

高等学校化学学报 ›› 2018, Vol. 39 ›› Issue (2): 255.doi: 10.7503/cjcu20170586
收稿日期:2017-08-31
出版日期:2018-02-10
发布日期:2017-12-20
作者简介:联系人简介: 赵一雷, 男, 博士, 教授, 博士生导师, 主要从事生物分子结构计算和动力学研究. E-mail:基金资助:Received:2017-08-31
Online:2018-02-10
Published:2017-12-20
Contact:
ZHAO Yilei
E-mail:yileizhao@sjtu.edu.cn
Supported by:摘要:
运用生物信息学方法分析了各物种漆酶的耐盐性差异, 并利用随机加速动力学模拟计算揭示了白腐菌漆酶内部连接三铜活性中心(TNC)的氯离子、 氧气和水分子的可能输运通路. 生物信息学系统发生树和结构比对分析表明, 担子真菌、 子囊真菌与细菌漆酶具有较高的结构保守性; 通过随机加速动力学模拟发现, 白腐菌T. versicolor漆酶内部有5条小分子输运通道(p1~p5), 其中p2和p5为新的输运通道; 与氧气和水分子输运不同, 氯离子在漆酶内部输运时受到明显约束, 以较高的几率通过p1和p4输运通道. 高耐盐漆酶的p1通道周边富集了更多酸性和芳香性氨基酸残基, 降低了氯离子的输运效率, 从而提高其耐盐性.
中图分类号:
TrendMD:
李文娟, 赵一雷. 白腐菌漆酶耐盐性的生物信息学研究及氯离子、 氧气和水分子输运通道分析. 高等学校化学学报, 2018, 39(2): 255.
LI Wenjuan, ZHAO Yilei. Salt Tolerance of T. Versicolor Laccase: Bioinformatics Study and Internal Transportation of Chloride, Dioxygen, and Water†. Chem. J. Chinese Universities, 2018, 39(2): 255.
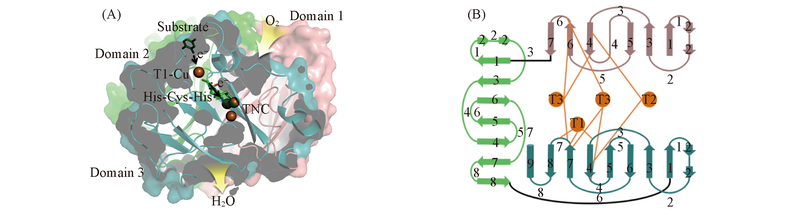
Scheme 1 Graphical representation of laccase structure with 3D surface model(A) and 2D topological diagram(B) Representative structure based on PDB# 1KYA in protein data bank, and the four copper atoms in orange and the three domains in salmon, green and teal, respectively.
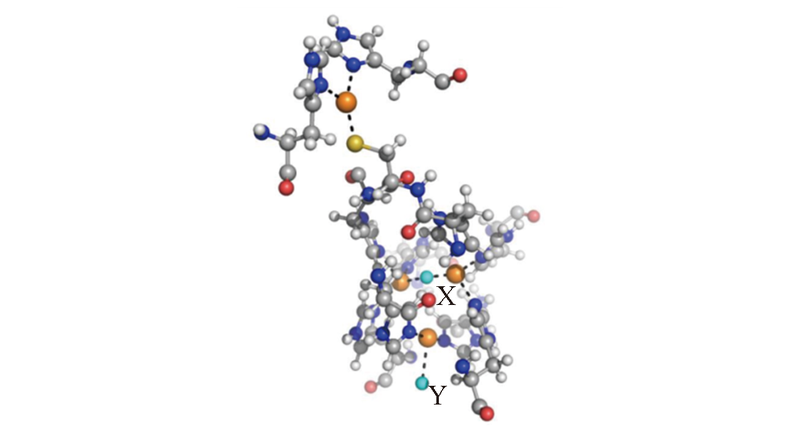
Fig.1 Active sites of single and tri-nuclear copper clustersX and Y were the positions of two oxygen atoms from dioxygen molecule, in which T3-Cu ligand X=OH-, O2 and Cl-, meanwhile T2-Cu ligand Y=OH-, H2O, and Cl-, respectively. The dashed lines denoted coordination to the copper cations.
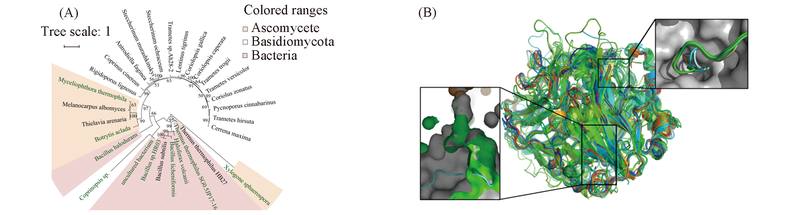
Fig.2 Phylogenetic tree of 29 laccase sequences based on sequence and structure alignment(A) and 26 superposed laccase structures with the POSA program(B) (A) Halotolerant laccases in green color; (B) the insets emphasize the comparison between T. versicolor(gray, from ascomycotas), basidio(green), and bacterial laccases(blue).
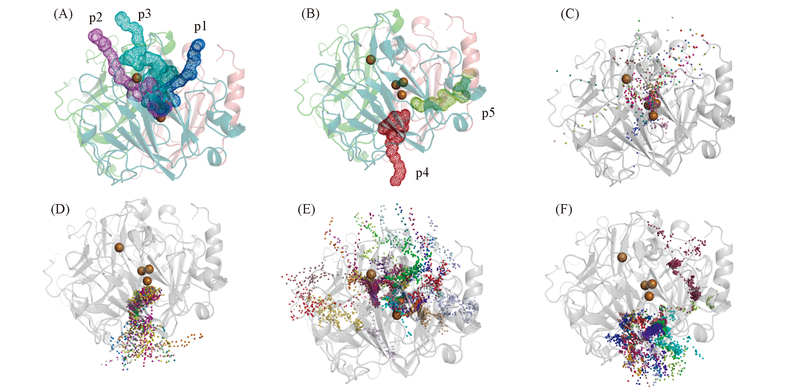
Fig.3 Transportation channels towards T3-Cu(A) and T2-Cu(B), and pathways exhibited with ligand snapshots in the RAMD trajectories of X-Cl-(C), Y-Cl-(D), X-O2(E) and Y-H2O(F) Each trajectory was shown in one color in Fig.3(C—F).
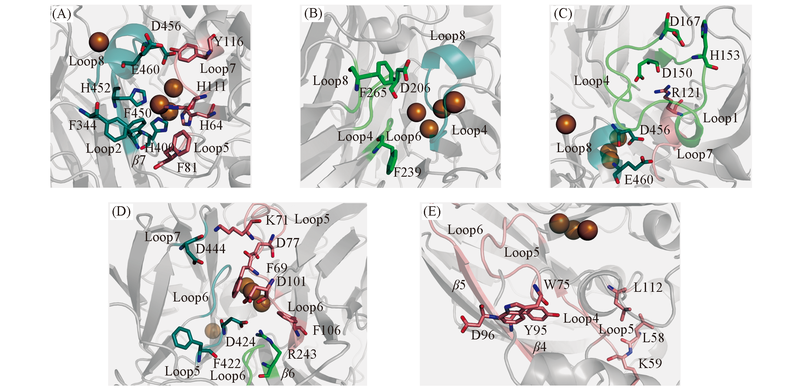
Fig.4 Constructions of tunnels p1(A), p2(B), p3(C), p4(D) and p5(E)The color-coding is same as Fig.1. The important residues surrounding the tunnel are shown as sticks.
| No. | Ligand | 1024 Accel./(kJ·mol-1· nm-1·g-1) | rmin/nm | Number of successful egress | Pathway | Frequency |
|---|---|---|---|---|---|---|
| Ⅰ | X=Cl- | 15.12 | 0.0120 | 18 | p1a | 12/18 |
| p2 | 3/18 | |||||
| p3 | 1/18 | |||||
| p4 | 1/18 | |||||
| Ⅱ | Y=Cl- | 7.56 | 0.0050 | 18 | p4 | 18/18 |
| Ⅱ | X=O2 | 5.29 | 0.0042 | 18 | p1a | 3/18 |
| p1b | 3/18 | |||||
| p1c | 1/18 | |||||
| p2 | 6/18 | |||||
| p3 | 1/18 | |||||
| Others | 4/18 | |||||
| Ⅳ | Y=H2O | 5.29 | 0.0042 | 13 | p4 | 11/13 |
| p5a | 1/13 | |||||
| p5b | 1/13 |
Table 1 Partition of internal transportation calculated with the RAMD simulations
| No. | Ligand | 1024 Accel./(kJ·mol-1· nm-1·g-1) | rmin/nm | Number of successful egress | Pathway | Frequency |
|---|---|---|---|---|---|---|
| Ⅰ | X=Cl- | 15.12 | 0.0120 | 18 | p1a | 12/18 |
| p2 | 3/18 | |||||
| p3 | 1/18 | |||||
| p4 | 1/18 | |||||
| Ⅱ | Y=Cl- | 7.56 | 0.0050 | 18 | p4 | 18/18 |
| Ⅱ | X=O2 | 5.29 | 0.0042 | 18 | p1a | 3/18 |
| p1b | 3/18 | |||||
| p1c | 1/18 | |||||
| p2 | 6/18 | |||||
| p3 | 1/18 | |||||
| Others | 4/18 | |||||
| Ⅳ | Y=H2O | 5.29 | 0.0042 | 13 | p4 | 11/13 |
| p5a | 1/13 | |||||
| p5b | 1/13 |
| [1] | Giardina P., Faraco V., Pezzella C., Piscitelli A., Vanhulle S., Sannia G., Cell Mol. Life Sci., 2010, 67(3), 369—385 |
| [2] | Riva S., Trends Biotechnol., 2006, 24(5), 219—226 |
| [3] | Jiao J., Yang X., Jin L. N., Gao J., Zhou Y., Xiao Y. Z., Zhang Y. J., Chem. J. Chinese Universities, 2016, 37(7), 1320—1327 |
| (焦晶, 杨雪, 金兰娜, 高键, 周洋, 肖亚中, 张应玖. 高等学校化学学报, 2016,37(7), 1320—1327) | |
| [4] | Dwivedi U. N., Singh P., Pandey V. P., Kumar A., J. Mol. Catal. B-Enzym., 2011, 68(2), 117—128 |
| [5] | Hakulinen N., Rouvinen J., Cell Mol. Life Sci., 2015, 72(5), 857—868 |
| [6] | Qi Y. B., Wang X. L., Shi T., Liu S. C., Xu Z. H., Li X. Q., Shi X. L., Xu P., Zhao Y. L., Phys. Chem. Chem. Phys., 2015, 17(44), 29597—29607 |
| [7] | Qi Y. B., Zhu J. R., Sun Y. J., Du Y., Chu J. J., Shi T., Zhao Y. L., Wang X. L., Chem. J. Chinese Universities, 2014, 35(4), 776—783 |
| (齐艳兵, 朱吉人, 孙尧金, 杜芸, 褚建君, 石婷, 赵一雷, 王晓雷. 高等学校化学学报, 2014,35(4), 776—783) | |
| [8] | Jones S. M., Solomon E. I., Cell Mol. Life Sci., 2015, 72(5), 869—883 |
| [9] | Piontek K., Antorini M., Choinowski T., J. Biol. Chem., 2002, 277(40), 37663—37669 |
| [10] | Bento I., Martins L. O., Gato Lopes G., Armenia Carrondo M., Lindley P. F., Dalton. Trans., 2005, 21(21), 3507—3513 |
| [11] | Lyashenko A. V., Bento I., Zaitsev V. N., Zhukhlistova N. E., Zhukova Y. N., Gabdoulkhakov A. G., Morgunova E. Y., Voelter W., Kachalova G. S., Stepanova E. V., Koroleva O. V., Lamzin V. S., Tishkov V. I., Betzel C., Lindley P. F., Mikhailov A. M., J. Biol. Inorg. Chem., 2006, 11(8), 963—973 |
| [12] | Kallio J. P., Rouvinen J., Kruus K., Hakulinen N., Biochemistry-US,2011, 50(21), 4396—4398 |
| [13] | Pietra F., Chem. Biodivers., 2016, 13(11), 1493—1501 |
| [14] | Damas J. M., Baptista A. M., Soares C. M., J. Chem. Theory Comput., 2014, 10(8), 3525—3531 |
| [15] | Farnet A. M., Gil G., Ferre E., Chemosphere,2008, 70(5), 895—900 |
| [16] | Pan T., Sun Y., Wang X. L., Shi T., Zhao Y. L., Chin. Chem. Lett., 2014, 25(7), 983—988 |
| [17] | Vaz-Dominguez C., Campuzano S., Rudiger O., Pita M., Gorbacheva M., Shleev S., Fernandez V. M., De Lacey A. L., Biosens. Bioelectron., 2008, 24(4), 531—537 |
| [18] | Kumar P., Prasad B., Mishra I. M., Chand S., J. Hazard. Mater., 2007, 149(1), 26—34 |
| [19] | Vishnu G., Palanisamy S., Joseph K., J. Clean. Prod., 2008, 16(10), 1081—1089 |
| [20] | Raseda N., Hong S., Kwon O., Ryu K., J. Microbiol. Biotechnol., 2014, 24(12),1673—1678 |
| [21] | Xu F., Biochemistry-US, 1996, 35(23), 7608—7614 |
| [22] | Robles A., Lucas R., Martí Nez-Cañamero M., Omar N. B., Pérez R., Gálvez A., Enzyme Microb. Technol., 2002, 31(4), 516—522 |
| [23] | Ruijssenaars H. J., Hartmans S., Appl. Microbiol. Biotechnol., 2004, 65(2), 177—182 |
| [24] | Jimenez-Juarez N., Roman-Miranda R., Baeza A., Sanchez-Amat A., Vazquez-Duhalt R., Valderrama B., J. Biotechnol., 2005, 117(1), 73—82 |
| [25] | Trovaslet M., Enaud E., Guiavarc’h Y., Corbisier A. M., Vanhulle S., Enzyme Microb. Technol., 2007, 41(3), 368—376 |
| [26] | Niladevi K. N., Jacob N., Prema P., Process Biochem., 2008, 43(6), 654—660 |
| [27] | Singh G., Sharma P., Capalash N., J. Gen. Appl. Microbiol., 2009, 55(4), 283—289 |
| [28] | Uthandi S., Saad B., Humbard M. A., Maupin-Furlow J. A., Appl. Environ. Microbiol., 2010, 76(3), 733—743 |
| [29] | Fang Z., Li T., Wang Q., Zhang X., Peng H., Fang W., Hong Y., Ge H., Xiao Y., Appl. Microbiol. Biotechnol., 2011, 89(4), 1103—1110 |
| [30] | Kittl R., Mueangtoom K., Gonaus C., Khazaneh S. T., Sygmund C., Haltrich D., Ludwig R., J. Biotechnol., 2012, 157(2), 304—314 |
| [31] | Qasemian L., Billette C., Guiral D., Alazard E., Moinard M., Farnet A. M., Fungal Biol., 2012, 116(10), 1090—1098 |
| [32] | Lu L., Wang T. N., Xu T. F., Wang J. Y., Wang C. L., Zhao M., Bioresour. Technol., 2013, 134, 81—86 |
| [33] | Brander S., Mikkelsen J. D., Kepp K. P., PLoS One, 2014, 9(6), e99402 |
| [34] | Dabirmanesh B., Khajeh K., Ghazi F., Ranjbar B., Etezad S. M., Int. J. Biol. Macromol., 2015, 79, 822—829 |
| [35] | Liu H., Cheng Y., Du B., Tong C., Liang S., Han S., Zheng S., Lin Y., PLoS One, 2015, 10(3), e0119833 |
| [36] | Imran M., Zhang B., Tang K., Liu J., Chem. Res. Chinese Universities, 2017, 33(1), 87—93 |
| [37] | Tamura K., Stecher G., Peterson D., Filipski A., Kumar S., Mol. Biol. Evol., 2013, 30(12), 2725—2729 |
| [38] | Li Z., Natarajan P., Ye Y., Hrabe T., Godzik A., Nucleic. Acids. Res., 2014, 42(W1), W240—W245 |
| [39] | Dolinsky T. J., Nielsen J. E., Mccammon J. A., Baker N. A., Nucleic. Acids. Res., 2004, 32(W1), W665—W667 |
| [40] | Jorgensen W. L., Chandrasekhar J., Madura J. D., Impey R. W., Klein M. L., J. Chem. Phys., 1983, 79(2), 926—935 |
| [41] | Case D.A., Darden T. A., Cheatham Ⅲ T. E., Simmerling C. L., Wang J., Duke R. E., Luo R., Walker R. C., Zhang W., Merz K. M., Roberts B., Hayik S., Roitberg A., Seabra G., Swails J., Götz A. W., Kolossváry I., Wong K. F., Paesani F., Vanicek J., Wolf R. M., Liu J., Wu X., Brozell S. R., Steinbrecher T., Gohlke H., Cai Q., Ye X., Wang J., Hsieh M. J., Cui G., Roe D. R., Mathews D. H., Seetin M. G., Salomon-Ferrer R., Sagui C., Babin V., Luchko T., Gusarov S., Kovalenko A., Kollman P. A., AMBER, Version 12, University of California,San Francisco, 2012 |
| [42] | Maestre-Reyna M., Liu W. C., Jeng W. Y., Lee C. C., Hsu C. A., Wen T. N., Wang A. H., Shyur L. F., PLoS One, 2015, 10(4), e0120601 |
| [43] | Wang J., Wolf R. M., Caldwell J. W., Kollman P. A., Case D. A., J. Comput. Chem., 2004, 25(9), 1157—1174 |
| [44] | Phillips J. C., Braun R., Wang W., Gumbart J., Tajkhorshid E., Villa E., Chipot C., Skeel R. D., Kale L., Schulten K., J. Comput. Chem., 2005, 26(16), 1781—1802 |
| [45] | Zhou C., Shi T., Zhao Y. L., Wang X. L., Chem. J. Chinese Universities, 2017, 38(10), 1813—1821 |
| (周超, 石婷, 赵一雷, 王晓雷. 高等学校化学学报, 2017,38(10), 1813—1821) | |
| [46] | Li W., Shen J., Liu G., Tang Y., Hoshino T., Proteins,2011, 79(1), 271—281 |
| [47] | Pavlova M., Klvana M., Prokop Z., Chaloupkova R., Banas P., Otyepka M., Wade R. C., Tsuda M., Nagata Y., Damborsky J., Nat. Chem. Biol., 2009, 5(10), 727—733 |
| [48] | Wang T., Duan Y., J. Am. Chem. Soc., 2007, 129(22), 6970—6971 |
| [49] | Kallio J., Auer S., Jänis J., Andberg M., Kruus K., Rouvinen J., Koivula A., Hakulinen N., J. Mol. Biol., 2009, 392(4), 895—909 |
| [50] | Solomon E.I., Augustine A. J., Yoon J.,Dalton. Trans., 2008, (30), 3921—3932 |
| [1] | 周宁, 唐小华, 曹红, 查飞, 李春, 谢春燕, 徐明平, 孙艺格. 石榴状凝胶微球固定化漆酶的制备、 表征及降解双酚A[J]. 高等学校化学学报, 2022, 43(2): 20210705. |
| [2] | 段秉怡, 王宇, 郭宁宁, 王润伟, 张宗弢, 裘式纶. 蛋黄-壳结构Fe3O4@SiO2@PMO磁性微球的制备及对漆酶的固定化[J]. 高等学校化学学报, 2019, 40(2): 210. |
| [3] | 焦晶, 杨雪, 金兰娜, 高键, 周洋, 肖亚中, 张应玖. 枯草杆菌漆酶重要功能位点的保守性与可变性分析[J]. 高等学校化学学报, 2016, 37(7): 1320. |
| [4] | 黎春怡, 黄卓烈, 王春晓. 壳寡糖化学修饰对云芝漆酶催化活性和酶学性质的影响[J]. 高等学校化学学报, 2015, 36(4): 704. |
| [5] | 齐艳兵, 朱吉人, 孙尧金, 杜芸, 褚建君, 石婷, 赵一雷, 王晓雷. 漆酶与酚类模式底物的结合及反应活性的理论研究[J]. 高等学校化学学报, 2014, 35(4): 776. |
| [6] | 王惠厦, 姚林, 丁彬, 罗健辉, 周歌, 江波. 疏水改性聚丙烯酰胺溶液的分子模拟[J]. 高等学校化学学报, 2013, 34(5): 1295. |
| [7] | 王苗苗, 李群艳, 韦奇, 聂祚仁. 介孔SiO2/Fe3O4中空磁性微球的漆酶固定化[J]. 高等学校化学学报, 2013, 34(2): 299. |
| [8] | 周海峰, 杨东杰, 伍晓蕾, 邱学青. 漆酶改性木质素磺酸钠的结构表征及吸附特征[J]. 高等学校化学学报, 2013, 34(1): 218. |
| [9] | 段淑娥, 翟云会, 屈颖娟, 马明阳. 银-组氨酸配合物-蒙脱石抗菌剂的制备及抗菌活性[J]. 高等学校化学学报, 2012, 33(12): 2617. |
| [10] | 兰甲宁, 纳薇, 韦奇, 李群艳, 王为, 聂祚仁. 环氧基修饰周期性介孔有机氧化硅对漆酶的固定化作用[J]. 高等学校化学学报, 2010, 31(8): 1579. |
| [11] | 黄俊, 王军涛, 李斌, 肖海燕, 刘诚, 丁莉芸, 姜德生 . 新型有机-无机纳米复合粒子的制备及其固定化漆酶研究[J]. 高等学校化学学报, 2006, 27(11): 2088. |
| [12] | 涂楚桥, 梁宏, 王光辉. 漆树漆酶的催化氧化作用 (ⅹⅢ)──Fe2+离子对漆酶催化氧化反应的影响[J]. 高等学校化学学报, 1999, 20(2): 243. |
| [13] | 吴兴国, 蔡汝秀, 黄厚评, 王鹏. 漆酶催化半醌自由基中间体动力学的模拟研究[J]. 高等学校化学学报, 1998, 19(8): 1223. |
| [14] | 望天志, 吴鼎泉, 万洪文, 屈松生, 杜予民. 微量热法研究漆酶和3,4,5-三羟基苯甲酸的反应[J]. 高等学校化学学报, 1997, 18(5): 786. |
| [15] | 熊亚, 谭智群, 刘义, 潘正君, 吴鼎泉, 康立山, 屈松生. 线粒体和亚线粒体体外代谢及漆酶影响的微量热研究[J]. 高等学校化学学报, 1997, 18(5): 753. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
