

Chem. J. Chinese Universities ›› 2018, Vol. 39 ›› Issue (11): 2395.doi: 10.7503/cjcu20180272
• Analytical Chemistry • Previous Articles Next Articles
LI Wenwen1,3, ZHU Airu2, LONG Yijing2, WANG Chunyan2, HAN Yuanping2, DUAN Yixiang2,3,*( )
)
Received:2018-04-09
Online:2018-11-10
Published:2018-10-16
Contact:
DUAN Yixiang
E-mail:yduan@scu.edu.cn
Supported by:CLC Number:
TrendMD:
LI Wenwen, ZHU Airu, LONG Yijing, WANG Chunyan, HAN Yuanping, DUAN Yixiang. Metabolomics Study of Serum and Liver in Type 2 Diabetes Mice Induced by High Fat Diet with Vitamin D Deficiency†[J]. Chem. J. Chinese Universities, 2018, 39(11): 2395.
| Parameter | Control(mean±SE) | HD(mean±SE) |
|---|---|---|
| Mass/g | 33.57±0.71 | 45.47±2.14* |
| Visceral fat coefficient(%) | 2.70±0.32 | 7.50±0.74* |
| FPB/(mmol·L-1) | 5.20±0.24 | 8.90±0.31* |
| HOMA-IR(fold) | 1.0 | 1.7* |
| Total cholesterol/(mmol·L-1) | 3.33±0.22 | 4.94±0.33* |
| Triglyceride/(mmol·L-1) | 0.63±0.05 | 0.90±0.07* |
| Free fatty acid/(mmol·L-1) | 0.89±0.04 | 1.19±0.03* |
| LDL-C/(mmol·L-1) | 0.42±0.04 | 0.78±0.11* |
| HDL-C/LDL-C | 7.14±0.48 | 5.33±0.38* |
Table 1 Biochemical parameters for HD group and control group
| Parameter | Control(mean±SE) | HD(mean±SE) |
|---|---|---|
| Mass/g | 33.57±0.71 | 45.47±2.14* |
| Visceral fat coefficient(%) | 2.70±0.32 | 7.50±0.74* |
| FPB/(mmol·L-1) | 5.20±0.24 | 8.90±0.31* |
| HOMA-IR(fold) | 1.0 | 1.7* |
| Total cholesterol/(mmol·L-1) | 3.33±0.22 | 4.94±0.33* |
| Triglyceride/(mmol·L-1) | 0.63±0.05 | 0.90±0.07* |
| Free fatty acid/(mmol·L-1) | 0.89±0.04 | 1.19±0.03* |
| LDL-C/(mmol·L-1) | 0.42±0.04 | 0.78±0.11* |
| HDL-C/LDL-C | 7.14±0.48 | 5.33±0.38* |
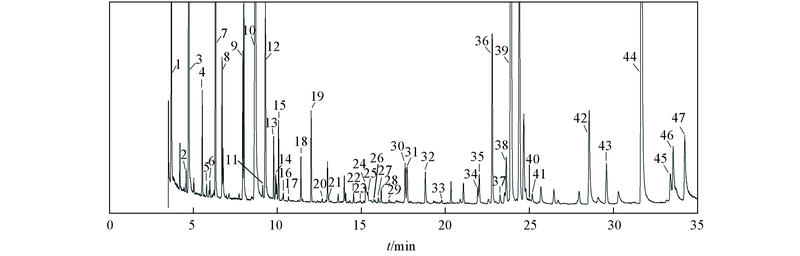
Fig.1 Total ion chromatogram of a serum samplePeak: 1. hexanol; 2. pyruvic acid**; 3. L-lactic acid**; 4. L-Alanine**; 5. hydroxylamine; 6. 2-hydroxybutyric acid*; 7. oxalic acid; 8. 3-hydroxybutyric acid; 9. L-valine; 10. urea**; 11. ethanol amine; 12. glycerol**; 13. L-isoleucine; 14. L-proline**; 15. glycine; 16. succinic acid*; 17. glyceric acid*; 18. L-serine**; 19. L-threonine**; 20. 2,4-dihydroxybutyric acid; 21. β-alanine; 22. malic acid; 23. erythritol; 24. L-methionine**; 25. 5-oxoproline; 26. erythronic acid; 27. creatinine*; 28. threonic acid**; 29. α-ketoglutarate; 30. L-glutamic acid*; 31. L-phenylalanine; 32. ribose; 33. ribitol; 34. L-ornithine; 35. citric acid; 36. 1-deoxyglucose*; 37. fructose**; 38. mannose; 39. glucose**; 40. glucitol; 41. L-tyrosine; 42. palmitic acid; 43. inositol*; 44. heptadecanoic acid; 45. linoleic acid; 46. oleic acid; 47. stearic acid. Metabolites confirmed with standards are in bold, statistical significance between control and HD group, * P<0.05, ** P<0.01.
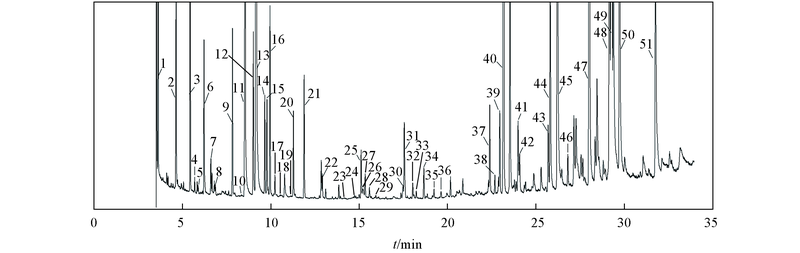
Fig.2 Total ion chromatogram of a liver samplePeak: 1. hexanol; 2. L-lactic acid*; 3. L-alanine; 4. hydroxylamine*; 5. 2-hydroxybutyric acid*; 6. oxalic acid; 7. 3-hydroxybutyric acid; 8. 2-aminobutyric acid*; 9. L-valine*; 10. 4-hydroxybutyric acid; 11. urea*; 12. ethanol amine**; 13. glycerol; 14. L-isoleucine*; 15. L-proline; 16. glycine; 17. succinic acid*; 18. glyceric acid*; 19. fumaric acid**; 20. L-serine; 21. L-threonine; 22. β-alanine**; 23. malic acid**; 24. erythritol**; 25. L-methionine*; 26. 5-oxoproline; 27. 4-aminobutyric acid; 28. 2,6-ditertbutylphenol; 29. cysteine; 30. L-glutamic acid; 31. L-phenylalanine; 32. xylose**; 33. lauric acid**; 34. ribose**; 35. xylitol; 36. ribitol; 37. myristic acid; 38. fructose**; 39. mannose**; 40. glucose**; 41. glucitol; 42. L-tyrosine; 43. palmitelaidic acid*; 44. palmitoleic acid; 45. palmitic acid**; 46. inositol; 47. heptadecanoic acid; 48. linoleic acid; 49. oleic acid*; 50. stearic acid; 51. arachidonic acid. Metabolites confirmed with standards are in bold, * P<0.05, ** P<0.01.
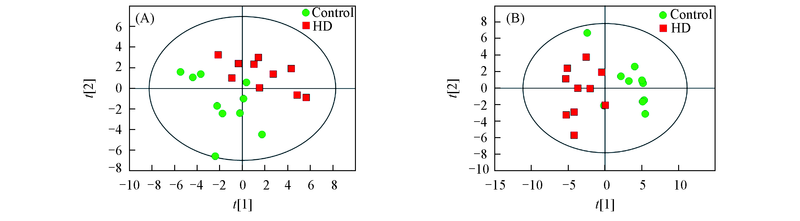
Fig.3 Principal component analysis score plots for metabolic profiling of serum(A) and liver(B) samples in HD group and control group(A) R2X[1]=0.14, R2X[2]=0.1; (B) R2X[1]=0.181, R2X[2]=0.0884.
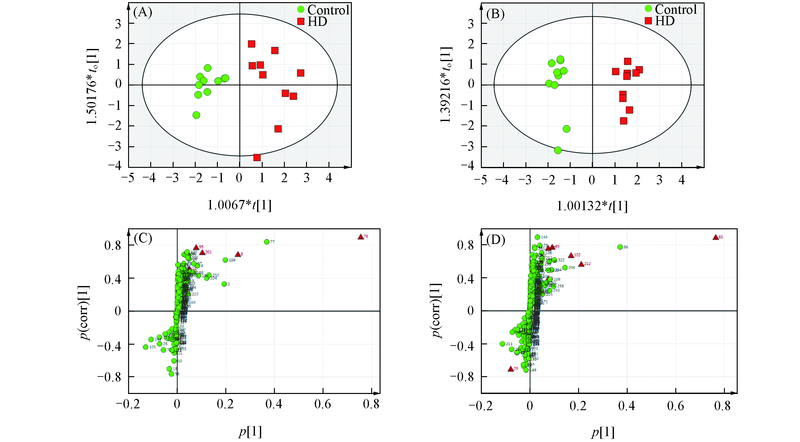
Fig.4 Orthogonal partial least squares-discriminant analysis(OPLS-DA) score plots for metabolic profiling of serum(A) and liver(B) samples in HD group and control group and S-plots of variablesin serum(C) and liver(D)(A) R2X[1]=0.35; R2X[X side comp. 1]=0.217; (B) R2X[1]=0.305; R2X[X side comp.1]=0.172; (C) R2X[1]=0.35; (D) R2X[1]=0.305. (C) and (D): red triangles are metabolic biomarkers with VIP>1 and statistical significance P<0.05.
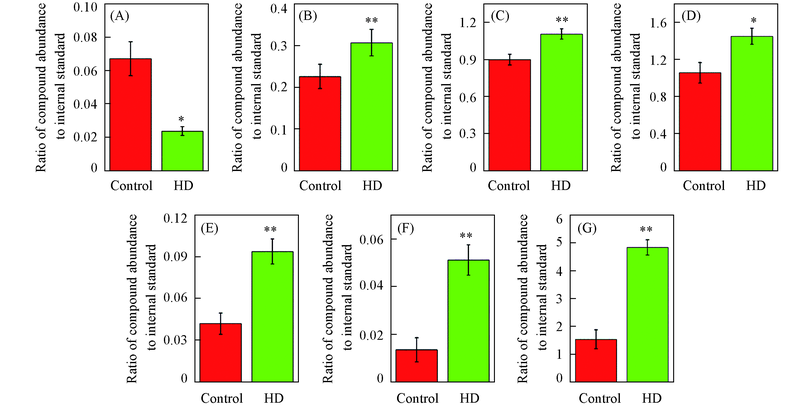
Fig.6 Liver metabolic biomarkers for T2DM(* P<0.05, ** P<0.01)(A) Ethanolamine; (B) lactic acid; (C) palmitic acid; (D) oleic acid; (E) ribose; (F) fructose; (G) glucose.
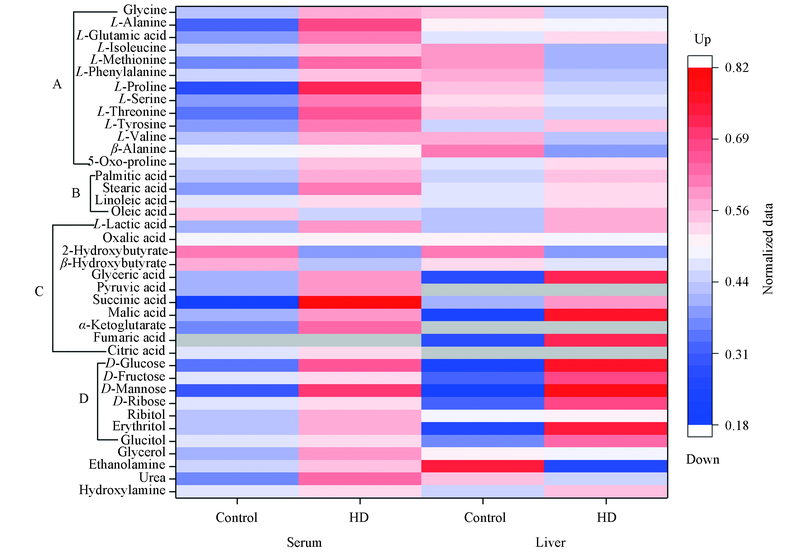
Fig.7 Heat map for serum and liver metabolites in HD group and control groupA, B, C and D represent amino acids, fatty acids, organic acids, and carbohydrates and alditol, respectively. Red color and blue color indicate up-regulated and down-regulated, grey color represents absence.
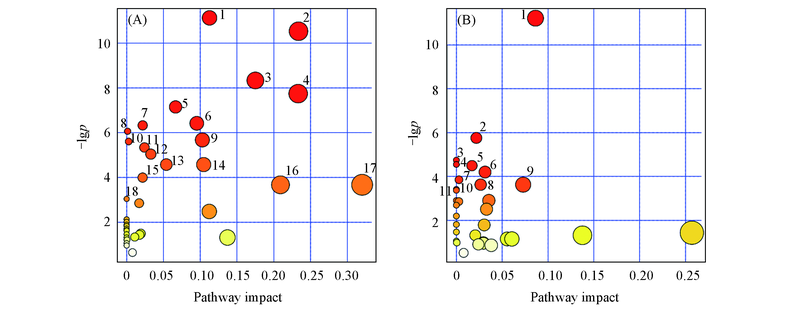
Fig.8 Metabo analyst pathway analysis for serum(A) and liver(B) metabolitesNumbers in the plots are metabolic pathways listed in Table 2 and Table 3, respectively.
| No. | Pathway name | P | Impact | KEGG |
|---|---|---|---|---|
| 1 | Aminoacyl-tRNA biosynthesis* | 0 | 0.113 | map00970 |
| 2 | Alanine, aspartate and glutamate metabolism* | 0 | 0.234 | map00250 |
| 3 | Arginine and proline metabolism* | 0 | 0.175 | map00330 |
| 4 | Glycine, serine and threonine metabolism* | 0 | 0.233 | map00260 |
| 5 | Cysteine and methionine metabolism* | 0.001 | 0.067 | map00270 |
| 6 | Glycolysis or Gluconeogenesis* | 0.002 | 0.095 | map00010 |
| 7 | Pentose phosphate pathway | 0.002 | 0.022 | map00030 |
| 8 | Propanoate metabolism | 0.002 | 0.001 | map00640 |
| 9 | Butanoate metabolism* | 0.003 | 0.103 | map00650 |
| 10 | Galactose metabolism* | 0.004 | 0.003 | map00052 |
| 11 | Ascorbate and aldarate metabolism | 0.005 | 0.024 | map00053 |
| 12 | Glyoxylate and dicarboxylate metabolism* | 0.006 | 0.033 | map00630 |
| 13 | Taurine and hypotaurine metabolism | 0.010 | 0.054 | map00430 |
| 14 | Citrate cycle(TCA cycle)* | 0.010 | 0.105 | map00020 |
| 15 | Valine, leucine and isoleucine biosynthesis | 0.018 | 0.022 | map00290 |
| 16 | Glycerolipid metabolism* | 0.025 | 0.209 | map00561 |
| 17 | Pyruvate metabolism* | 0.025 | 0.320 | map00620 |
| 18 | Phenylalanine metabolism | 0.048 | 0 | map00360 |
Table 2 Significant pathways for serum metabolites
| No. | Pathway name | P | Impact | KEGG |
|---|---|---|---|---|
| 1 | Aminoacyl-tRNA biosynthesis* | 0 | 0.113 | map00970 |
| 2 | Alanine, aspartate and glutamate metabolism* | 0 | 0.234 | map00250 |
| 3 | Arginine and proline metabolism* | 0 | 0.175 | map00330 |
| 4 | Glycine, serine and threonine metabolism* | 0 | 0.233 | map00260 |
| 5 | Cysteine and methionine metabolism* | 0.001 | 0.067 | map00270 |
| 6 | Glycolysis or Gluconeogenesis* | 0.002 | 0.095 | map00010 |
| 7 | Pentose phosphate pathway | 0.002 | 0.022 | map00030 |
| 8 | Propanoate metabolism | 0.002 | 0.001 | map00640 |
| 9 | Butanoate metabolism* | 0.003 | 0.103 | map00650 |
| 10 | Galactose metabolism* | 0.004 | 0.003 | map00052 |
| 11 | Ascorbate and aldarate metabolism | 0.005 | 0.024 | map00053 |
| 12 | Glyoxylate and dicarboxylate metabolism* | 0.006 | 0.033 | map00630 |
| 13 | Taurine and hypotaurine metabolism | 0.010 | 0.054 | map00430 |
| 14 | Citrate cycle(TCA cycle)* | 0.010 | 0.105 | map00020 |
| 15 | Valine, leucine and isoleucine biosynthesis | 0.018 | 0.022 | map00290 |
| 16 | Glycerolipid metabolism* | 0.025 | 0.209 | map00561 |
| 17 | Pyruvate metabolism* | 0.025 | 0.320 | map00620 |
| 18 | Phenylalanine metabolism | 0.048 | 0 | map00360 |
| No. | Pathway name | P | Impact | KEGG |
|---|---|---|---|---|
| 1 | Propanoate metabolism# | 0 | 0.086 | map00640 |
| 2 | Pentose phosphate pathway | 0.003 | 0.022 | map00030 |
| 3 | Amino sugar and nucleotide sugar metabolism* | 0.009 | 0 | map00520 |
| 4 | Fatty acid biosynthesis | 0.011 | 0 | map00061 |
| 5 | Starch and sucrose metabolism* | 0.011 | 0.017 | map00500 |
| 6 | Citrate cycle(TCA cycle)* | 0.015 | 0.031 | map00020 |
| 7 | Alanine, aspartate and glutamate metabolism | 0.021 | 0.003 | map00250 |
| 8 | Valine, leucine and isoleucine biosynthesis# | 0.027 | 0.027 | map00290 |
| 9 | Pantothenate and CoA biosynthesis# | 0.027 | 0.073 | map00770 |
| 10 | Aminoacyl-tRNA biosynthesis# | 0.032 | 0 | map00970 |
| 11 | Glycolysis or Gluconeogenesis | 0.034 | 0 | map00010 |
Table 3 Significant pathways for liver metabolites
| No. | Pathway name | P | Impact | KEGG |
|---|---|---|---|---|
| 1 | Propanoate metabolism# | 0 | 0.086 | map00640 |
| 2 | Pentose phosphate pathway | 0.003 | 0.022 | map00030 |
| 3 | Amino sugar and nucleotide sugar metabolism* | 0.009 | 0 | map00520 |
| 4 | Fatty acid biosynthesis | 0.011 | 0 | map00061 |
| 5 | Starch and sucrose metabolism* | 0.011 | 0.017 | map00500 |
| 6 | Citrate cycle(TCA cycle)* | 0.015 | 0.031 | map00020 |
| 7 | Alanine, aspartate and glutamate metabolism | 0.021 | 0.003 | map00250 |
| 8 | Valine, leucine and isoleucine biosynthesis# | 0.027 | 0.027 | map00290 |
| 9 | Pantothenate and CoA biosynthesis# | 0.027 | 0.073 | map00770 |
| 10 | Aminoacyl-tRNA biosynthesis# | 0.032 | 0 | map00970 |
| 11 | Glycolysis or Gluconeogenesis | 0.034 | 0 | map00010 |
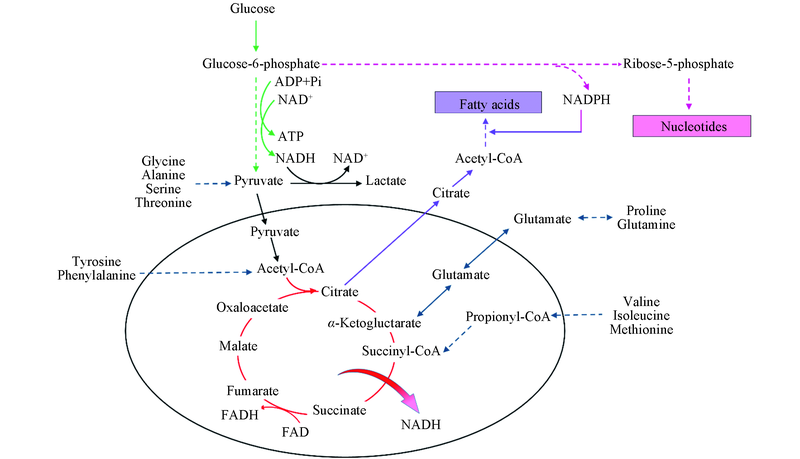
Fig.9 Network of metabolites and pathways in this workRed is TCA cycle; green is glycolysis; blue is amino acids into TCA cycle; pink is pentose phosphate pathway; purple is fatty acid synthesis.
| [1] | American Diabetes Association, Diabetes Care, 2014, 37(Suppl.1), S81—S90 |
| [2] | Taskinen M. R., Curr. Mol. Med., 2005, 5(3), 297—308 |
| [3] | Mathers C. D., Loncar D., PLoS Med., 2006, 3(11), e442 |
| [4] | International Diabetes Federation. IDF Diabetes Atlas: Eighth Edition., 2017, |
| [5] | Alberti K.G. M. M. Zimmet P. Z., Diabet Med., 1998, 15(7), 539—553 |
| [6] | de Lusignan S., Khunti K., Belsey J., Hattersley A., van Vlymen J., Gallagher H., Millett C., Hague N. J., Tomson C., Harris K., Majeed A., Diabet Med., 2010, 27(2), 203—209 |
| [7] | Zhang A. H., Qiu S., Xu H. Y., Sun H., Wang X. J., Clin. Chim. Acta, 2014, 429, 106—110 |
| [8] | Meiss E., Werner P., John C., Scheja L., Herbach N., Heeren J., Fischer M., Metabolomics, 2016, 12(3), 52 |
| [9] | Fiehn O., Plant Mol .Biol., 2002, 48, 155—171 |
| [10] | Vinayavekhin N., Homan E. A., Saghatelian A., ACS Chem. Biol., 2010, 5(1), 91—103 |
| [11] | Johnson C. H., Ivanisevic J., Benton H. P., Siuzdak G., Anal. Chem., 2015, 87(1), 147—156 |
| [12] | Oakes N. D., Cooney G. J., Camilleri S., Chisholm D. J., Kraegen E. W., Diabetes, 1997, 46(11), 1768—1774 |
| [13] | Lottenberg A. M., Afonso Mda S., Lavrador M. S., Machado R. M., Nakandakare E. R., J. Nutr. Biochem., 2012, 23(9), 1027—1040 |
| [14] | Kratz M., Baars T., Guyenet S., Eur. J. Nutr., 2013, 52(1), 1—24 |
| [15] | Flores M., Nutr. Res. Rev., 2007, 18(2), 175—182 |
| [16] | Cao L. J., Jiang P., Li H. D., He X., Zhu W. Y., Central South Pharmacy, 2014, 12(9), 844—847 |
| (曹玲娟, 江沛, 李焕德, 何昕, 朱文叶. 中南医学, 2014, 12(9), 844—847) | |
| [17] | Chiu K. C., Chu A., Go V. L. W., Saad M. F., Am. J. Clin. Nutr., 2004, 79(5), 820—825 |
| [18] | McCarty M. F., Thomas C. A., Med. Hypotheses, 2003, 61(5/6), 535—542 |
| [19] | Chagas C. E., Borges M. C., Martini L. A., Rogero M. M., Nutrients, 2012, 4(1), 52—67 |
| [20] | Wu S. M., Feng B., Cheng J. H., Li H. J., Fang J. J., Yan X. Z., Wei L., Dong F. T., Chem J Chinese Universities, 2012, 33(6), 1188—1194 |
| (吴胜明, 封波, 程建华, 李海静, 方均建, 颜贤忠, 魏来, 董方霆. 高等学校化学学报, 2012, 33(6), 1188—1194) | |
| [21] | Styczynski M. P., Moxley J. F., Tong L. V., Walther J. L., Jensen K. L., Stephanopoulos G. N., Anal. Chem., 2007, 79(3), 966—973 |
| [22] | Valenza F., Aletti G., Fossali T., Chevallard G., Sacconi F., Irace M., Gattinoni L., Crit. Care, 2005, 9(6), 588—593 |
| [23] | Felig P., Metabolism, 1973, 22(2), 179—207 |
| [24] | Attie A. D., Krauss R. M., Gray-Keller M. P., Brownlie A., Miyazaki M., Kastelein J. J., Lusis A. J., Stalenhoef A. F. H., Stoehr J. P., Hayden M. R., Ntambi J. M., J. Lipid. Res., 2002, 43(11), 1899—1907 |
| [25] | Lee J. J., Lambert J. E., Hovhannisyan Y., Ramos-Roman M. A., Trombold J. R., Wagner D. A., Parks E. J., Am. J. Clin. Nutr., 2015, 101(1), 34—43 |
| [26] | Wang T. J., Larson M. G., Vasan R. S., Cheng S., Rhee E. P., McCabe E., Lewis G. D., Fox C. S., Jacques P. F., Fernandez C., O’Donnell C. J., Carr S. A., Mootha V. K., Florez J. C., Souza A., Melander O., Clish C. B., Gerszten R. E., Nat. Med., 2011, 17(4), 448—453 |
| [27] | Huffman K. M., Shah S. H., Stevens R. D., Bain J. R., Muehlbauer M., Slentz C. A., Tanner C. J., Kuchibhatla M., Houmard J. A., Newgard C. B., Kraus W. E., Diabetes Care, 2009, 32(9), 1678—1683 |
| [28] | Satapati S., Sunny N. E., Kucejova B., Fu X., He T. T., Mendez-Lucas A., Shelton J. M., Perales J. C., Browning J. D., Burgess S. C., J. Lipid. Res., 2012, 53(6), 1080—1092 |
| [29] | Vessby B., Curr. Opin Lipidol, 2003, 14(1), 15—19 |
| [1] | HUANG Yu, GU Caiyun, WU Hanzhong, XIA Xiaoshuang, LI Xin. UPLC-QTOF-MS-based Metabolomics Study on Ischemic Stroke Patients† [J]. Chem. J. Chinese Universities, 2017, 38(10): 1742. |
| [2] | LI Penghui, DENG Lingli, LUO Jiao, LI Wei, NING Jing, DING Jianhua, WU Xiaoping. EESI-MS Detection and Statistical Analysis of Multi-batch of Exhaled Breath Metabolomics Data of Liver Failure Patients [J]. Chem. J. Chinese Universities, 2016, 37(4): 626. |
| [3] | ZHANG Ruixing, LIU Shu, PI Zifeng, SONG Fengrui, LIU Zhiqiang. Cell Metabolomics Study of Hg(Ⅱ) Ion Effect on the Metabolic Pathways of Cells [J]. Chem. J. Chinese Universities, 2014, 35(6): 1146. |
| [4] | MEI Zhen, CAI Wen-Sheng, SHAO Xue-Guang. Rapid Analysis of Complex Samples by GC-MS with Immune Algorithm [J]. Chem. J. Chinese Universities, 2013, 34(4): 819. |
| [5] | HUANG Min, GAO Jian-Yi, ZHAI Zhi-Gang, LIANG Qiong-Lin, WANG Yi-Ming, BAI Yan-Qiang, LUO Guo-An. Influence of Caffeine on the Sedative Effects of Promethazine Based on Pharmacokinetics and Metabolomics [J]. Chem. J. Chinese Universities, 2013, 34(4): 829. |
| [6] | WU Sheng-Ming, FENG Bo, CHENG Jian-Hua, LI Hai-Jing, FANG Jun-Jian, YAN Xian-Zhong, WEI Lai, DONG Fang-Ting. GC/TOF-MS-based Analytical Method of Endogenous Metabolites for Mouse Serum [J]. Chem. J. Chinese Universities, 2012, 33(06): 1188. |
| [7] | DONG Ji-Yang, LI Wei, DENG Ling-Li, XU Jing-Jing, Julian L. Griffin, CHEN Zhong*. New Variable Scaling Method for NMR-based Metabolomics Data Analysis [J]. Chem. J. Chinese Universities, 2011, 32(2): 262. |
| [8] | CHEN Jun1, LIANG Qiong-Lin1, LI Hui2, LUO Guo-An1*, WANG Yi-Ming1. Detection of Testosterone, Dihydrotestosterone, Dehydroepiandrosterone in Emasculated Rat Prostate via GC/MRM [J]. Chem. J. Chinese Universities, 2008, 29(11): 2155. |
| [9] | HUANG Bao-Yong, PAN Can-Beng, WANG Yi-Ru, CAO Zhe, ZHANG Wei, LI Wen-Ming, JIANG Shu-Ren. Rapid Determination of Pesticide Multiresidues in Vegetable by Gas Chromatography\|mass Spectrometry and Compensation for Matrix Effect with Protectants [J]. Chem. J. Chinese Universities, 2006, 27(2): 227. |
| [10] | YI Lun-Zhao, LIANG Yi-Zeng, ZENG Zhong-Da, YUAN Da-Lin, WANG Ping. AMWFA Method Applied to Comparative Analysis of Two-dimensional Data with Overlapped Peaks [J]. Chem. J. Chinese Universities, 2006, 27(11): 2052. |
| [11] | SHEN Peng, KANG Yu-Fei, CHENG Yi-Yu . Pattern Feature Discovery for Metabonomics of Breast Cancer and HPLC/MS/MS Analysis of Characteristic Metabolites [J]. Chem. J. Chinese Universities, 2005, 26(10): 1798. |
| [12] | WANG Yi-Ru, WANG Xiao-Ru, Frank S. C. Lee. Quantitative Analysis by Solid-phase Microextraction in Nonequilibrium Situations [J]. Chem. J. Chinese Universities, 1999, 20(5): 699. |
| [13] | WANG Rui, CHEN Yao-Zu. Analysis of Volatile Oil of Aconitum Sungpanense and A. karakclicum var. patentipilum by GC/MS [J]. Chem. J. Chinese Universities, 1992, 13(8): 1087. |
| [14] | XU You-xuan, ZHU Shao-tang, ZHANG Chang-jiu, WU Yun, ZHANG You-jie. Determination of Morphine in Urine of Drug Addicts by Gas Chromatography-Mass Spectrometry [J]. Chem. J. Chinese Universities, 1992, 13(5): 608. |
| [15] | Li Zhaolin, Chen Ning, Xue Dunyuan, Li Haiquan, Chen Yaozu . Study on Chemical Constituents of Volatile Oil of Fresh Flowers of Elaeagnus Angustifolia L [J]. Chem. J. Chinese Universities, 1989, 10(8): 804. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||