

高等学校化学学报 ›› 2021, Vol. 42 ›› Issue (7): 2161.doi: 10.7503/cjcu20210245
收稿日期:2021-04-12
出版日期:2021-07-10
发布日期:2021-06-22
通讯作者:
张增辉
E-mail:zhzhang@phy.ecnu.edu.cn
基金资助:
WEN Wei1, HUANG Dading1, BAO Jingxiao1, ZHANG John Z. H.1,2,3( )
)
Received:2021-04-12
Online:2021-07-10
Published:2021-06-22
Contact:
ZHANG John Z. H.
E-mail:zhzhang@phy.ecnu.edu.cn
摘要:
通过分子动力学模拟检测了2种程序性细胞死亡蛋白(PD-1)/单克隆抗体(Pembrolizumab和Nivolumab)复合物, 并使用高效的计算丙氨酸扫描方法预测了单抗与PD-1的结合热点, 将它们与对PD-1/PD-L1结合重要的热点残基进行对比分析. 结果显示, Pembrolizumab以类似于PD-L1的方式与PD-1结合, 而Nivolumab则以不同的方式与PD-1结合. 2个PD-1/mAb复合物中共有的热点只有PD-1K131. 同时发现, 与PD-1K131结合的单抗的关键残基通常都受范德华(vdW)能量控制. 2种单克隆抗体上热点的自由能贡献都以vdW能量为主, 这表明在下一代PD-1新抗体的设计中需要提高静电型热点残基的数量.
中图分类号:
TrendMD:
温炜, 黄达锭, 鲍劲霄, 张增辉. PD-1与单克隆抗体残基特异性结合机制的计算丙氨酸扫描研究. 高等学校化学学报, 2021, 42(7): 2161.
WEN Wei, HUANG Dading, BAO Jingxiao, ZHANG John Z. H.. Residue Specific Binding Mechanisms of PD-1 to Its Monoclonal Antibodies by Computational Alanine Scanning. Chem. J. Chinese Universities, 2021, 42(7): 2161.
| Species | Chain | Mutation | ΔΔEvdW/(kJ·mol-1) | ΔΔEele/ (kJ·mol-1) | ΔΔGgb/ (kJ·mol-1) | ΔΔGnp/(kJ·mol-1) | ΔΔH/ (kJ·mol-1) | -TΔΔS/(kJ·mol-1) | ΔΔG/ (kJ·mol-1) | SD* |
|---|---|---|---|---|---|---|---|---|---|---|
| PD?1 | Z | D85A | -3.13 | 85.36 | -56.39 | -0.04 | 25.79 | -9.24 | 16.55 | 1.30 |
| Z | R86A | 23.99 | -42.89 | 37.87 | 2.30 | 21.28 | -6.15 | 15.13 | 4.56 | |
| Z | L128A | 18.35 | 0.46 | -3.64 | 0.29 | 15.47 | -1.59 | 13.88 | 0.92 | |
| Z | K131A | 14.59 | 48.53 | -44.10 | 3.43 | 22.45 | -9.82 | 12.62 | 3.30 | |
| Z | Q75A | 13.88 | 7.23 | -7.61 | 1.63 | 15.13 | -3.68 | 11.45 | 5.23 | |
| Z | I126A | 13.54 | -0.54 | -1.34 | -0.13 | 11.49 | -1.25 | 10.24 | 1.21 | |
| Z | K78A | 10.12 | -31.35 | 35.74 | -0.33 | 14.21 | -4.93 | 9.24 | 0.42 | |
| Z | V64A | 8.53 | 0.42 | -0.29 | -0.25 | 8.36 | -0.54 | 7.82 | 1.0 | |
| Z | N66A | 7.27 | 5.69 | -3.64 | -0.29 | 8.99 | -1.38 | 7.65 | 1.46 | |
| Z | Q88A | 8.65 | -6.40 | 6.10 | -0.04 | 8.28 | -1.09 | 7.19 | 1.09 | |
| Z | Y68A | 7.48 | 0.54 | -1.17 | 0.08 | 6.90 | -0.50 | 6.40 | 1.08 | |
| Pembrolizumab | A | F103A | 27.04 | 1.63 | -2.34 | 0.50 | 26.84 | -2.22 | 24.62 | 1.13 |
| A | Y33A | 18.77 | 11.24 | -5.81 | 0.29 | 24.49 | -1.84 | 22.66 | 0.42 | |
| A | R102A | 24.24 | -17.77 | 14.88 | 1.88 | 23.24 | -5.48 | 17.77 | 2.88 | |
| A | M105A | 17.77 | 4.51 | -6.90 | 0.21 | 15.55 | -3.01 | 12.54 | 4.51 | |
| B | Y34A | 15.72 | 7.19 | -10.62 | 1.88 | 14.21 | -2.72 | 11.49 | 0.88 | |
| A | N59A | 12.58 | 8.61 | -9.03 | -0.04 | 12.12 | -1.97 | 10.16 | 0.88 | |
| A | Y101A | 13.46 | 4.35 | -7.48 | 2.01 | 12.33 | -2.30 | 10.03 | 2.22 | |
| B | Y57A | 11.16 | 5.10 | -6.31 | 1.38 | 11.37 | -2.51 | 8.86 | 2.59 | |
| B | Y36A | 8.53 | 14.96 | -13.67 | 0.21 | 10.03 | -2.88 | 7.15 | 4.22 | |
| B | Y53A | 7.48 | 14.59 | -12.83 | 0.25 | 9.49 | -3.22 | 6.27 | 1.59 | |
| A | Y35A | 5.27 | 11.66 | -8.19 | -0.50 | 8.23 | -2.01 | 6.19 | 0.50 | |
| B | R96A | 10.58 | -30.47 | 27.17 | 0.92 | 8.19 | -2.84 | 5.39 | 2.97 | |
| A | D100A | 3.51 | 28.01 | -24.87 | 0.08 | 6.69 | -1.34 | 5.39 | 0.96 | |
| A | N52A | 5.89 | 9.74 | -9.49 | 0.04 | 6.23 | -1.51 | 4.72 | 2.21 |
Table 1 Computational alanine scanning result for the PD-1/pembrolizumab system
| Species | Chain | Mutation | ΔΔEvdW/(kJ·mol-1) | ΔΔEele/ (kJ·mol-1) | ΔΔGgb/ (kJ·mol-1) | ΔΔGnp/(kJ·mol-1) | ΔΔH/ (kJ·mol-1) | -TΔΔS/(kJ·mol-1) | ΔΔG/ (kJ·mol-1) | SD* |
|---|---|---|---|---|---|---|---|---|---|---|
| PD?1 | Z | D85A | -3.13 | 85.36 | -56.39 | -0.04 | 25.79 | -9.24 | 16.55 | 1.30 |
| Z | R86A | 23.99 | -42.89 | 37.87 | 2.30 | 21.28 | -6.15 | 15.13 | 4.56 | |
| Z | L128A | 18.35 | 0.46 | -3.64 | 0.29 | 15.47 | -1.59 | 13.88 | 0.92 | |
| Z | K131A | 14.59 | 48.53 | -44.10 | 3.43 | 22.45 | -9.82 | 12.62 | 3.30 | |
| Z | Q75A | 13.88 | 7.23 | -7.61 | 1.63 | 15.13 | -3.68 | 11.45 | 5.23 | |
| Z | I126A | 13.54 | -0.54 | -1.34 | -0.13 | 11.49 | -1.25 | 10.24 | 1.21 | |
| Z | K78A | 10.12 | -31.35 | 35.74 | -0.33 | 14.21 | -4.93 | 9.24 | 0.42 | |
| Z | V64A | 8.53 | 0.42 | -0.29 | -0.25 | 8.36 | -0.54 | 7.82 | 1.0 | |
| Z | N66A | 7.27 | 5.69 | -3.64 | -0.29 | 8.99 | -1.38 | 7.65 | 1.46 | |
| Z | Q88A | 8.65 | -6.40 | 6.10 | -0.04 | 8.28 | -1.09 | 7.19 | 1.09 | |
| Z | Y68A | 7.48 | 0.54 | -1.17 | 0.08 | 6.90 | -0.50 | 6.40 | 1.08 | |
| Pembrolizumab | A | F103A | 27.04 | 1.63 | -2.34 | 0.50 | 26.84 | -2.22 | 24.62 | 1.13 |
| A | Y33A | 18.77 | 11.24 | -5.81 | 0.29 | 24.49 | -1.84 | 22.66 | 0.42 | |
| A | R102A | 24.24 | -17.77 | 14.88 | 1.88 | 23.24 | -5.48 | 17.77 | 2.88 | |
| A | M105A | 17.77 | 4.51 | -6.90 | 0.21 | 15.55 | -3.01 | 12.54 | 4.51 | |
| B | Y34A | 15.72 | 7.19 | -10.62 | 1.88 | 14.21 | -2.72 | 11.49 | 0.88 | |
| A | N59A | 12.58 | 8.61 | -9.03 | -0.04 | 12.12 | -1.97 | 10.16 | 0.88 | |
| A | Y101A | 13.46 | 4.35 | -7.48 | 2.01 | 12.33 | -2.30 | 10.03 | 2.22 | |
| B | Y57A | 11.16 | 5.10 | -6.31 | 1.38 | 11.37 | -2.51 | 8.86 | 2.59 | |
| B | Y36A | 8.53 | 14.96 | -13.67 | 0.21 | 10.03 | -2.88 | 7.15 | 4.22 | |
| B | Y53A | 7.48 | 14.59 | -12.83 | 0.25 | 9.49 | -3.22 | 6.27 | 1.59 | |
| A | Y35A | 5.27 | 11.66 | -8.19 | -0.50 | 8.23 | -2.01 | 6.19 | 0.50 | |
| B | R96A | 10.58 | -30.47 | 27.17 | 0.92 | 8.19 | -2.84 | 5.39 | 2.97 | |
| A | D100A | 3.51 | 28.01 | -24.87 | 0.08 | 6.69 | -1.34 | 5.39 | 0.96 | |
| A | N52A | 5.89 | 9.74 | -9.49 | 0.04 | 6.23 | -1.51 | 4.72 | 2.21 |
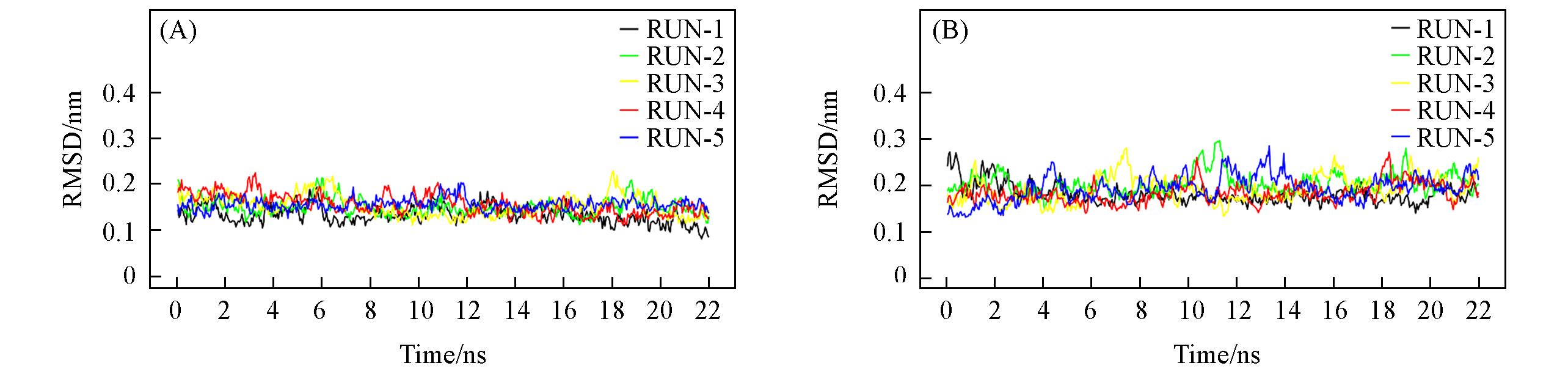
Fig.1 RMSDs of the heavy atoms in backbone of the PD?1/pembrolizumab(A) and PD?1/nivolumab(B) complexes relative to the initial crystal structures5 Trajectories were ran with 22 ns each for a total of 110 ns for each system.
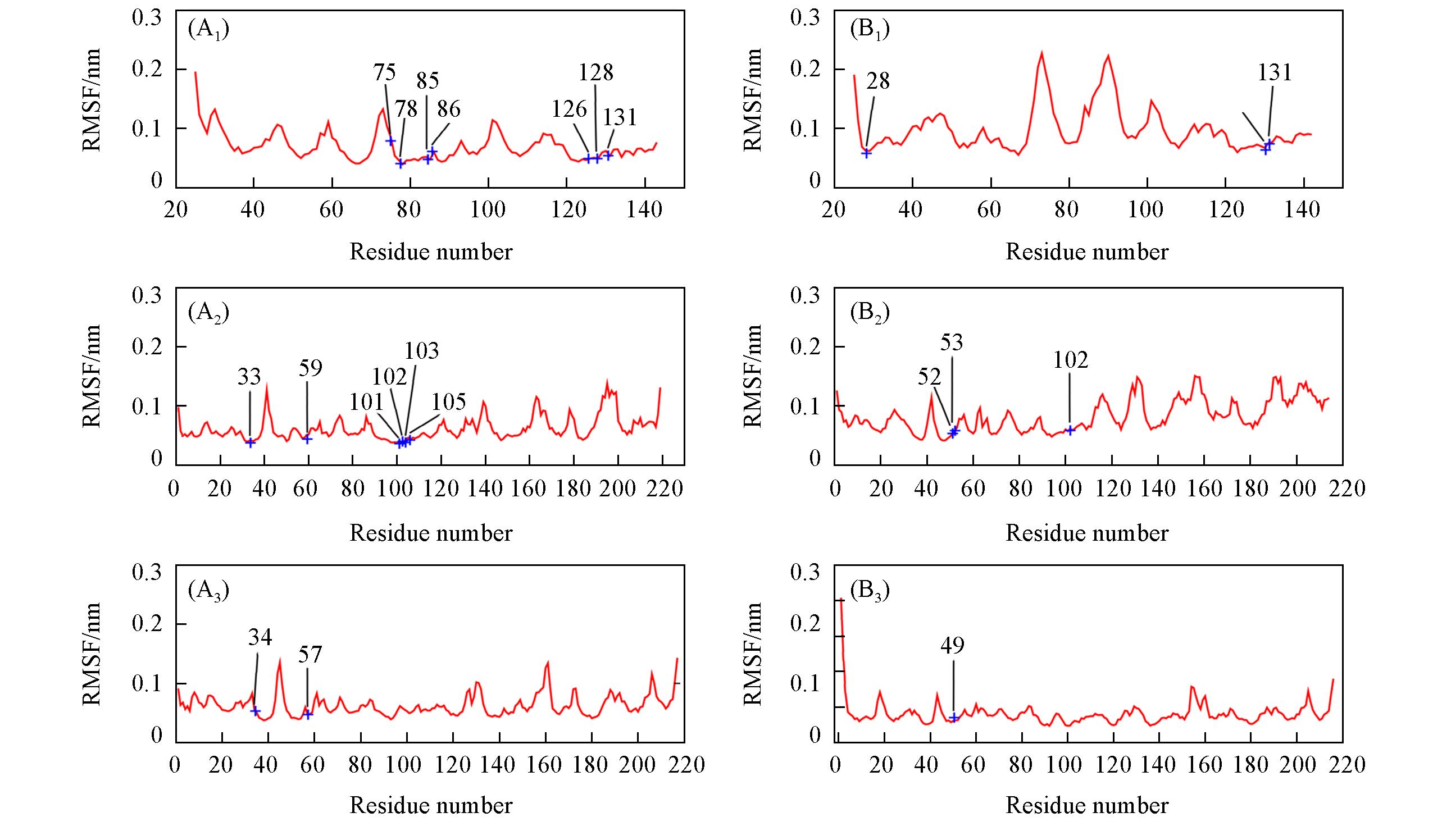
Fig.2 RMSFs of the Cα atoms calculated from the 22 ns trajectory of PD?1/pembrolizumab(A1—A3) and PD?1/nivolumab(B1—B3) trajectories(A1) PD-1; (A2) pembrolizumab-A; (A3) pembrolizumab-B; (B1) PD-1; (B2) nivolumab-H; (B3) nivolumab-L.The hot spots predicted by MM/GB/ASIE method are labeled in blue.
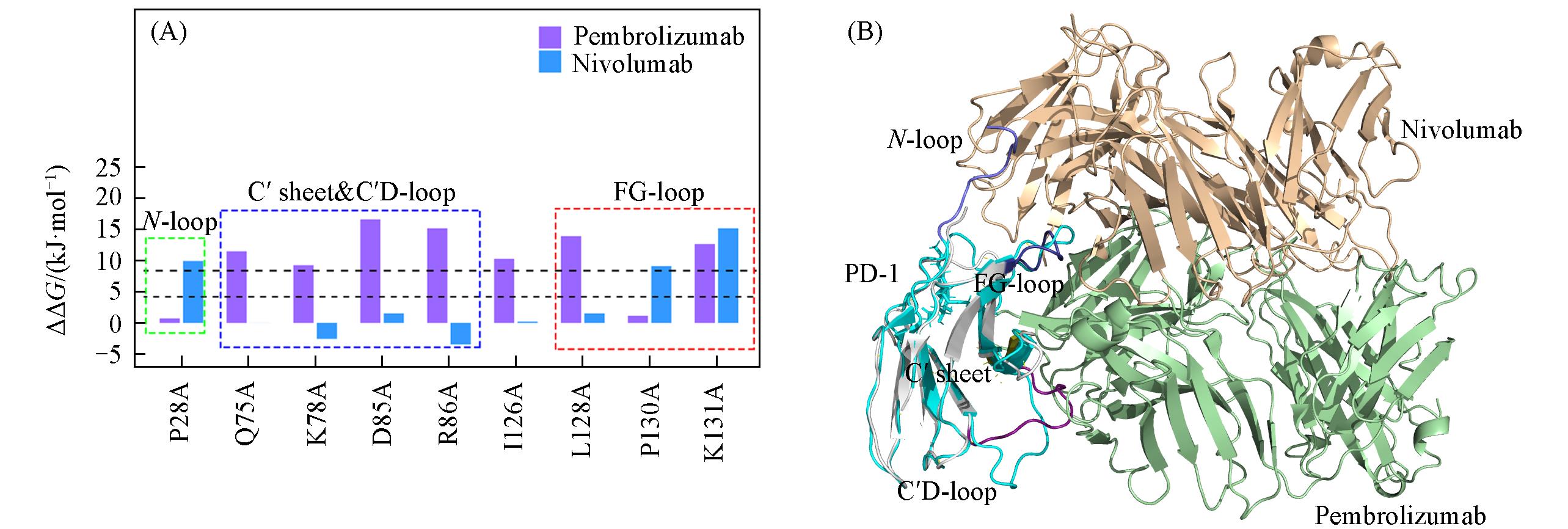
Fig.3 Hot spots predicted on PD?1 in two PD?1/mAb systems compared and arranged according to the sequence of residues on PD?1(A) and the superposition diagram of two PD?1/mAb complexes(PD?1/pembrolizumab and PD?1/nivolumab)(B)(A) The overlapped binding region(FG-loop in PD-1) with pembrolizumab and nivolumab is circled by the red dotted line; the blue dotted line circles the C′ sheet and C′D-loop regions that specifically bind to pembrolizumab; the red dotted line circles the N-loop region that specifically binds to nivolumab. The black horizontal dotted line indicates the distinguishing value between hotspot and warm spot. (B) PD-1(except C′D-loop, C′sheet and FG-loop) in PD-1/pembrolizumab is in white and PD-1 in PD-1/nivolumab is in cyan. The heavy chain and light chain of pembrolizumab are palegreen cartoons and those of nivolumab are in wheat. C′D-loop, C′sheet and FG-loop on PD-1 in PD-1/pembrolizumab complex are in deeppurple, deepolive and deepblue, respectively. N-loop on PD-l in PD-1/nivolumab is in light blue. The color of tags in these regions is the same as that of residues in cartoon form.
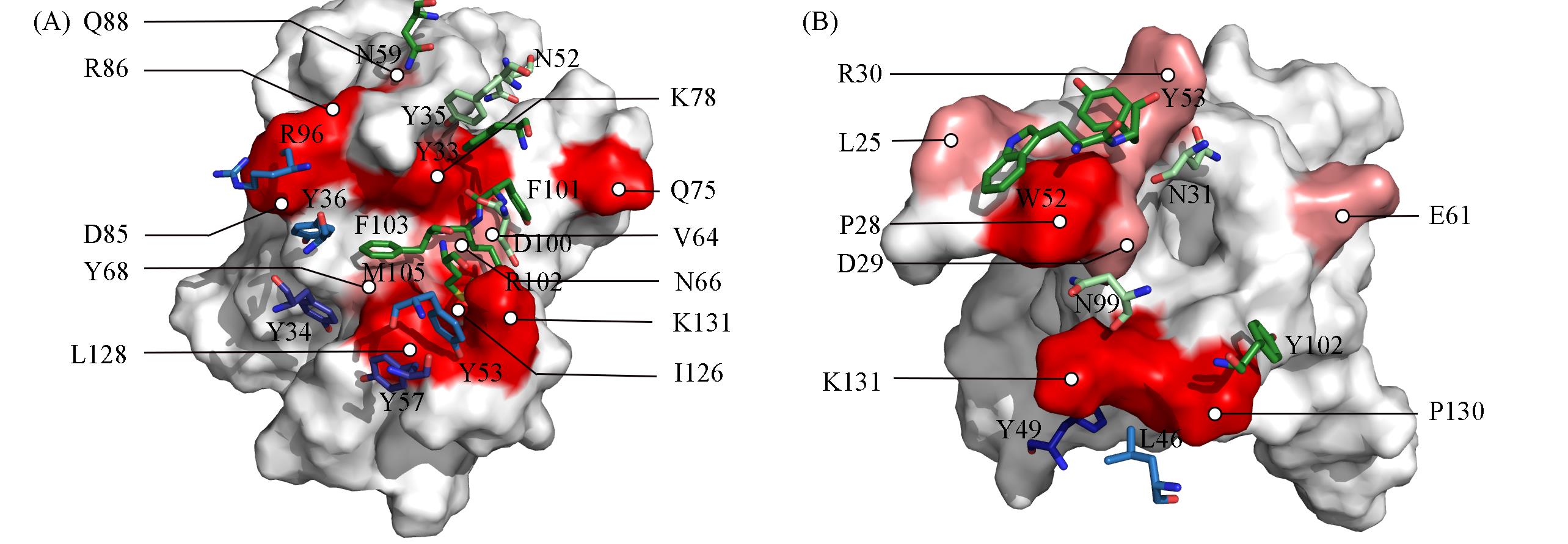
Fig.4 Details of the interaction of hotspots and warm spots at the PD?1/pembrolizumab(A) and PD?1/nivolumab(B) interfaces(A) Predicted spots are colored according to their location: on PD-1, red; on heavy chain of mAb, green; on light chain of mAb, cyan. Predicted residues are also colored by ΔΔG values: darker color for hot spots(typically defined as > 8.37 kJ/mol); lighter color for warm spots, 4.18 kJ/mol to 8.37 kJ/mol. For clarity, the surface of PD-1 is demonstrated and that of mAb is hidden. The PD-L1 residues are labeled in black font and are colored on the surface, and the mAb residues are labeled in white font and are represented as sticks. The atom coloring style is red for oxygen and blue for nitrogen.
| Species | Chain | Mutation | ??EvdW/(kJ·mol-1) | ??Eele/ (kJ·mol-1) | ??Ggb/ (kJ·mol-1) | ??Gnp/ (kJ·mol-1) | ??H/ (kJ·mol-1) | -T??S/(kJ·mol-1) | ??G/ (kJ·mol-1) | SD* |
|---|---|---|---|---|---|---|---|---|---|---|
| PD?1 | G | K131A | 22.82 | 12.04 | -15.13 | 2.72 | 22.45 | -7.32 | 15.13 | 4.56 |
| G | P28A | 13.54 | 0.17 | -3.72 | 0.67 | 10.62 | -0.75 | 9.91 | 1.34 | |
| G | P130A | 11.70 | 1.76 | -3.68 | 0.50 | 10.28 | -1.21 | 9.07 | 1.71 | |
| G | R30A | 14.38 | -34.78 | 31.89 | 1.25 | 12.75 | -4.51 | 8.23 | 3.47 | |
| G | L25A | 14.88 | 1.42 | -9.28 | 2.13 | 9.15 | -1.17 | 7.98 | 1.80 | |
| G | D29A | 6.35 | 60.44 | -54.72 | 0.46 | 12.54 | -6.48 | 6.06 | 1.42 | |
| G | E61A | 5.56 | 74.74 | -71.19 | 0.75 | 9.86 | -4.93 | 4.93 | 3.80 | |
| Nivolumab | H | Y53A | 14.96 | 3.85 | -4.77 | 1.55 | 15.59 | -1.63 | 13.96 | 0.67 |
| L | Y49A | 13.71 | 6.27 | -7.98 | 1.42 | 13.42 | -2.05 | 11.33 | 1.80 | |
| H | Y102A | 11.12 | 2.51 | -3.97 | 1.46 | 11.12 | -0.92 | 10.20 | 4.60 | |
| H | W52 | 24.79 | 6.27 | -14.92 | 1.30 | 21.61 | -7.65 | 9.78 | 2.55 | |
| L | L46A | 9.82 | -1.59 | 0.08 | -0.42 | 7.90 | -0.71 | 7.23 | 1.05 | |
| H | N99A | 6.98 | 25.67 | -20.73 | 0.59 | 12.54 | -5.48 | 7.02 | 2.04 | |
| H | N31A | 7.48 | 22.20 | -20.15 | 0.79 | 10.32 | -4.68 | 5.64 | 0.71 |
Table 2 Computational alanine scanning result for the PD-1/nivolumab system
| Species | Chain | Mutation | ??EvdW/(kJ·mol-1) | ??Eele/ (kJ·mol-1) | ??Ggb/ (kJ·mol-1) | ??Gnp/ (kJ·mol-1) | ??H/ (kJ·mol-1) | -T??S/(kJ·mol-1) | ??G/ (kJ·mol-1) | SD* |
|---|---|---|---|---|---|---|---|---|---|---|
| PD?1 | G | K131A | 22.82 | 12.04 | -15.13 | 2.72 | 22.45 | -7.32 | 15.13 | 4.56 |
| G | P28A | 13.54 | 0.17 | -3.72 | 0.67 | 10.62 | -0.75 | 9.91 | 1.34 | |
| G | P130A | 11.70 | 1.76 | -3.68 | 0.50 | 10.28 | -1.21 | 9.07 | 1.71 | |
| G | R30A | 14.38 | -34.78 | 31.89 | 1.25 | 12.75 | -4.51 | 8.23 | 3.47 | |
| G | L25A | 14.88 | 1.42 | -9.28 | 2.13 | 9.15 | -1.17 | 7.98 | 1.80 | |
| G | D29A | 6.35 | 60.44 | -54.72 | 0.46 | 12.54 | -6.48 | 6.06 | 1.42 | |
| G | E61A | 5.56 | 74.74 | -71.19 | 0.75 | 9.86 | -4.93 | 4.93 | 3.80 | |
| Nivolumab | H | Y53A | 14.96 | 3.85 | -4.77 | 1.55 | 15.59 | -1.63 | 13.96 | 0.67 |
| L | Y49A | 13.71 | 6.27 | -7.98 | 1.42 | 13.42 | -2.05 | 11.33 | 1.80 | |
| H | Y102A | 11.12 | 2.51 | -3.97 | 1.46 | 11.12 | -0.92 | 10.20 | 4.60 | |
| H | W52 | 24.79 | 6.27 | -14.92 | 1.30 | 21.61 | -7.65 | 9.78 | 2.55 | |
| L | L46A | 9.82 | -1.59 | 0.08 | -0.42 | 7.90 | -0.71 | 7.23 | 1.05 | |
| H | N99A | 6.98 | 25.67 | -20.73 | 0.59 | 12.54 | -5.48 | 7.02 | 2.04 | |
| H | N31A | 7.48 | 22.20 | -20.15 | 0.79 | 10.32 | -4.68 | 5.64 | 0.71 |
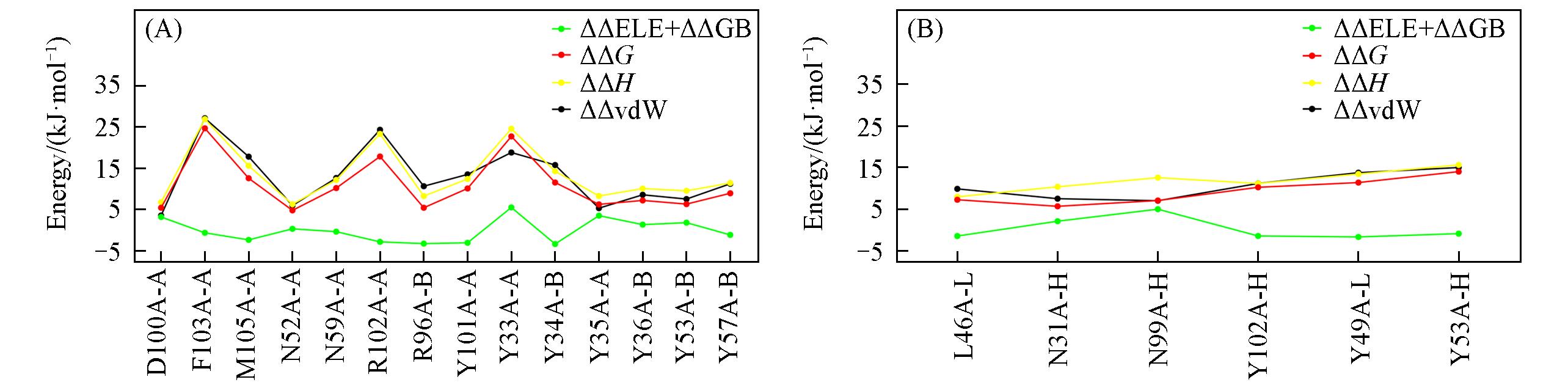
Fig.5 Composition of free energy contribution for hotspots and warm spots predicted by MM/GB/ASIE method(A) Pembrolizumab; (B) nivolumab. Hydrophobic item(vdW) is plotted in black and polar item(ELE+GB) is plotted in green. Enthalpy item and entropy item are plotted in yellow and red, respectively.
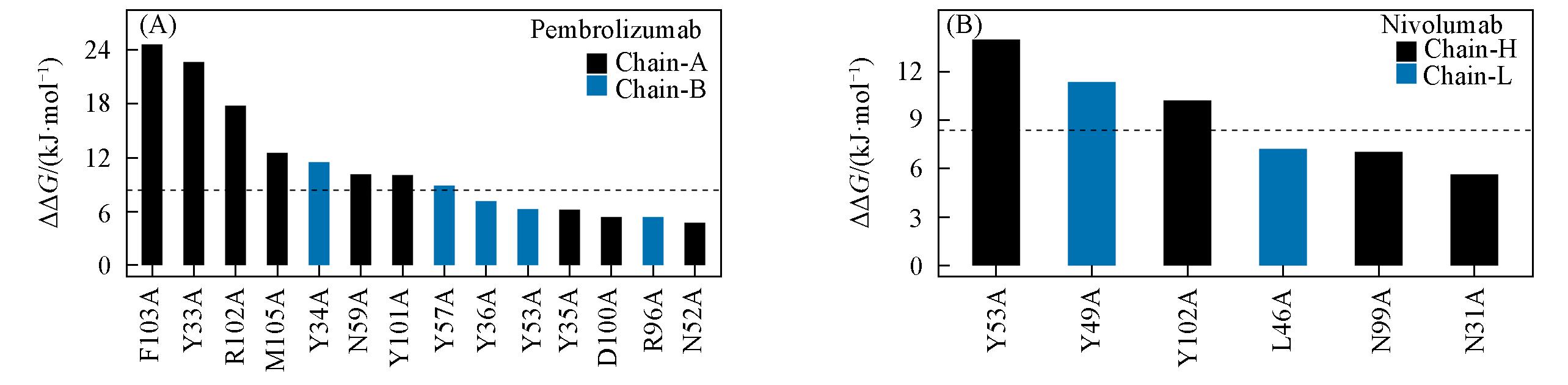
Fig.6 Hot and warm spots predicted on the PD?1/pembrolizumab(A) and PD?1/nivolumab(B)The residues on heavy chain of mAbs are in black, and residues on light chain of mAbs are in blue. The black horizontal dotted line indicates the value of hotspot.
| Species | Residue | Interaction | Spot type(mAb) | |
|---|---|---|---|---|
| PD?1 | mAb | |||
| PD?1/pembrolizumab | PD?1K131 | Y57 | cation?π(vdW) | Hot |
| PD?1K131 | Y53 | cation?π(vdW) | Warm | |
| PD?1K131 | D110 | Electrostatic | Warm | |
| PD?1/nivolumab | PD?1K131 | LY49 | cation?π(vdW) | Hot |
| PD?1K131 | HY102 | cation?π(vdW) | Hot | |
Table 3 Residues on the mAbs interacting with the only overlapping hotspots PD-1K131 and their main interaction types and the spot types of themselves
| Species | Residue | Interaction | Spot type(mAb) | |
|---|---|---|---|---|
| PD?1 | mAb | |||
| PD?1/pembrolizumab | PD?1K131 | Y57 | cation?π(vdW) | Hot |
| PD?1K131 | Y53 | cation?π(vdW) | Warm | |
| PD?1K131 | D110 | Electrostatic | Warm | |
| PD?1/nivolumab | PD?1K131 | LY49 | cation?π(vdW) | Hot |
| PD?1K131 | HY102 | cation?π(vdW) | Hot | |
| 1 | Nouri Rouzbahani F., Shirkhoda M., Memari F., Dana H., Mahmoodi Chalbatani G., Mahmoodzadeh H., Samarghandi N., Gharagozlou E., Mohammadi Hadloo M. H., Maleki A. R., Sadeghian E., Nia E., Nia N., Hadjilooei F., Rezaeian O., Meghdadi S., Miri S., Jafari F., Rayzan E., Marmari V., Pak. J. Biol. Sci.,2018, 21(3), 135—150 |
| 2 | Freeman G. J., Long A. J., Iwai Y., Bourque K., Chernova T., Nishimura H., Fitz L. J., Malenkovich N., Okazaki T., Byrne M. C., Horton H. F., Fouser L., Carter L., Ling V., Bowman M. R., Carreno B. M., Collins M., Wood C. R., Honjo T., J. Exp. Med.,2000, 192(7), 1027—1034 |
| 3 | Francisco L. M., Salinas V. H., Brown K. E., Vanguri V. K., Freeman G. J., Kuchroo V. K., Sharpe A. H., J. Exp. Med.,2009, 206(13), 3015—3029 |
| 4 | Zang X., Allison J. P., Clin. Cancer Res.,2007, 13(18 Pt 1), 5271—5279 |
| 5 | Pardoll D. M., Nat. Rev. Cancer,2012, 12(4), 252—264 |
| 6 | Akbay E. A., Koyama S., Carretero J., Altabef A., Tchaicha J. H., Christensen C. L., Mikse O. R., Cherniack A. D., Beauchamp E. M., Pugh T. J., Wilkerson M. D., Fecci P. E., Butaney M., Reibel J. B., Soucheray M., Cohoon T. J., Janne P. A., Meyerson M., Hayes D. N., Shapiro G. I., Shimamura T., Sholl L. M., Rodig S. J., Freeman G. J., Hammerman P. S., Dranoff G., Wong K. K., Cancer Discov.,2013, 3(12), 1355—1363 |
| 7 | Shi L., Chen S., Yang L., Li Y., J. Hematol. Oncol.,2013, 6(1), 74 |
| 8 | Zhou G. W., Xiong Y., Chen S., Xia F., Li Q., Hu J., Medicine(Baltimore),2016, 95(35), e4611 |
| 9 | Mariathasan S., Turley S. J., Nickles D., Castiglioni A., Yuen K., Wang Y., Kadel E. E. III, Koeppen H., Astarita J. L., Cubas R., Jhunjhunwala S., Banchereau R., Yang Y., Guan Y., Chalouni C., Ziai J., Senbabaoglu Y., Santoro S., Sheinson D., Hung J., Giltnane J. M., Pierce A. A., Mesh K., Lianoglou S., Riegler J., Carano R. A. D., Eriksson P., Hoglund M., Somarriba L., Halligan D. L., van der Heijden M. S., Loriot Y., Rosenberg J. E., Fong L., Mellman I., Chen D. S., Green M., Derleth C., Fine G. D., Hegde P. S., Bourgon R., Powles T., Nature,2018, 554(7693), 544—548 |
| 10 | Garon E. B., Rizvi N. A., Hui R., Leighl N., Balmanoukian A. S., Eder J. P., Patnaik A., Aggarwal C., Gubens M., Horn L., Carcereny E., Ahn M. J., Felip E., Lee J. S., Hellmann M. D., Hamid O., Goldman J. W., Soria J. C., Dolled⁃Filhart M., Rutledge R. Z., Zhang J., Lunceford J. K., Rangwala R., Lubiniecki G. M., Roach C., Emancipator K., Gandhi L., N. Engl. J. Med.,2015, 372(21), 2018—2028 |
| 11 | Chen L., Han X., J. Clin. Invest.,2015, 125(9), 3384—3391 |
| 12 | Li Y., Li F., Jiang F., Lv X., Zhang R., Lu A., Zhang G., Int. J. Mol. Sci.,2016, 17(7), 1151—1172 |
| 13 | Syn N. L., Teng M. W. L., Mok T. S. K., Soo R. A., Lancet Oncol.,2017, 18(12), e731—e741 |
| 14 | Callahan M. K., Postow M. A., Wolchok J. D., Immunity,2016, 44(5), 1069—1078 |
| 15 | Lee J. Y., Lee H. T., Shin W., Chae J., Choi J., Kim S. H., Lim H., Won Heo T., Park K. Y., Lee Y. J., Ryu S. E., Son J. Y., Lee J. U., Heo Y. S., Nat. Commun.,2016, 7, 13354 |
| 16 | Tan S., Zhang H., Chai Y., Song H., Tong Z., Wang Q., Qi J., Wong G., Zhu X., Liu W. J., Gao S., Wang Z., Shi Y., Yang F., Gao G. F., Yan J., Nat. Commun.,2017, 8, 14369 |
| 17 | Huang D., Wen W., Liu X., Li Y., Zhang J. Z. H., RSC Advances,2019, 9(26), 14944—14956 |
| 18 | Zhan M. M., Hu X. Q., Liu X. X., Ruan B. F., Xu J., Liao C., Drug Discov. Today,2016, 21(6), 1027—1036 |
| 19 | Adams J. L., Smothers J., Srinivasan R., Hoos A., Nat. Rev. Drug Discov.,2015, 14(9), 603—622 |
| 20 | Berman H., Henrick K., Nakamura H., Nat. Struct. Biol.,2003, 10(12), 980 |
| 21 | Waterhouse A., Bertoni M., Bienert S., Studer G., Tauriello G., Gumienny R., Heer F. T., de Beer T. A. P., Rempfer C., Bordoli L., Lepore R., Schwede T., Nucleic Acids Res.,2018, 46(W1), W296—W303 |
| 22 | Jorgensen W. L., Chandrasekhar J., Madura J. D., Impey R. W., Klein M. L., J. Chem. Phys.,1983, 79(2), 926—935 |
| 23 | Case V. B. D. A., Berryman J. T., Betz R. M., Cai Q., Cerutti D. S., Cheatham T. E., Darden T. A., Duke R. E., Gohlke H., Goetz A. W., Gusarov S., Homeyer N., Janowski P., Kaus J., Kolossváry I., Kovalenko A., Lee T. S., LeGrand S., Luchko T., Luo R., Madej B., Merz K. M., Paesani F., Roe D. R., Roitberg A., Sagui C., Salomon⁃Ferrer R., Seabra G., Simmerling C. L., Smith W., Swails J., Walker, Wang J., Wolf R. M., Wu X., Kollman P. A., AMBER14, University of California, San Francisco,2014 |
| 24 | Maier J. A., Martinez C., Kasavajhala K., Wickstrom L., Hauser K. E., Simmerling C., J. Chem. Theory Comput.,2015, 11(8), 3696—3713 |
| 25 | Darden T., York D., Pedersen L., J. Chem. Phys.,1993, 98(12), 10089—10092 |
| 26 | Ryckaert J. P., Ciccotti G., Berendsen H. J. C., J. Comput. Phys.,1977, 23(3), 327—341 |
| 27 | Kollman P.A., Massova I., Reyes C., Kuhn B., Huo S., Chong L., Lee M., Lee T., Duan Y., Wang W., Donini O., Cieplak P., Srinivasan J., Case D. A., Cheatham T. E., Acc. Chem. Res,2000, 33(12), 889—897 |
| 28 | Zwanzig R., J. Stat. Phys.,1973, 9(3), 215—220 |
| 29 | Berendsen H. J. C., Postma J. P. M., van Gunsteren W. F., DiNola A., Haak J. R., J. Chem. Phys.,1984, 81(8), 3684— 3690 |
| 30 | Qiu L., Yan Y., Sun Z., Song J., Zhang J. Z. H., Wires. Comput. Mol. Sci,2018, 8(2), e1342 |
| 31 | Duan L., Liu X., Zhang J. Z., J. Am. Chem. Soc.,2016, 138(17), 5722—5728 |
| 32 | Yan Y., Yang M., Ji C. G., Zhang J. Z. H., J. Chem. Inf. Model.,2017, 57(5), 1112—1122 |
| 33 | Huang D., Qi Y., Song J., Zhang J. Z. H., J. Comput. Chem.,2019, 40(9), 1045—1056 |
| 34 | Sitkoff D., Sharp K. A., Honig B., J. Phys. Chem.,2002, 98(7), 1978—1988 |
| [1] | 黄池宝, 张道海, 曾伯平, 刘其斌, 陈华仕, 康帅, 陈晓远. 用于糖链抗原CA19-9实时动态活检的免疫双光子荧光标记试剂盒[J]. 高等学校化学学报, 2016, 37(4): 638. |
| [2] | 范佳, 宋健, 毕丽荣, 周广宇, 张皓, 魏景艳, 杨柏. 量子点标记的斑点免疫渗滤分析定量检测cTnI[J]. 高等学校化学学报, 2009, 30(11): 2180. |
| [3] | 黎根,刘宁,刘志强,刘淑莹 . β2-微球蛋白连续表位的免疫亲和质谱研究[J]. 高等学校化学学报, 2007, 28(1): 92. |
| [4] | 李伟国, 祁超, 杜丽, 刘立岩, 赵大庆. 稀土离子对钙调蛋白与单克隆抗体分子识别的影响[J]. 高等学校化学学报, 2003, 24(8): 1346. |
| [5] | 李伟国, 祁超, 杜丽, 刘仔, 赵大庆. 稀土离子诱导钙调蛋白构象变化后的单克隆抗体制备[J]. 高等学校化学学报, 2002, 23(4): 526. |
| [6] | 祁超, 丁兰, 丁天兵, 刘志强, 张玉静, 刘立岩, 赵大庆. 抗meso-四(α,α,α,α-O-苯乙酰苯)卟啉抗体的酶学性质研究[J]. 高等学校化学学报, 2002, 23(4): 530. |
| [7] | 吴一卉, 马雪艳, 丁兰, 刘兰英, 赵大庆, 倪嘉缵. 具有cGPX活力的含硒抗体酶的酶学及动力学性质研究[J]. 高等学校化学学报, 2000, 21(11): 1621. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||