuzarte P. C., David M., Dursi L. J., Timp W., Nat. Meth., 2017, 14(4), 407—410 |
| 53 | Rand A. C., Jain M., Eizenga J. M., Musselman⁃Brown A., Olsen H. E., Akeson M., Paten B., Nat. Meth., 2017, 14(4), 411—413 |
| 54 | Liu Q., Fang L., Yu G., Wang D., Xiao C. L., Wang K., Nat. Commun., 2019, 10(1), 2449 |
| 55 | McIntyre A. B. R., Alexander N., Grigorev K., Bezdan D., Sichtig H., Chiu C. Y., Mason C. E., Nat. Commun., 2019, 10(1), 579 |
| 56 | Ni P., Huang N., Zhang Z., Wang D. P., Liang F., Miao Y., Xiao C. L., Luo F., Wang J., Bioinformatics, 2019, 35(22), 4586—4595 |
| 57 | Abante J., Kambhampati S., Feinberg A. P., Goutsias J., Sci. Rep., 2021, 11(1), 21619 |
| 58 | Tourancheau A., Mead E. A., Zhang X. S., Fang G., Nat. Meth., 2021, 18(5), 491—498 |
| 59 | Gao Y., Liu X., Wu B., Wang H., Xi F., Kohnen M. V., Reddy A. S. N., Gu L., Genome Biol., 2021, 22(1), 22 |
| 60 | Hassan D., Acevedo D., Daulatabad S. V., Mir Q., Janga S. C., Methods, 2022, 203, 478—487 |
| 61 | Liu Q., Georgieva D. C., Egli D., Wang K., BMC Genomics, 2019, 20(1), 78 |
| 62 | Stoiber M., Quick J., Egan R., Eun Lee J., Celniker S., Neely R. K., Loman N., Pennacchio L. A., Brown J., bioRxiv, 2017, 094672 |
| 63 | Lorenz D. A., Sathe S., Einstein J. M., Yeo G. W., RNA, 2020, 26(1), 19—28 |
| 64 | Leger A., Amaral P. P., Pandolfini L., Capitanchik C., Capraro F., Miano V., Migliori V., Toolan⁃Kerr P., Sideri T., Enright A. J., Tzelepis K., van Werven F. J., Luscombe N. M., Barbieri I., Ule J., Fitzgerald T., Birney E., Leonardi T., Kouzarides T., Nat. Commun., 2021, 12(1), 7198 |
| 65 | Pratanwanich P. N., Yao F., Chen Y., Koh C. W. Q., Wan Y. K., Hendra C., Poon P., Goh Y. T., Yap P. M. L., Chooi J. Y., Chng W. J., Ng S. B., Thiery A., Goh W. S. S., Göke J., Nat. Biotechnol., 2021, 39(11), 1394—1402 |
| 66 | Ueda H., bioRxiv., 2021, 295089 |
| 67 | Parker M. T., Barton G. J., Simpson G. G., bioRxiv., 2021, 448494 |
| 68 | Begik O., Lucas M. C., Pryszcz L. P., Ramirez J. M., Medina R., Milenkovic I., Cruciani S., Liu H., Vieira H. G. S., Sas⁃Chen A., Mattick J. S., Schwartz S., Novoa E. M., Nat. Biotechnol., 2021, 39(10), 1278—1291 |
| 69 | Liu H., Begik O., Lucas M. C., Ramirez J. M., Mason C. E., Wiener D., Schwartz S., Mattick J. S., Smith M. A., Novoa E. M., Nat. Commun., 2019, 10(1), 4079 |
| 70 | Jenjaroenpun P., Wongsurawat T., Wadley T. D., Wassenaar Trudy M., Liu J., Dai Q., Wanchai V., Akel N. S., Jamshidi⁃Parsian A., Franco A. T., Boysen G., Jennings M. L., Ussery D. W., He C., Nookaew I., Nucleic Acids Res., 2020, 49(2), e7 |
| 71 | Parker M. T., Knop K., Sherwood A. V., Schurch N. J., Mackinnon K., Gould P. D., Hall A. J. W., Barton G. J., Simpson G. G., eLife, 2020, 9, e49658 |
| 72 | Price A. M., Hayer K. E., McIntyre A. B. R., Gokhale N. S., Abebe J. S., Della Fera A. N., Mason C. E., Horner S. M., Wilson A. C., Depledge D. P., Weitzman M. D., Nat. Commun., 2020, 11(1), 6016 |
| 73 | Laszlo A. H., Derrington I. M., Brinkerhoff H., Langford K. W., Nova I. C., Samson J. M., Bartlett J. J., Pavlenok M., Gundlach J. H., Proc. Natl. Acad. Sci., 2013, 110(47), 18904—18909 |
| 74 | Schreiber J., Wescoe Z. L., Abu⁃Shumays R., Vivian J. T., Baatar B., Karplus K., Akeson M., Proc. Natl. Acad. Sci., 2013, 110(47), 18910—18915 |
| 75 | Wescoe Z. L., Schreiber J., Akeson M., J. Am. Chem. Soc., 2014, 136(47), 16582—16587 |
| 76 | Wang Y., Patil K. M., Yan S., Zhang P., Guo W., Wang Y., Chen H. Y., Gillingham D., Huang S., Angew. Chem. Int. Ed., 2019, 58(25), 8432—8436 |
| 77 | Ma F., Yan S., Zhang J., Wang Y., Wang L., Wang Y., Zhang S., Du X., Zhang P., Chen H. Y., Huang S., ACS Sensors, 2021, 6(8), 3082—3092 |
| 78 | Jain M., Koren S., Miga K. H., Quick J., Rand A. C., Sasani T. A., Tyson J. R., Beggs A. D., Dilthey A. T., Fiddes I. T., Malla S., Marriott H., Nieto T., O'Grady J., Olsen H. E., Pedersen B. S., Rhie A., Richardson H., Quinlan A. R., Snutch T. P., Tee L., Paten B., Phillippy A. M., Simpson J. T., Loman N. J., Loose M., Nat. Biotechnol., 2018, 36(4), 338—345 |
| 79 | Lee I., Razaghi R., Gilpatrick T., Molnar M., Gershman A., Sadowski N., Sedlazeck F. J., Hansen K. D., Simpson J. T., Timp W., Nat. Meth., 2020, 17(12), 1191—1199 |
| 80 | Bicci I., Calabrese C., Golder Z. J., Gomez⁃Duran A., Chinnery Patrick F., Nucleic Acids Res., 2021, 49(22), 12757—12768 |
| 81 | Jenjaroenpun P., Wongsurawat T., Pereira R., Patumcharoenpol P., Ussery D. W., Nielsen J., Nookaew I., Nucleic Acids Res., 2018, 46(7), e38 |
| 82 | Kot W., Olsen N. S., Nielsen T. K., Hutinet G., de Crécy⁃Lagard V., Cui L., Dedon P. C., Carstens A. B., Moineau S., Swairjo M. A., Hansen L. H., Nucleic Acids Res., 2020, 48(18), 10383—10396 |
| 83 | Afonin A. M., Gribchenko E. S., Zorin E. A., Sulima A. S., Zhukov V. A., Microorganisms, 2021, 9(12), 2458 |
| 84 | Shipony Z., Marinov G. K., Swaffer M. P., Sinnott⁃Armstrong N. A., Skotheim J. M., Kundaje A., Greenleaf W. J., Nat. Meth., 2020, 17(3), 319—327 |
| 85 | Wang Y., Wang A., Liu Z., Thurman A. L., Powers L. S., Zou M., Zhao Y., Hefel A., Li Y., Zabner J., Au K. F., Genome Res., 2019, 29(8), 1329—1342 |
| 86 | Gigante S., Gouil Q., Lucattini A., Keniry A., Beck T., Tinning M., Gordon L., Woodruff C., Speed T. P., Blewitt M. E., Ritchie M. E., Nucleic Acids Res., 2019, 47(8), e46 |
| 87 | Shah K., Cao W., Ellison C. E., G3 Genes Genom. Genet., 2019, 9(6), 1893—1900 |
| 88 | Urban J. M., Foulk M. S., Bliss J. E., Coleman C. M., Lu N., Mazloom R., Brown S. J., Spradling A. C., Gerbi S. A., BMC Genomics, 2021, 22(1), 643 |
| 89 | Wanner N., Larsen P. A., McLain A., Faulk C., BMC Genomics, 2021, 22(1), 726 |
| 90 | Dimond J. L., Nguyen N., Roberts S. B., G3 Genes Genom. Genet., 2021, 11(7), jkab148 |
| 91 | Zhang G., Diao S., Song Y., He C., Zhang J., Tree Physiol., 2022, tpab177 |
| 92 | Toh T. B., Lim J. J., Chow E. K. H., Clin. Transl. Med., 2019, 8(1), e13 |
| 93 | Davenport C. F., Scheithauer T., Dunst A., Bahr F. S., Dorda M., Wiehlmann L., Tran D. D. H., Int. J. Mol. Sci., 2021, 22(8), 3937 |
| 94 | Gilpatrick T., Lee I., Graham J. E., Raimondeau E., Bowen R., Heron A., Downs B., Sukumar S., Sedlazeck F. J., Timp W., Nat. Biotechnol., 2020, 38(4), 433—438 |
| 95 | Zhang J., Xie S., Xu J., Liu H., Wan S., Front. Genet., 2021, 12(657), 672804 |
| 96 | McKelvey B. A., Gilpatrick T., Wang Y., Timp W., Umbricht C. B., Zeiger M. A., Thyroid, 2020, 30(10), 1470—1481 |
| 97 | Goldsmith C., Cohen D., Dubois A., Martinez M. G., Petitjean K., Corlu A., Testoni B., Hernandez⁃Vargas H., Chemin I., Microb. Genom., 2021, 7(5), 000507 |
| 98 | Wongsurawat T., Jenjaroenpun P., De Loose A., Alkam D., Ussery D. W., Nookaew I., Leung Y. K., Ho S. M., Day J. D., Rodriguez A., Acta Neuropathol. Com., 2020, 8(1), 87 |
| 99 | Euskirchen P., Bielle F., Labreche K., Kloosterman W. P., Rosenberg S., Daniau M., Schmitt C., Masliah⁃Planchon J., Bourdeaut F., Dehais C., Marie Y., Delattre J. Y., Idbaih A., Acta Neuropathol., 2017, 134(5), 691—703 |
| 100 | Djirackor L., Halldorsson S., Niehusmann P., Leske H., Capper D., Kuschel L. P., Pahnke J., Due⁃Tønnessen B. J., Langmoen I. A., Sandberg C. J., Euskirchen P., Vik⁃Mo E. O., Neuro⁃Oncol. Adv., 2021, 3(1), vdab149 |
| 101 | Stark R., Grzelak M., Hadfield J., Nat. Rev. Genet., 2019, 20(11), 631—656 |
| 102 | Smith A. M., Jain M., Mulroney L., Garalde D. R., Akeson M., PLoS One, 2019, 14(5), e0216709 |
| 103 | Wang Y., Wang H., Xi F., Wang H., Han X., Wei W., Zhang H., Zhang Q., Zheng Y., Zhu Q., Kohnen M. V., Reddy A. S. N., Gu L., J. Integr. Plant Biol., 2020, 62(12), 1823—1838 |
| 104 | Zhang J., Yan S., Chang L., Guo W., Wang Y., Wang Y., Zhang P., Chen H. Y., Huang S., iScience, 2020, 23(3), 100916 |
| 105 | Workman R. E., Tang A. D., Tang P. S., Jain M., Tyson J. R., Razaghi R., Zuzarte P. C., Gilpatrick T., Payne A., Quick J., Sadowski N., Holmes N., de Jesus J. G., Jones K. L., Soulette C. M., Snutch T. P., Loman N., Paten B., Loose M., Simpson J. T., Olsen H. E., Brooks A. N., Akeson M., Timp W., Nat. Meth., 2019, 16(12), 1297—1305 |
| 106 | Bilinovich S. M., Uhl K. L., Lewis K., Soehnlen X., Williams M., Vogt D., Prokop J. W., Campbell D. B., Dev. Neurosci., 2020, 42(5/6), 195—207 |
| 107 | Huang S., Zhang W., Katanski C. D., Dersh D., Dai Q., Lolans K., Yewdell J., Eren A. M., Pan T., Genome Biol., 2021, 22(1), 330 |
| 108 | Li R., Ren X., Ding Q., Bi Y., Xie D., Zhao Z., Genome Res., 2020, 30(2), 287—298 |
| 109 | Watabe E., Togo⁃Ohno M., Ishigami Y., Wani S., Hirota K., Kimura⁃Asami M., Hasan S., Takei S., Fukamizu A., Suzuki Y., Suzuki T., Kuroyanagi H., EMBO J., 2021, 40(14), e106434 |
| 110 | Zhang S., Li R., Zhang L., Chen S., Xie M., Yang L., Xia Y., Foyer C. H., Zhao Z., Lam H. M., Nucleic Acids Res., 2020, 48(14), 7700—7711 |
| 111 | Xu T., Wu X., Wong C. E., Fan S., Zhang Y., Zhang S., Liang Z., Yu H., Shen L., Adv. Sci., 2022, 9(6), 2103628 |
| 112 | Srinivas K. P., Depledge D. P., Abebe J. S., Rice S. A., Mohr I., Wilson A. C., Proc. Natl. Acad. Sci., 2021, 118(30), e2104805118 |
| 113 | Adrian, Viehweger, Sebastian, Krautwurst, Kevin, Lamkiewicz, Ramakanth, Madhugiri, John, Ziebuhr, Genome Res., 2019, 29(9), 1545—1554 |
| 114 | Kim D., Lee J. Y., Yang J. S., Kim J. W., Kim V. N., Chang H., Cell, 2020, 181(4), 914—921 |
| 115 | Chang J. J. Y., Rawlinson D., Pitt M. E., Taiaroa G., Gleeson J., Zhou C., Mordant F. L., de Paoli⁃Iseppi R., Caly L., Purcell D. F. J., Stinear T. P., Londrigan S. L., Clark M. B., Williamson D. A., Subbarao K., Coin L. J. M., Cell Rep., 2021, 35(6), 109108 |
| 116 | Fleming A. M., Mathewson N. J., Howpay Manage S. A., Burrows C. J., ACS Central Sci., 2021, 7(10), 1707—1717 |
| 117 | Campos J. H. C., Maricato J. T., Braconi C. T., Antoneli F., Janini L. M. R., Briones M. R. S., Viruses, 2021, 13(11), 2108 |
| 118 | Zhang X., Hao H., Ma L., Zhang Y., Hu X., Chen Z., Liu D., Yuan J., Hu Z., Guan W., Meng X. J., mBio., 2021, 12(4), e01067⁃21 |
| 119 | Brinkerhoff H., Kang A. S. W., Liu J., Aksimentiev A., Dekker C., Science, 2021, 374(6574), 1509—1513 |
| 120 | Chen Z., Wang Z., Xu Y., Zhang X., Tian B., Bai J., Chemical Science, 2021, 12(47), 15750—15756 |
| 121 | Yan S., Zhang J., Wang Y., Guo W., Zhang S., Liu Y., Cao J., Wang Y., Wang L., Ma F., Zhang P., Chen H. Y., Huang S., Nano Lett., 2021, 21(15), 6703—6710 |
| [1] | 方鑫, 赵瑞奇, 莫婧, 王雅芬, 翁小成. 检测核酸表观遗传修饰的测序方法[J]. 高等学校化学学报, 2023, 44(3): 20220342. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||