

Chem. J. Chinese Universities ›› 2015, Vol. 36 ›› Issue (11): 2226.doi: 10.7503/cjcu20150628
• Physical Chemistry • Previous Articles Next Articles
WANG Yan2, ZHENG Qingchuan1,2,*( )
)
Received:2015-08-07
Online:2015-11-10
Published:2015-10-23
Contact:
ZHENG Qingchuan
E-mail:zhengqc@jlu.edu.cn
CLC Number:
TrendMD:
WANG Yan, ZHENG Qingchuan. Molecular Dynamics Simulation on the Conformational Changes of CYP2E1 Enzyme Under Different Concentrations of Ethyl Alcohol[J]. Chem. J. Chinese Universities, 2015, 36(11): 2226.
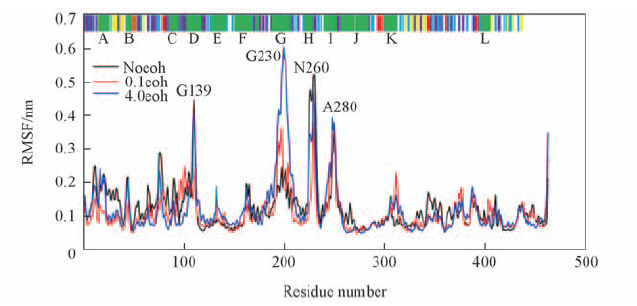
Fig.2 Root-mean-square fluctuations(RMSF) of noeoh, 0.1eoh and 4.0eoh systems Noeoh system is represented by black lines, 0.1eoh system red lines and 4.0eoh system blue lines. The results from DSSP are shown in the top panel of this Figure. Helices are indicated in green(and labeled at the top of Figure), β-sheets in yellow and turns in purple. A, B etc. stand for the well-known helix of the CYPs family.
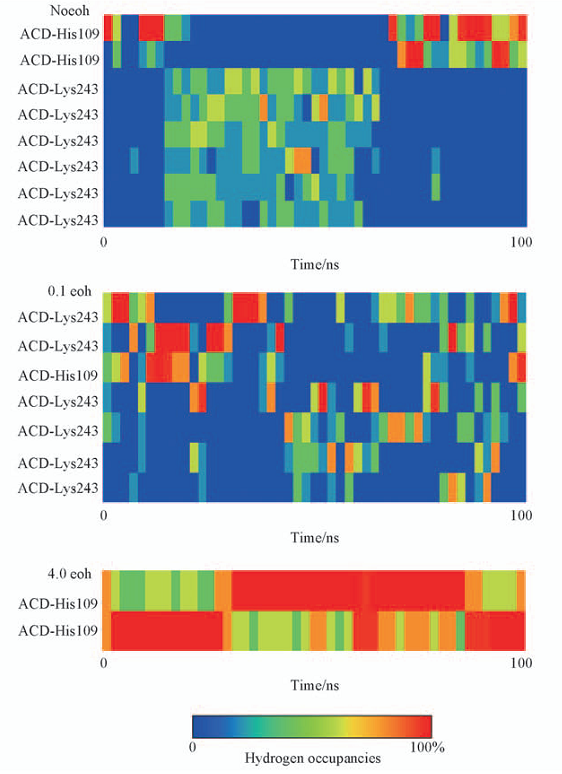
Fig.3 Percentage of occupancy of hydrogen bonds in different systems All these hydrogen bonds are monitored during the whole simulation time of each system.
| System | H-Bond | Occupancy(%) | Distance/nm | Angle/(°) |
|---|---|---|---|---|
| Noeoh | ACD@O2-H109@HE2 | 31.5 | 0.282 | 24.24 |
| ACD@O1-H109@HE2 | 17.4 | 0.289 | 28.27 | |
| ACD@O2-K243@HZ1 | 15.2 | 0.287 | 26.96 | |
| ACD@O2-K243@HZ3 | 14.6 | 0.285 | 25.47 | |
| ACD@O2-K243@HZ2 | 12.5 | 0.288 | 28.99 | |
| ACD@O1-K243@HZ1 | 11.3 | 0.296 | 33.47 | |
| ACD@O1-K243@HZ2 | 9.7 | 0.292 | 33.05 | |
| ACD@O1-K243@HZ3 | 9.0 | 0.297 | 37.29 | |
| 0.1eoh | ACD@O1-K243@HZ1 | 29.3 | 0.278 | 20.20 |
| ACD@O1-K243@HZ2 | 24.3 | 0.277 | 20.98 | |
| ACD@O2-H109@HE2 | 20.8 | 0.278 | 36.56 | |
| ACD@O1-K243@HZ3 | 19.3 | 0.280 | 21.44 | |
| ACD@O2-K243@HZ1 | 13.8 | 0.284 | 21.93 | |
| ACD@O2-K243@HZ3 | 11.2 | 0.283 | 22.37 | |
| ACD@O2-K243@HZ2 | 7.0 | 0.292 | 25.33 | |
| 4.0eoh | ACD@O1-H109@HE2 | 76.6 | 0.285 | 26.60 |
| ACD@O2-H109@HE2 | 74.7 | 0.290 | 29.89 |
Table 1 Properties of hydrogen bonds between AA and its adjacent residues, including occupancy, distance and angle
| System | H-Bond | Occupancy(%) | Distance/nm | Angle/(°) |
|---|---|---|---|---|
| Noeoh | ACD@O2-H109@HE2 | 31.5 | 0.282 | 24.24 |
| ACD@O1-H109@HE2 | 17.4 | 0.289 | 28.27 | |
| ACD@O2-K243@HZ1 | 15.2 | 0.287 | 26.96 | |
| ACD@O2-K243@HZ3 | 14.6 | 0.285 | 25.47 | |
| ACD@O2-K243@HZ2 | 12.5 | 0.288 | 28.99 | |
| ACD@O1-K243@HZ1 | 11.3 | 0.296 | 33.47 | |
| ACD@O1-K243@HZ2 | 9.7 | 0.292 | 33.05 | |
| ACD@O1-K243@HZ3 | 9.0 | 0.297 | 37.29 | |
| 0.1eoh | ACD@O1-K243@HZ1 | 29.3 | 0.278 | 20.20 |
| ACD@O1-K243@HZ2 | 24.3 | 0.277 | 20.98 | |
| ACD@O2-H109@HE2 | 20.8 | 0.278 | 36.56 | |
| ACD@O1-K243@HZ3 | 19.3 | 0.280 | 21.44 | |
| ACD@O2-K243@HZ1 | 13.8 | 0.284 | 21.93 | |
| ACD@O2-K243@HZ3 | 11.2 | 0.283 | 22.37 | |
| ACD@O2-K243@HZ2 | 7.0 | 0.292 | 25.33 | |
| 4.0eoh | ACD@O1-H109@HE2 | 76.6 | 0.285 | 26.60 |
| ACD@O2-H109@HE2 | 74.7 | 0.290 | 29.89 |
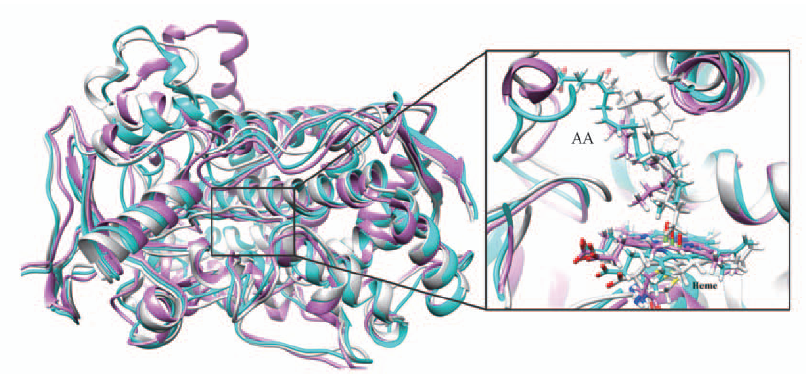
Fig.5 Alignment of three representative structures for noeoh, 0.1eoh and 4.0eoh systems All the structures are in ribbon style. Noeoh is colored in light gray, 0.1eoh in deep sky cyan and 4.0eoh in orchid. A close view for AA molecule and heme are presented on the right.
| Structure | Population(%) | RMSD to noeoh structure/nm |
|---|---|---|
| C4 | 41.2 | 0.116 |
| C3 | 38.5 | 0.132 |
Table 2 Populations of representative structures and their RMSD values compared to noeoh representative structure
| Structure | Population(%) | RMSD to noeoh structure/nm |
|---|---|---|
| C4 | 41.2 | 0.116 |
| C3 | 38.5 | 0.132 |
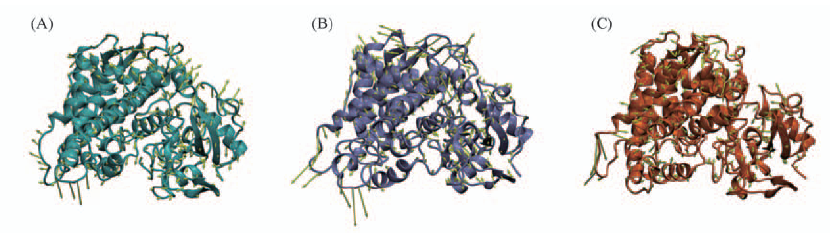
Fig.6 Porcupine plots in stereo showing noeoh(A), 0.1eoh(B) and 4.0eoh(C) with cones signifying the first eigenvectors movements Noeoh system is colored in cyan, N219D in purple and S366C in orange.
| System | ΔEele/(kJ·mol-1) | ΔEvdw/(kJ·mol-1) | ΔGGB/(kJ·mol-1) | ΔGSA/(kJ·mol-1) | -TΔS/(kJ·mol-1) | Δ |
|---|---|---|---|---|---|---|
| Noeoh | -633.80 | -162.00 | 675.70 | -22.27 | 75.72 | -66.64 |
| 0.1eoh | -1040.14 | -156.35 | 995.97 | -29.22 | 180.25 | -49.44 |
| 4.0eoh | -1047.84 | -165.18 | 1008.24 | -18.63 | 209.93 | -13.48 |
Table 3 Binding free energies and their components for noeoh, 0.1eoh and 4.0eoh
| System | ΔEele/(kJ·mol-1) | ΔEvdw/(kJ·mol-1) | ΔGGB/(kJ·mol-1) | ΔGSA/(kJ·mol-1) | -TΔS/(kJ·mol-1) | Δ |
|---|---|---|---|---|---|---|
| Noeoh | -633.80 | -162.00 | 675.70 | -22.27 | 75.72 | -66.64 |
| 0.1eoh | -1040.14 | -156.35 | 995.97 | -29.22 | 180.25 | -49.44 |
| 4.0eoh | -1047.84 | -165.18 | 1008.24 | -18.63 | 209.93 | -13.48 |
| [1] | Danielson P., Curr. Drug Metab., 2002, 3, 561—597 |
| [2] | Nelson D. R., Arch. Biochem. Biophys., 1999, 369, 1—10 |
| [3] | Watanabe K. P., Kawai Y. K., Ikenaka Y., Kawata M., Ikushiro S. I., Sakaki T., Ishizuka M., PLoS One, 2013, 8, e75689 |
| [4] | Scott E. E., Halpert J. R., Trends Biochem. Sci., 2005, 30, 5—7 |
| [5] | Park H., Lee S., Suh J., J. Am. Chem. Soc., 2005, 127, 13634—13642 |
| [6] | Walsh A. A., Szklarz G. D., Scott E. E., J. Biol. Chem., 2013, 288, 12932—12943 |
| [7] | Miller G. P., Expert. Opin. Drug Met., 2008, 4, 1053—1064 |
| [8] | Nanji A. A., Zhao S., Sadrzadeh S., Dannenberg A. J., Tahan S. R., Waxman D. J., Alcoholism: Clinical and Experimental Research, 1994, 18, 1280—1285 |
| [9] | Barnett C., Rudd S., Flatt P., Ioannides C., Biochem. Pharmacol., 1993, 45, 313—319 |
| [10] | Johansson I., Ekstroem G., Scholte B., Puzycki D., Joernvall H., Ingelman-Sundberg M., Biochemistry, 1988, 27, 1925—1934 |
| [11] | Raucy J. L., Lasker J., Ozaki K., Zoleta V., Toxicol. Sci., 2004, 79, 233—241 |
| [12] | DeVore N. M., Meneely K. M., Bart A. G., Stephens E. S., Battaile K. P., Scott E. E., Febs. J., 2012, 279, 1621—1631 |
| [13] | Li J., Wei D. Q., Wang J. F., Li Y. X., J. Chem. Inf. Model., 2011, 51, 3217—3225 |
| [14] | Porubsky P. R., Battaile K. P., Scott E. E., J. Biol. Chem., 2010, 285, 22282—22290 |
| [15] | Caro A. A., Cederbaum A. I., Free Radical Biol. Med., 2006, 40, 364—375 |
| [16] | Wu D., Cederbaum A. I., Hepatology, 2002, 35, 1420—1430 |
| [17] | Cowpland C., Su G.M., Murray M., Puddey I. B., Croft K. D., Clin. Exp. Pharmacol. P., 2006, 33, 183—188 |
| [18] | Lieber C. S., DeCarli L. M., Science, 1968, 162, 917—918 |
| [19] | Discovery Studio Version 2.5, Accelrys Inc., San Diego, CA, USA, 2009 |
| [20] | Olsson M. H., Søndergaard C. R., Rostkowski M., Jensen J. H., J. Chem. Theory Comput., 2011, 7, 525—537 |
| [21] | Case D.E., Darden T., Cheatham I. T., Simmerling C., Wang J., Duke R., Luo R., Walker R., Zhang W., Merz K., AMBER11, University of California, San Francisco, 2010 |
| [22] | Frisch M.J., Trucks G. W., Schlegel H. B., Scuseria G.E., Robb M. A., Cheeseman J. R., Scalmani G., Barone V., Mennucci B., Peters-son G. A., Nakatsuji H., Caricato M., Li X., Hratchian H. P., Izmay-lov A. F., Bloino J., Zheng G., Sonnenberg J. L., Hada M., Ehara M., Toyota K., Fukuda R., Hasegawa J., Ishida M., Nakajima T., Honda Y., Kitao O., Nakai H., Vreven T., Montgomery J. A. Jr., Peralta J. E., Ogliaro F., Bearpark M., Heyd J. J., Brothers E., Kudin K. N., Staro-verov V. N., Kobayashi R., Normand J., Raghavachari K., Rendell A., Burant J. C., Iyengar S. S., Tomasi J., Cossi M., Rega N., Millam N. J., Klene M., Knox J. E., Cross J. B., Bakken V., Adamo C., Jaramillo J., Gomperts R., Stratmann R. E., Yazyev O., Austin A. J., Cammi R., Pomelli C., Ochterski J. W., Martin R. L., Morokuma K., Zakrzewski V. G., Voth G. A., Salvador P., Dannenberg J. J., Dapprich S., Daniels A. D., Farkas Ö., Foresman J. B., Ortiz J. V., Cioslowski J., Fox D. J., Gaussian 09. D01, Gaussian Inc., Wallingford CT, 2009 |
| [23] | Becke A. D., J. Chem. Phys., 1993, 98, 5648—5652 |
| [24] | Lee C., Yang W., Parr R. G., Phys. Rev. B, 1988, 37, 785—789 |
| [25] | Rydberg P., Olsen L., Norrby P. O., Ryde U., J. Chem. Theory Comput., 2007, 3, 1765—1773 |
| [26] | Cui Y. L., Zheng Q. C., Zhang J. L., Zhang H. X., Biopolymers, 2015, 103, 53—66 |
| [27] | Hornak V., Abel R., Okur A., Strockbine B., Roitberg A., Simmerling C., Proteins: Structure Function and Bioinformatics, 2006, 65, 712—725 |
| [28] | Jorgensen W. L., Chandrasekhar J., Madura J. D., Impey R. W., Klein M. L., J. Chem. Phys., 1983, 79, 926—935 |
| [29] | Shao J., Tanner S. W., Thompson N., Cheatham T. E., J. Chem. Theory Comput., 2007, 3, 2312—2334 |
| [30] | Humphrey W., Dalke A., Schulten K., J. Mol. Graphics Modell., 1996, 14, 33—38 |
| [31] | Pettersen E. F., Goddard T. D., Huang C. C., Couch G. S., Greenblatt D. M., Meng E. C., Ferrin T. E., J. Comput. Chem., 2004, 25, 1605—1612 |
| [32] | DeLano W. L., PyMOL, Schrodinger, New York, 2002 |
| [33] | Amadei A., Linssen A., Berendsen H. J., Proteins: Structure Function , and Bioinformatics, 1993, 17, 412—425 |
| [34] | Bakan A., Meireles L. M., Bahar I., Bioinformatics, 2011, 27, 1575—1577 |
| [35] | Xue Q., Zhang J. L., Zheng Q. C., Cui Y. L., Chen L., Chu W. T., Zhang H. X., Langmuir, 2013, 29, 11135—11144 |
| [36] | Cui Y.L., Zheng Q. C., Zhang J. L., Xue Q., Wang Y., Zhang H. X.,J. Chem. Inf. Model., 2013, 3308—3317 |
| [37] | Wang Y., Zheng Q. C., Kong C. P., Tian Y., Zhan J., Zhang J. L., Zhang H. X., Mol. Biosyst., 2015, 11(1), 252—261 |
| [38] | Swanson J. M. J., Henchman R. H., McCammon J. A., Biophysical Journal, 2004, 86, 67—74 |
| [39] | Hou T. J., Zhang W., Case D. A., Wang W., J. Mol. Biol., 2008, 376, 1201—1214 |
| [40] | Joosten R. P., Te Beek T. A., Krieger E., Hekkelman M. L., Hooft R. W., Schneider R., Sander C., Vriend G., Nucleic Acids Res., 2011, 39, D411—D419 |
| [41] | Wu Y. J., Cui Y. L., Zheng Q. C., Zhang H. X., Chem. J. Chinese Universities, 2014, 35(12), 2605—2611 |
| [42] | Wu Y. J., Zheng Q. C., Xu Y., Chu W. L., Cui Y., Wang Y., Zhang H. X., Chem. Res. Chinese Universities, 2014, 30(6), 1011—1017 |
| [43] | Meng X. Y., Li Z., Niu R. J., Zhang H. X., Zheng Q. C., Chem. Res. Chinese Universities, 2012, 28(1), 137—141 |
| [1] | LIU Bei, GAO Pei, LI Shenshen, XIAO Yunqin, XIAO Jijun. Molecular Dynamics Investigation of Adhesion Between CL-20/TNT Co-crystal Surfaces and Adhesives of PLA, PCL, P(LA-co-CL)† [J]. Chem. J. Chinese Universities, 2018, 39(11): 2556. |
| [2] | REN Jiayi, YANG Zhiwei, ZHENG Nianjue, LI Junqi, YANG Chunlong, LIN Shujian, YANG Bing, HUANG Junqing, LIAO Huaxin, YUAN Xiaohui, OU Shiyi. Effect of Force Fields and Water Models on EGFRvⅢ-MR1(scFv)Complex by Molecular Dynamics Simulation, MM-PBSA Calculation, and ITC Experiment† [J]. Chem. J. Chinese Universities, 2017, 38(11): 2070. |
| [3] | LIU Dong-Mei, ZHAO Li, XIAO Ji-Jun, CHEN Jun, JI Guang-Fu, ZHU Wei, ZHAO Feng, WU Qiang, XIAO He-Min. Sensitivity Criterion and Mechanical Properties Prediction of HMX and RDX Crystals at Different Temperatures Comparative Study with Molecular Dymamics Simulation [J]. Chem. J. Chinese Universities, 2013, 34(11): 2558. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||